Summary information and primary citation
- PDB-id
- 2zo0; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- ligase-DNA
- Method
- X-ray (2.19 Å)
- Summary
- Mouse np95 sra domain DNA specific complex 1
- Reference
- Hashimoto H, Horton JR, Zhang X, Bostick M, Jacobsen SE, Cheng X (2008): "The SRA domain of UHRF1 flips 5-methylcytosine out of the DNA helix." Nature, 455, 826-829. doi: 10.1038/nature07280.
- Abstract
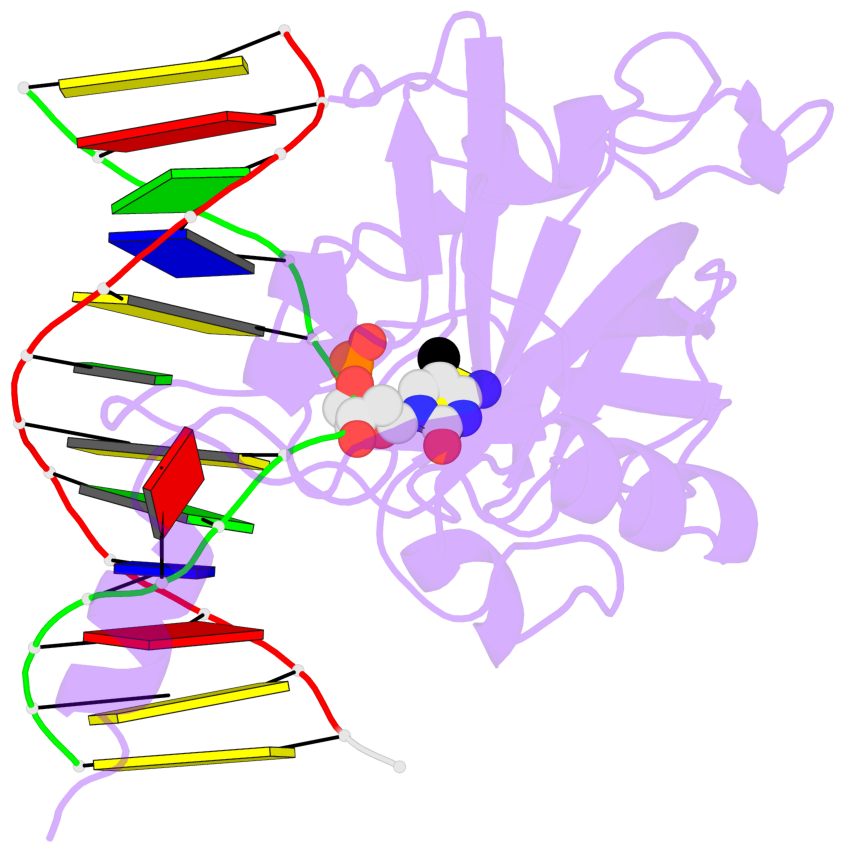
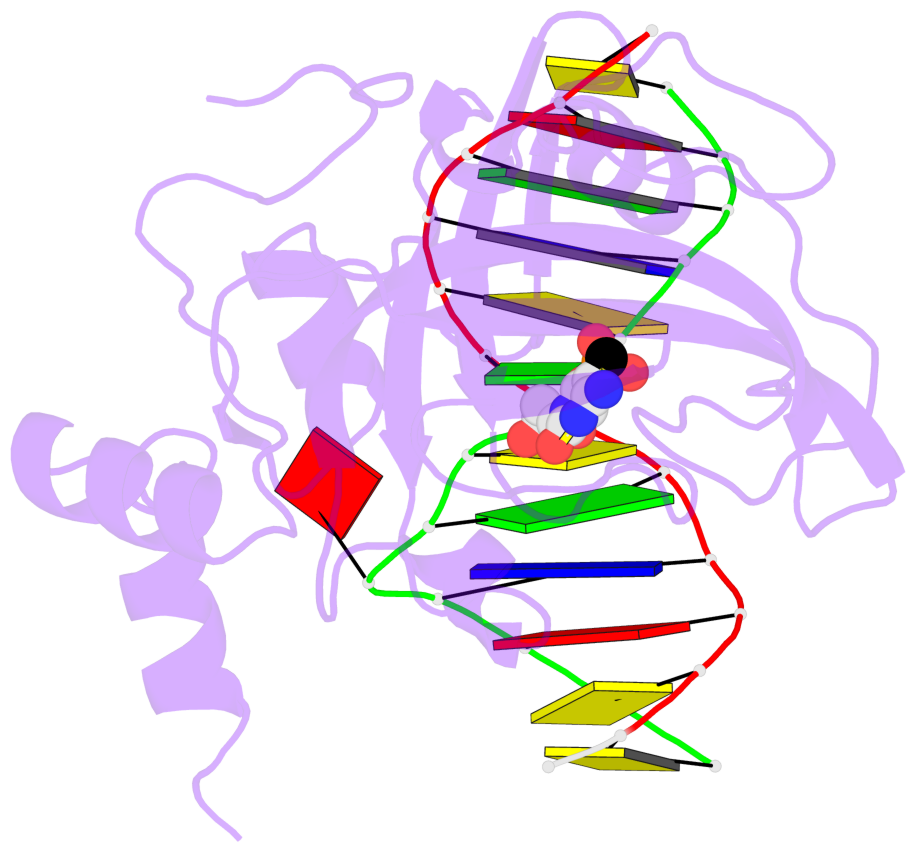
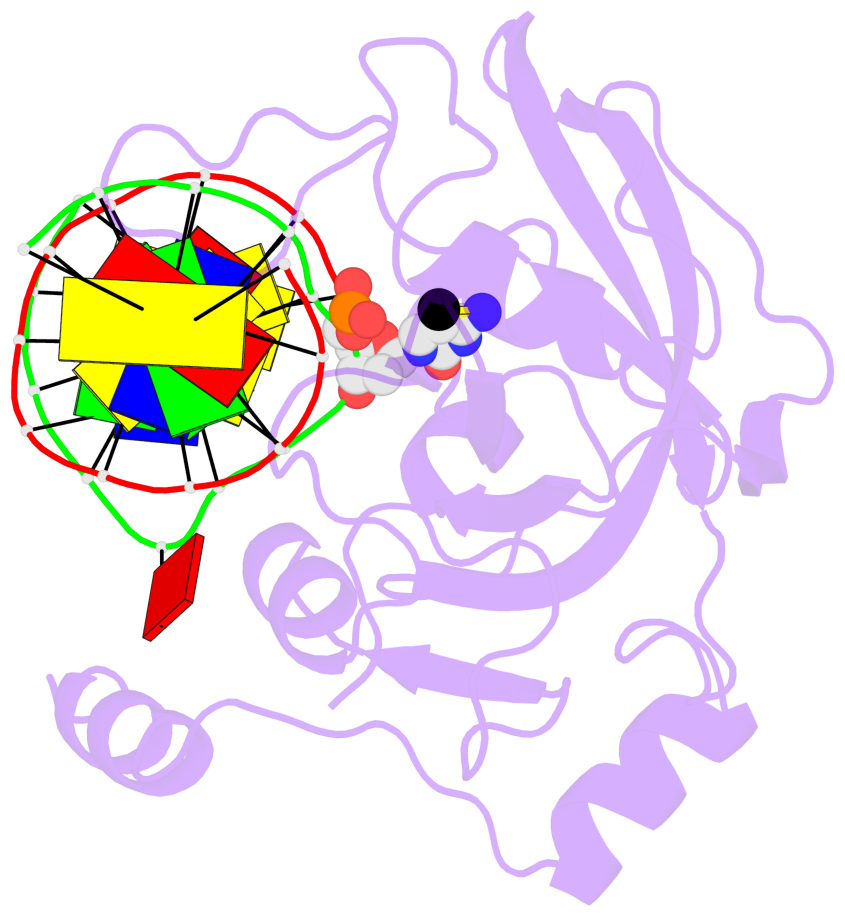
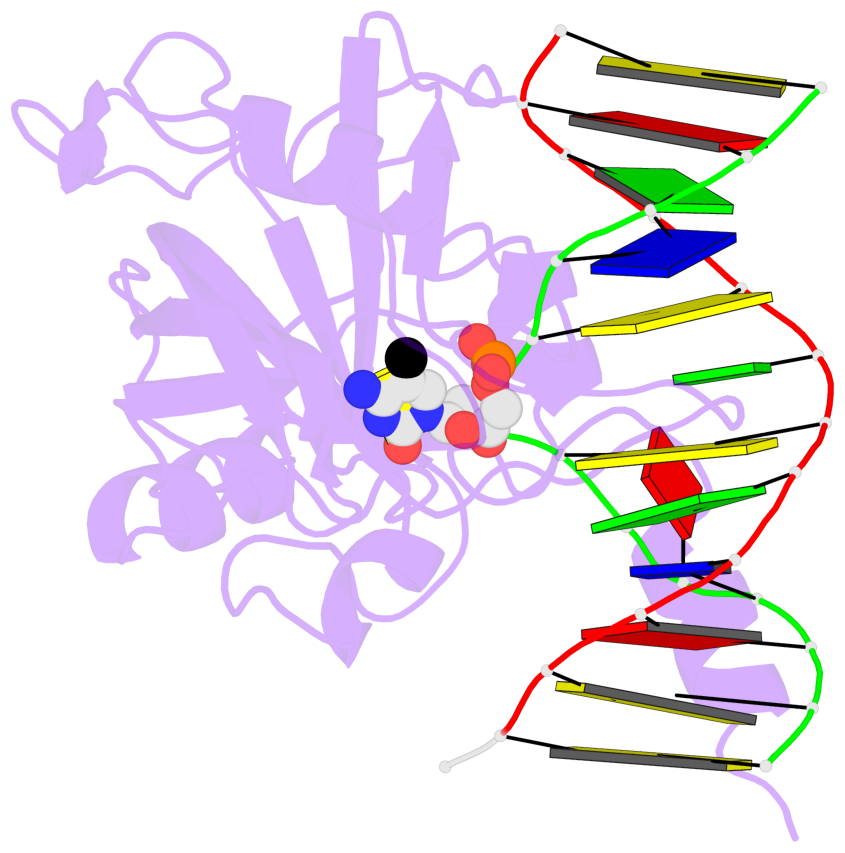
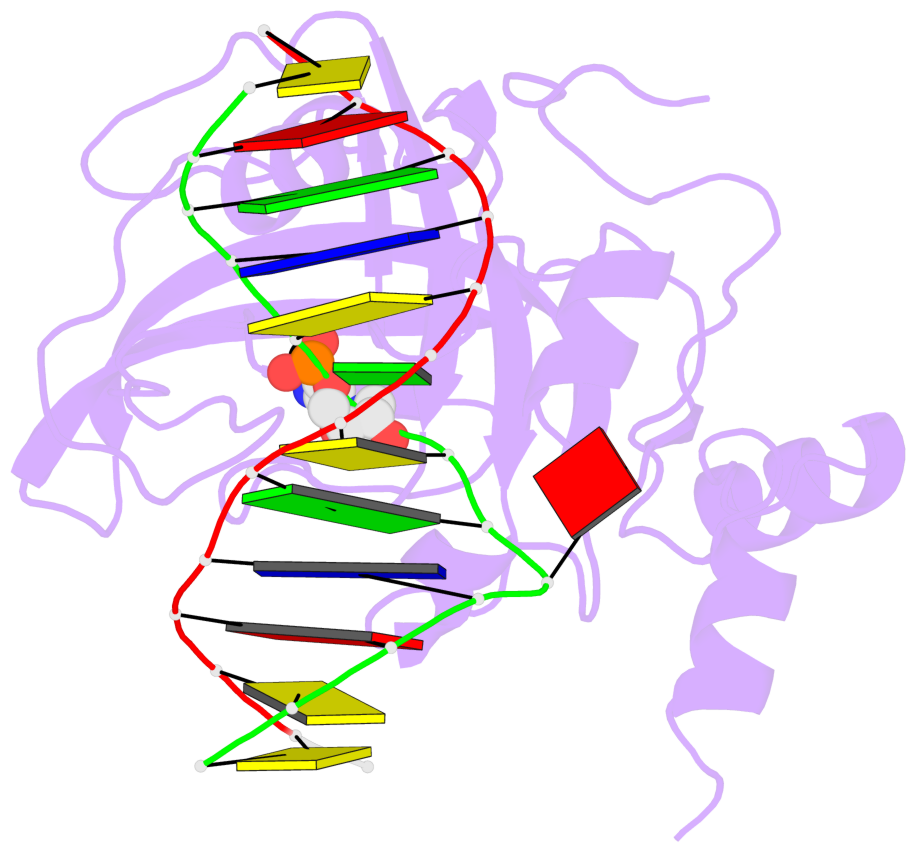
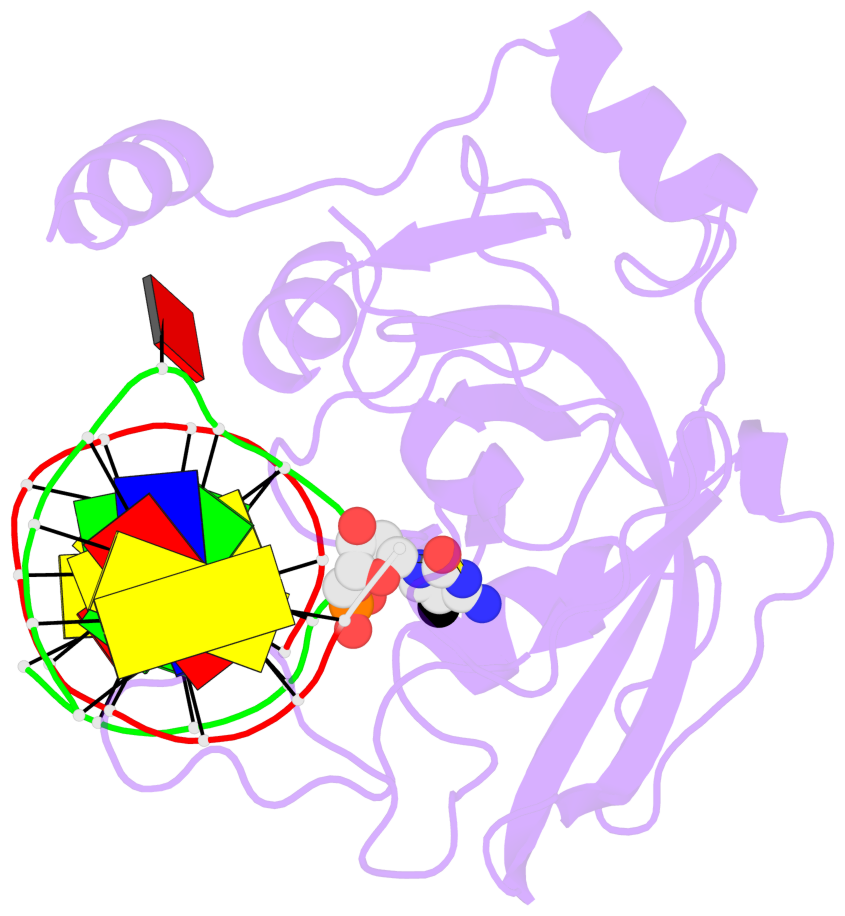
- Maintenance methylation of hemimethylated CpG dinucleotides at DNA replication forks is the key to faithful mitotic inheritance of genomic methylation patterns. UHRF1 (ubiquitin-like, containing PHD and RING finger domains 1) is required for maintenance methylation by interacting with DNA nucleotide methyltransferase 1 (DNMT1), the maintenance methyltransferase, and with hemimethylated CpG, the substrate for DNMT1 (refs 1 and 2). Here we present the crystal structure of the SET and RING-associated (SRA) domain of mouse UHRF1 in complex with DNA containing a hemimethylated CpG site. The DNA is contacted in both the major and minor grooves by two loops that penetrate into the middle of the DNA helix. The 5-methylcytosine has flipped completely out of the DNA helix and is positioned in a binding pocket with planar stacking contacts, Watson-Crick polar hydrogen bonds and van der Waals interactions specific for 5-methylcytosine. Hence, UHRF1 contains a previously unknown DNA-binding module and is the first example of a non-enzymatic, sequence-specific DNA-binding protein domain to use the base flipping mechanism to interact with DNA.
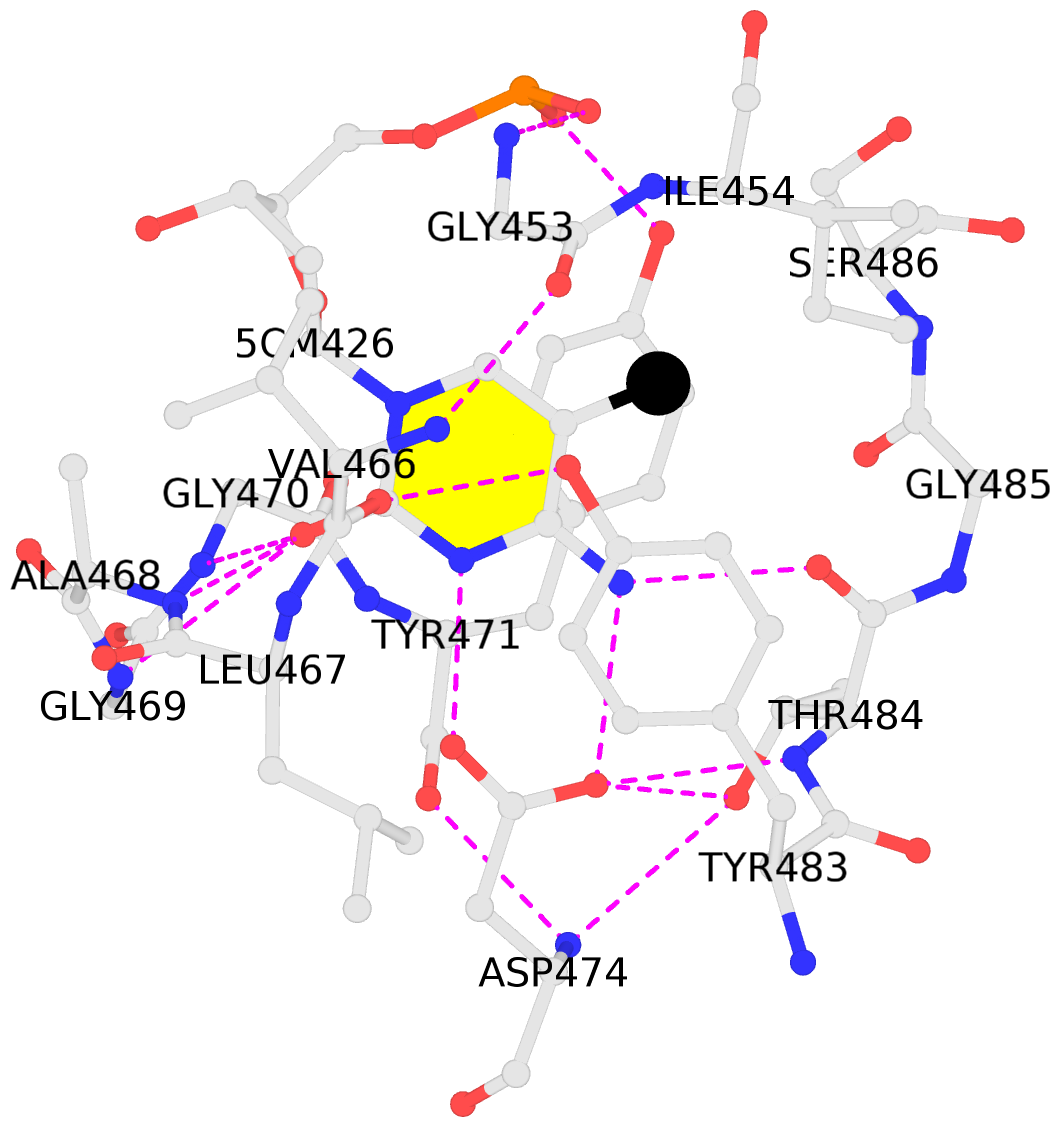
- The contacts include paired nucleotides (mostly a G in Watson-Crick G-C pairing), and
amino-acids within a 4.5-A distance cutoff to base atoms of 5mC.
- The structure is oriented in the base reference frame of 5mC, allowing for easy comparison
and direct superimposition between entries.
- The black sphere (•) denotes the 5-methyl carbon atom in 5mC.
No. 1 E.5CM426: stacking-with-B.TYR471 stacking-with-B.TYR483 not-WC-paired not-in-duplex |
 |
|