Summary information and primary citation
- PDB-id
- 3ssc; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein-DNA
- Method
- X-ray (2.1 Å)
- Summary
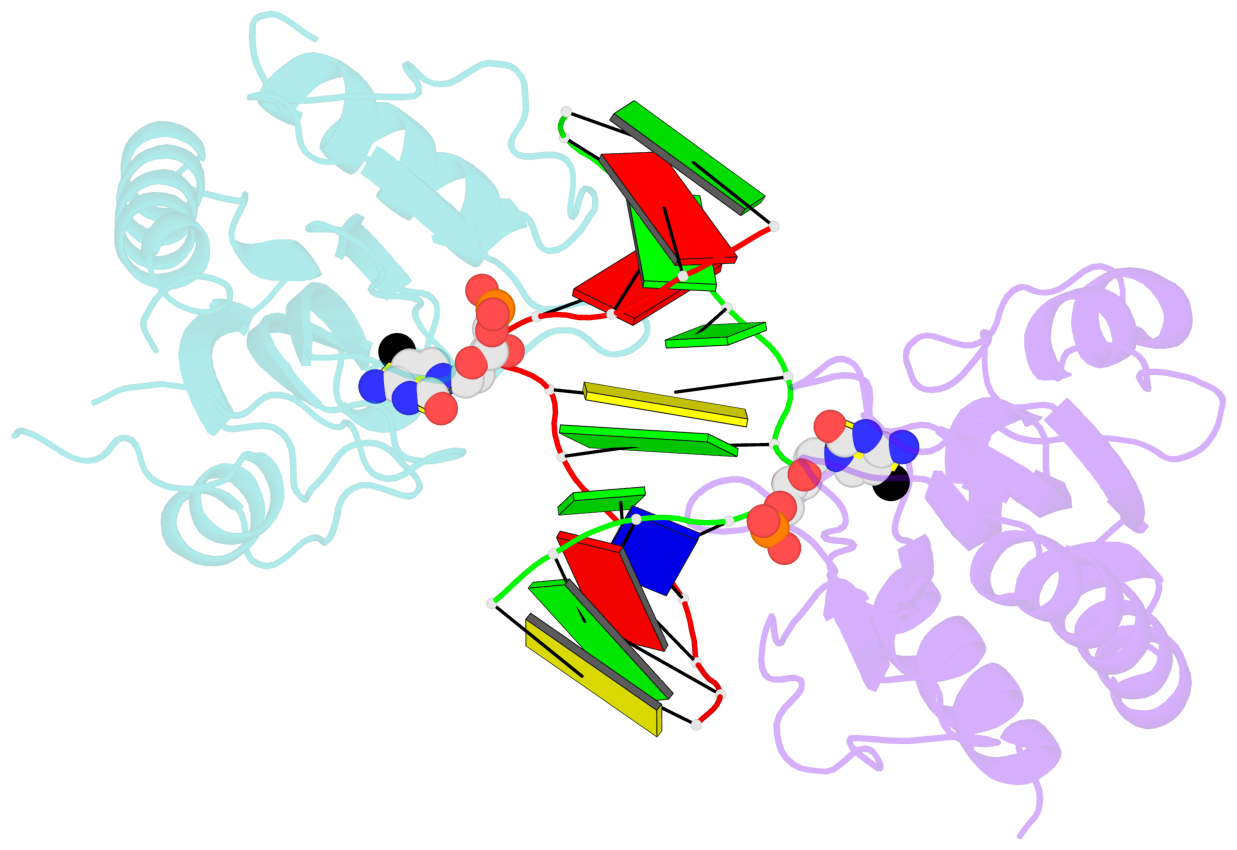
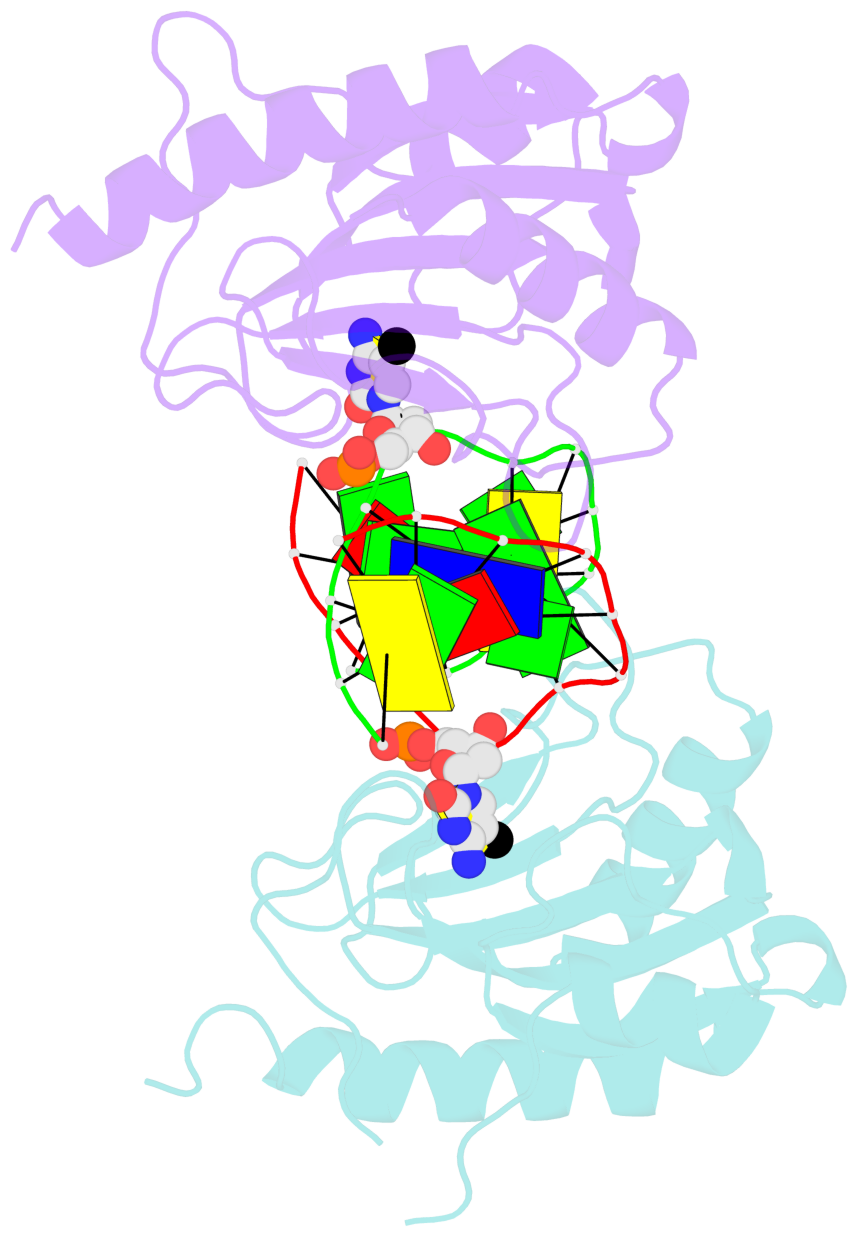
- DNA binding domain of restriction endonuclease bound to DNA
- Reference
- Sukackaite R, Grazulis S, Tamulaitis G, Siksnys V (2012): "The recognition domain of the methyl-specific endonuclease McrBC flips out 5-methylcytosine." Nucleic Acids Res., 40, 7552-7562. doi: 10.1093/nar/gks332.
- Abstract
- DNA cytosine methylation is a widespread epigenetic mark. Biological effects of DNA methylation are mediated by the proteins that preferentially bind to 5-methylcytosine (5mC) in different sequence contexts. Until now two different structural mechanisms have been established for 5mC recognition in eukaryotes; however, it is still unknown how discrimination of the 5mC modification is achieved in prokaryotes. Here we report the crystal structure of the N-terminal DNA-binding domain (McrB-N) of the methyl-specific endonuclease McrBC from Escherichia coli. The McrB-N protein shows a novel DNA-binding fold adapted for 5mC-recognition. In the McrB-N structure in complex with methylated DNA, the 5mC base is flipped out from the DNA duplex and positioned within a binding pocket. Base flipping elegantly explains why McrBC system restricts only T4-even phages impaired in glycosylation [Luria, S.E. and Human, M.L. (1952) A nonhereditary, host-induced variation of bacterial viruses. J. Bacteriol., 64, 557-569]: flipped out 5-hydroxymethylcytosine is accommodated in the binding pocket but there is no room for the glycosylated base. The mechanism for 5mC recognition employed by McrB-N is highly reminiscent of that for eukaryotic SRA domains, despite the differences in their protein folds.
- The contacts include paired nucleotides (mostly a G in Watson-Crick G-C pairing), and
amino-acids within a 4.5-A distance cutoff to base atoms of 5mC.
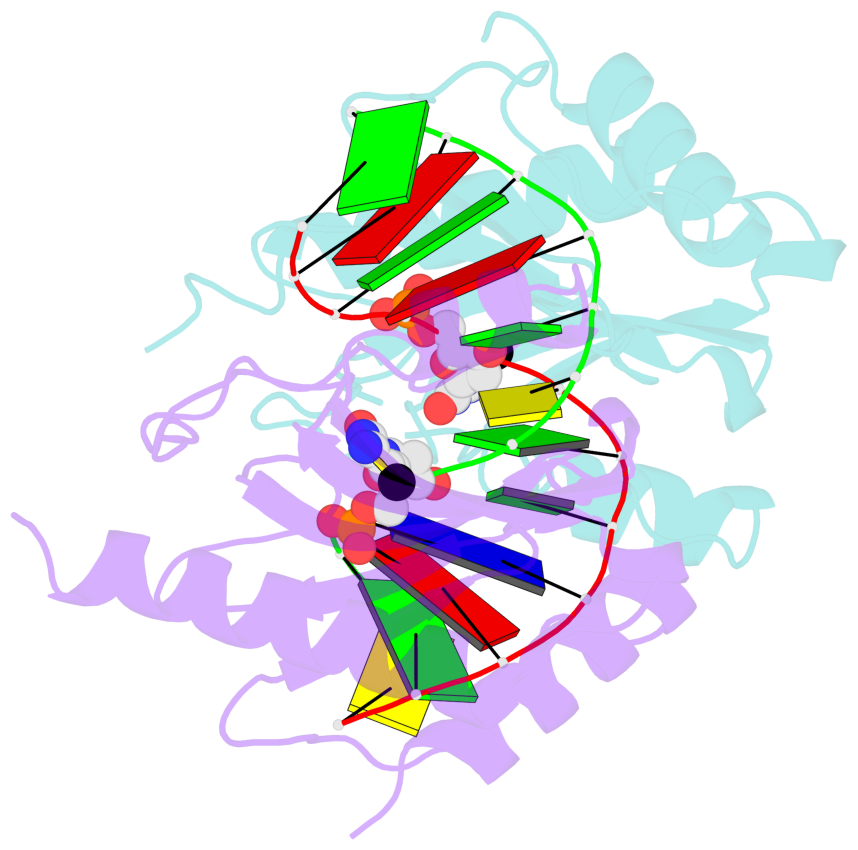
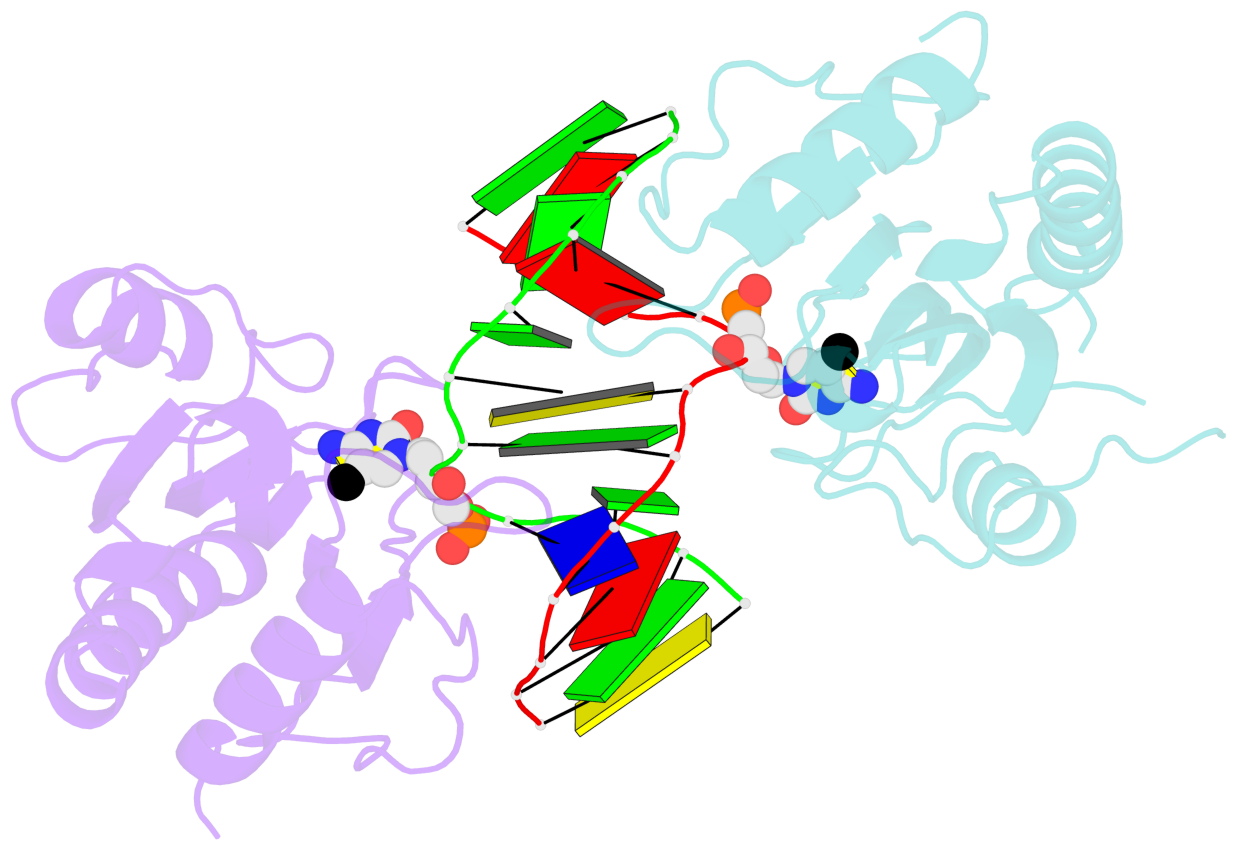
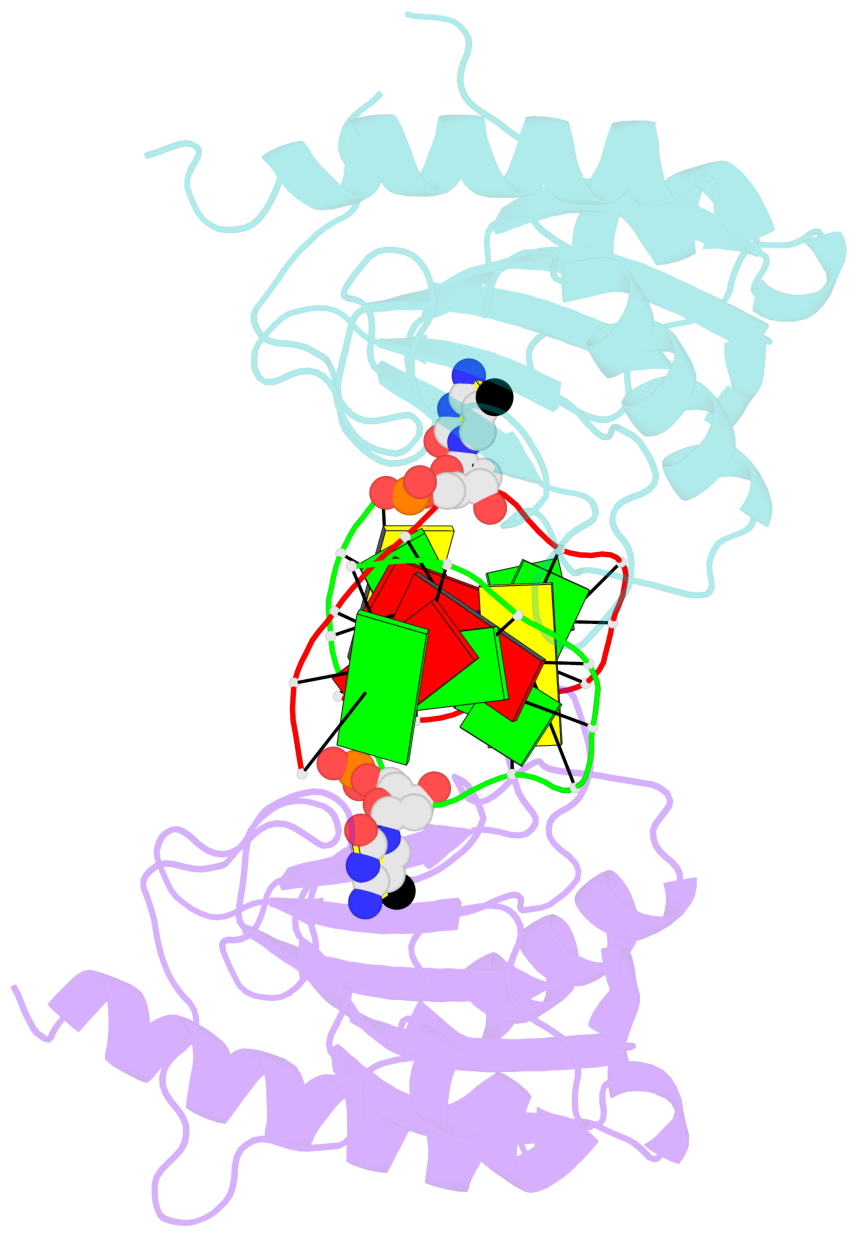
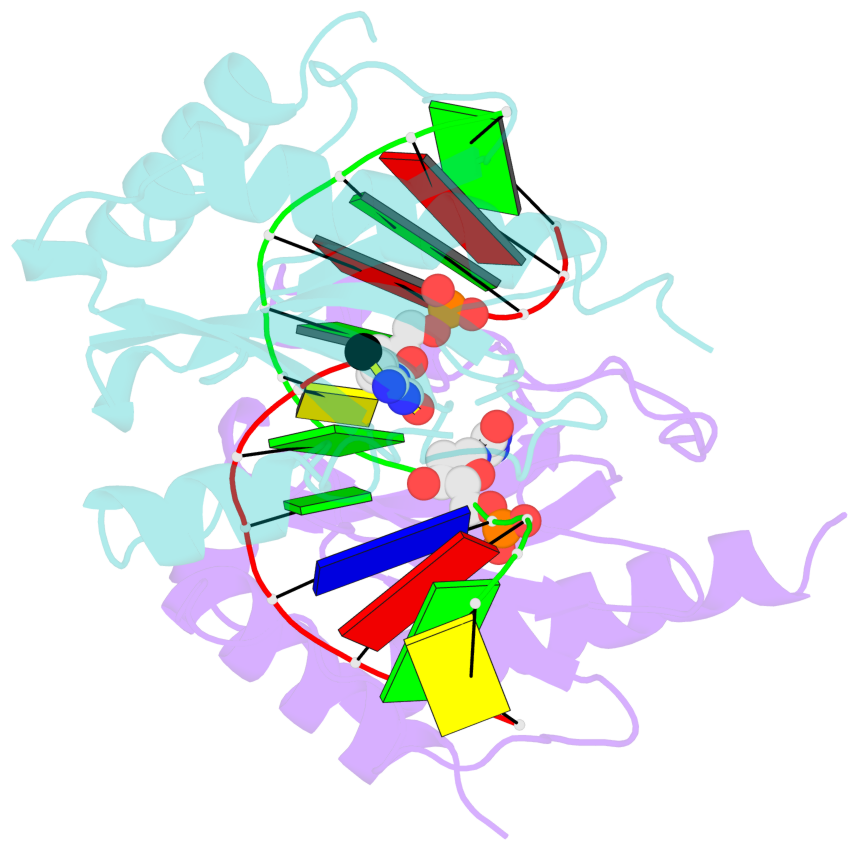
- The structure is oriented in the base reference frame of 5mC, allowing for easy comparison
and direct superimposition between entries.
- The black sphere (•) denotes the 5-methyl carbon atom in 5mC.
No. 1 C.5CM6: stacking-with-B.TYR117 not-WC-paired not-in-duplex |
 |
|
No. 2 D.5CM6: stacking-with-A.TYR117 not-WC-paired not-in-duplex |
 |
|