Summary information and primary citation
- PDB-id
- 6mht; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-DNA
- Method
- X-ray (2.05 Å)
- Summary
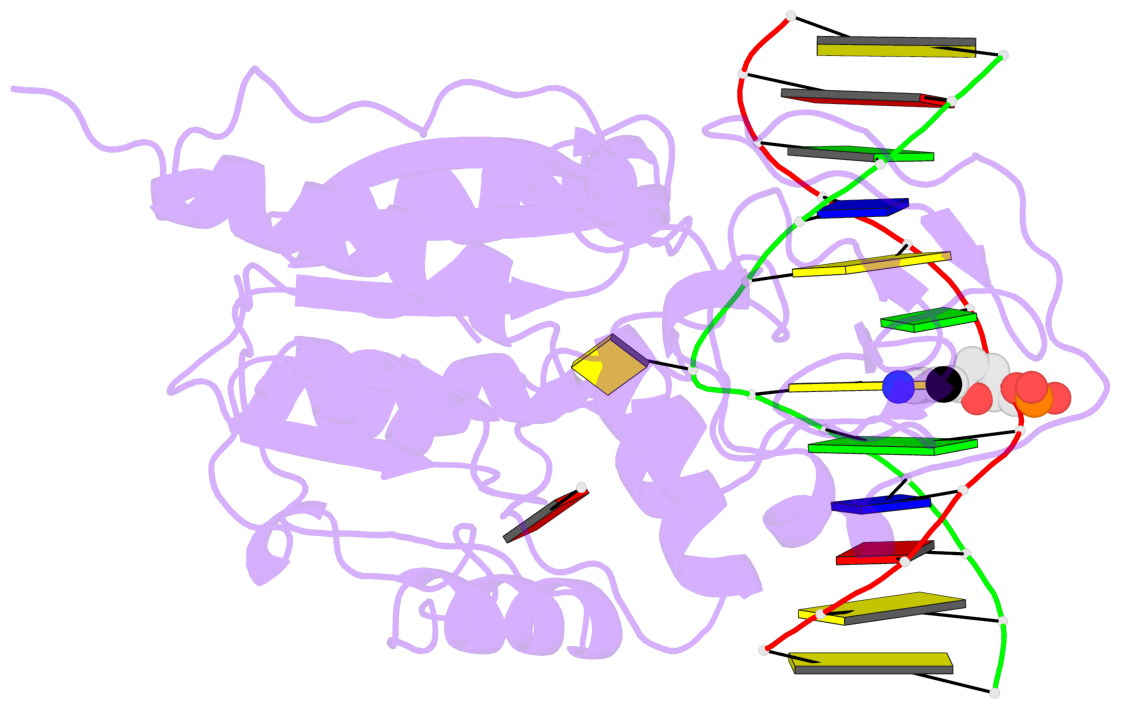
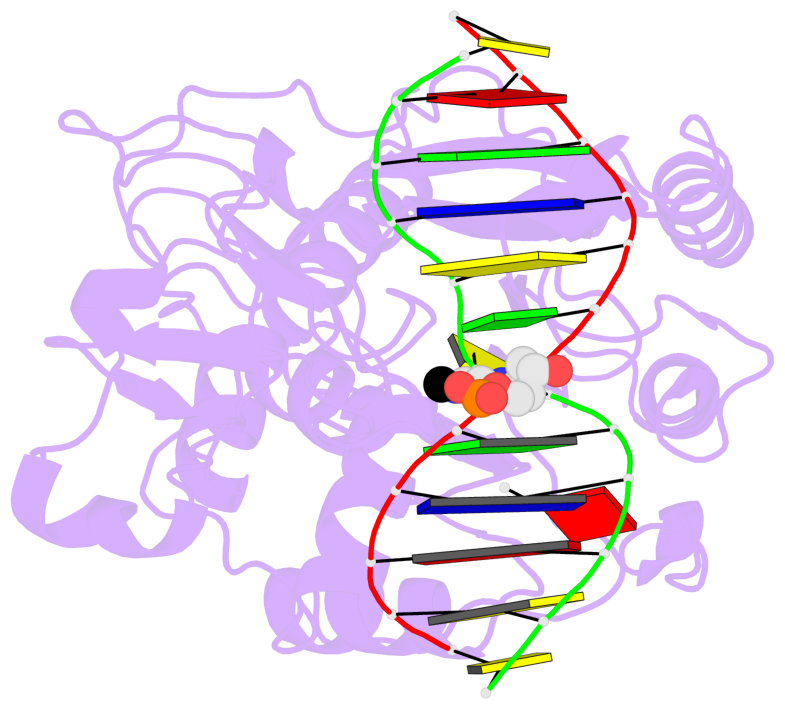
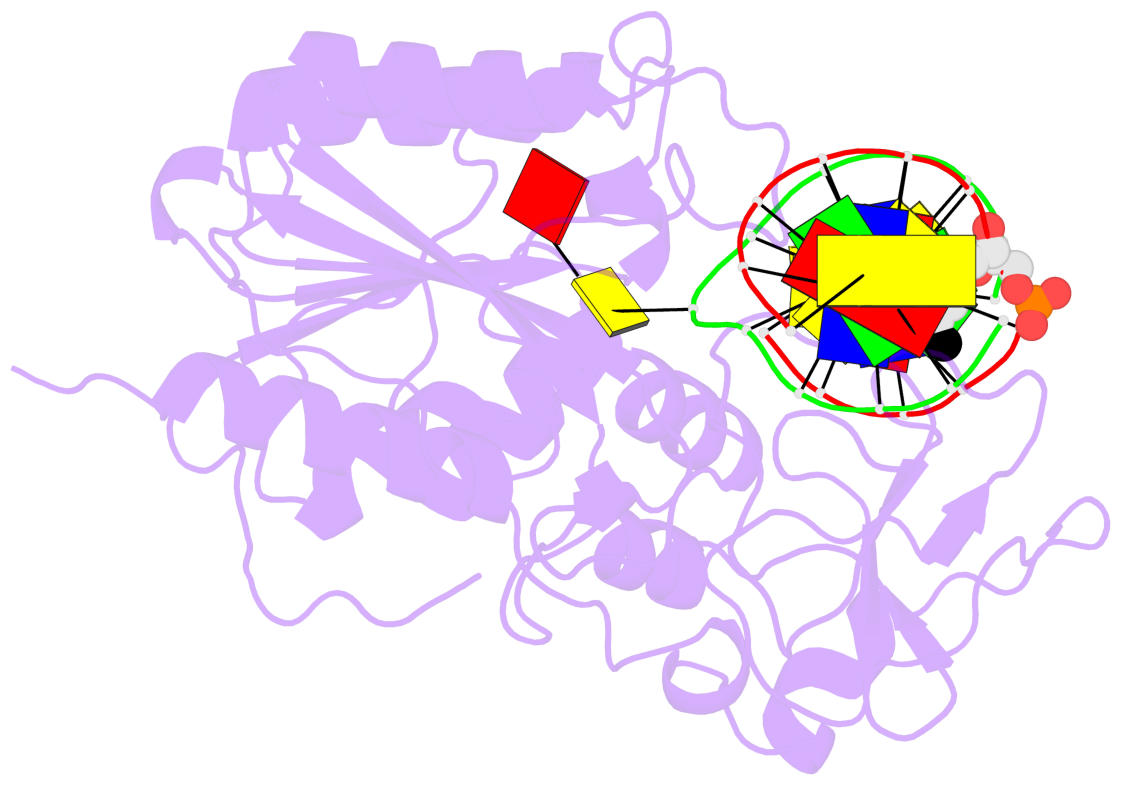
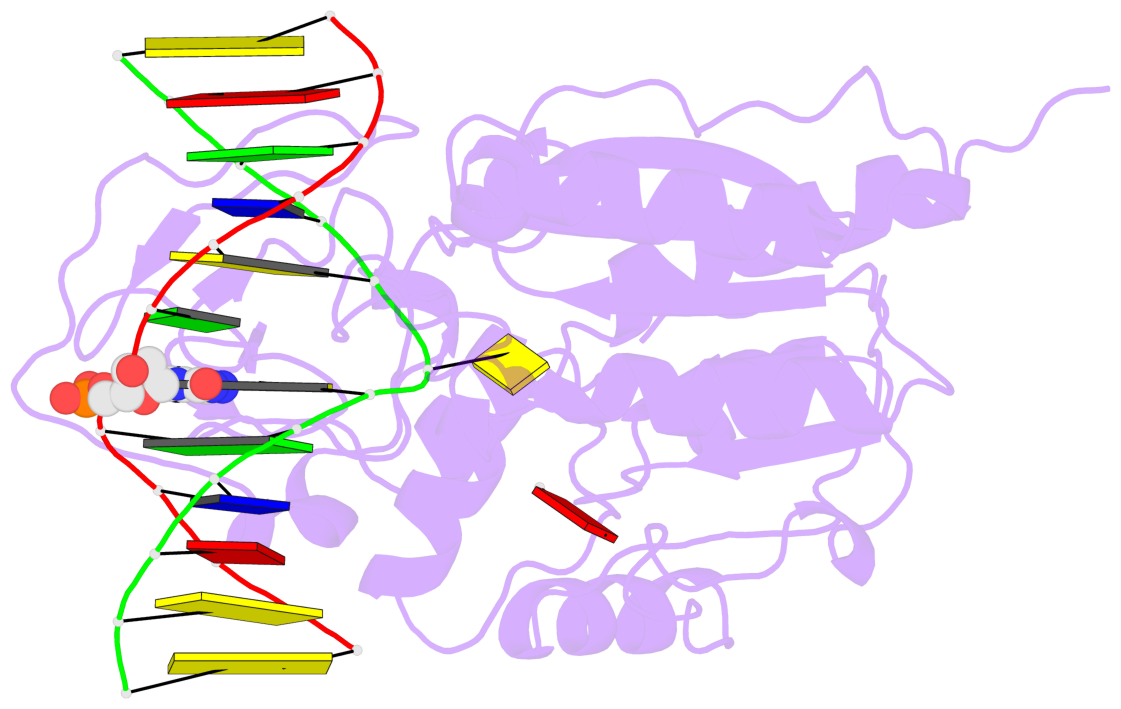
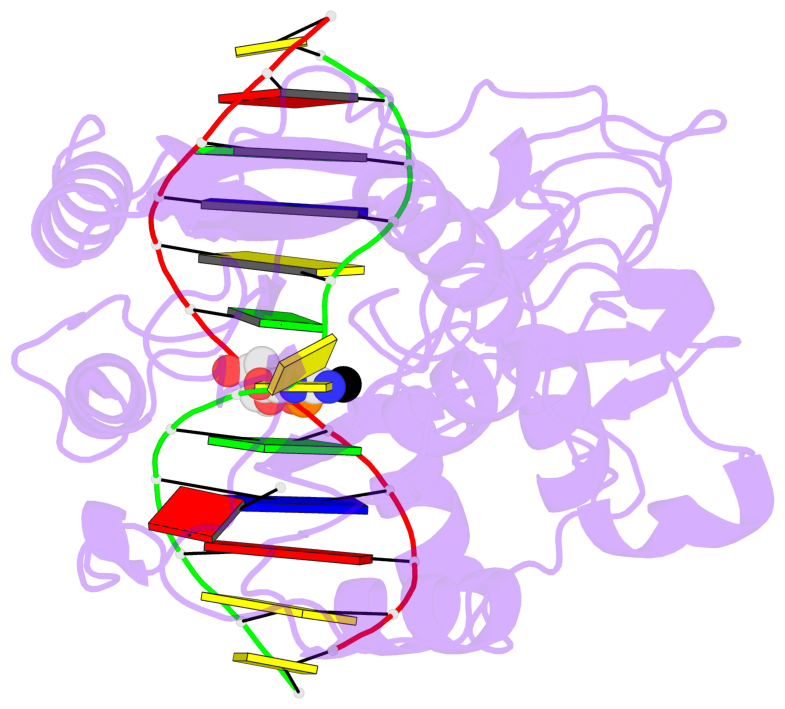
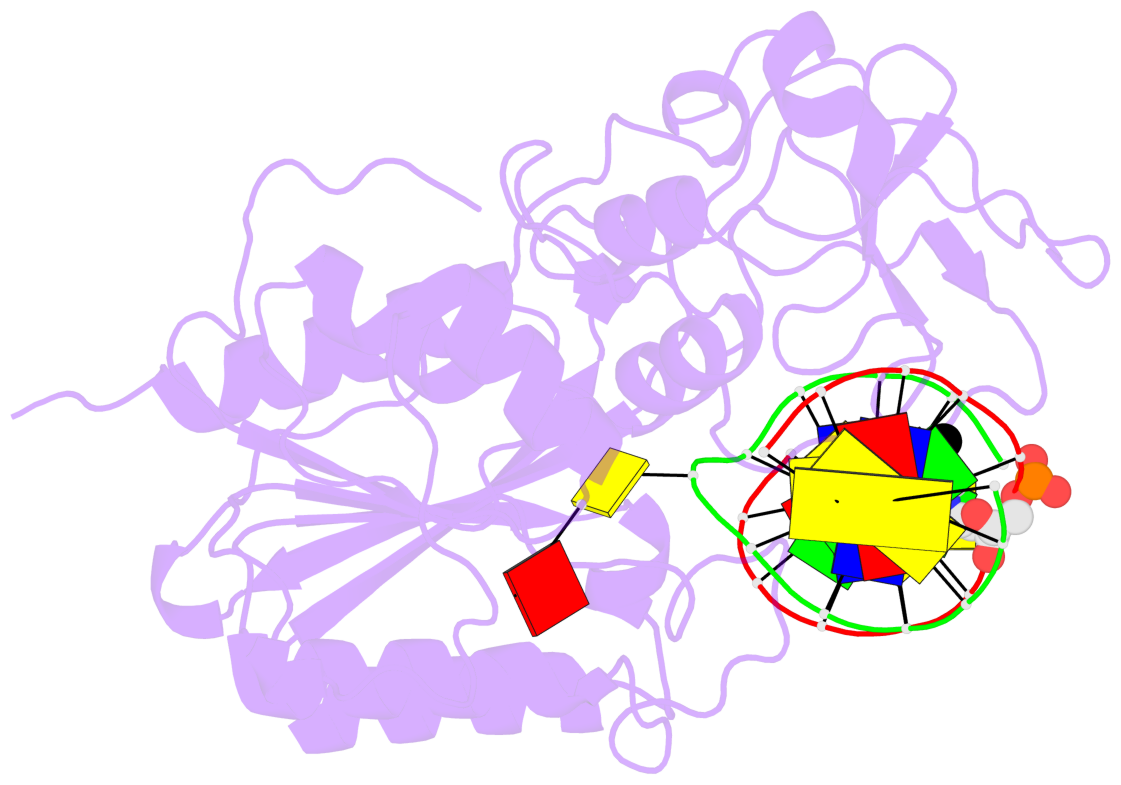
- Ternary structure of hhai methyltransferase with adohcy and DNA containing 4'-thio-2'deoxycytidine at the target
- Reference
- Kumar S, Horton JR, Jones GD, Walker RT, Roberts RJ, Cheng X (1997): "DNA containing 4'-thio-2'-deoxycytidine inhibits methylation by HhaI methyltransferase." Nucleic Acids Res., 25, 2773-2783. doi: 10.1093/nar/25.14.2773.
- Abstract
- 4'-Thio-2'-deoxycytidine was synthesized as a 5'- protected phosphoramidite compatible with solid phase DNA synthesis. When incorporated as the target cytosine (C*) in the GC*GC recognition sequence for the DNA methyltransferase M. HhaI, methyl transfer was strongly inhibited. In contrast, these same oligonucleotides were normal substrates for the cognate restriction endonuclease R. HhaI and its isoschizomer R. Hin P1I. M. HhaI was able to bind both 4'-thio-modified DNA and unmodified DNA to equivalent extents under equilibrium conditions. However, the presence of 4'-thio-2'-deoxycytidine decreased the half-life of the complex by >10-fold. The crystal structure of a ternary complex of M. HhaI, AdoMet and DNA containing 4'-thio-2'-deoxycytidine was solved at 2.05 A resolution with a crystallographic R-factor of 0.186 and R-free of 0.231. The structure is not grossly different from previously solved ternary complexes containing M. HhaI, DNA and AdoHcy. The difference electron density suggests partial methylation at C5 of the flipped target 4'-thio-2'-deoxycytidine. The inhibitory effect of the 4'sulfur atom on enzymatic activity may be traced to perturbation of a step in the methylation reaction after DNA binding but prior to methyl transfer. This inhibitory effect can be partially overcome after a considerably long time in the crystal environment where the packing prevents complex dissociation and the target is accurately positioned within the active site.
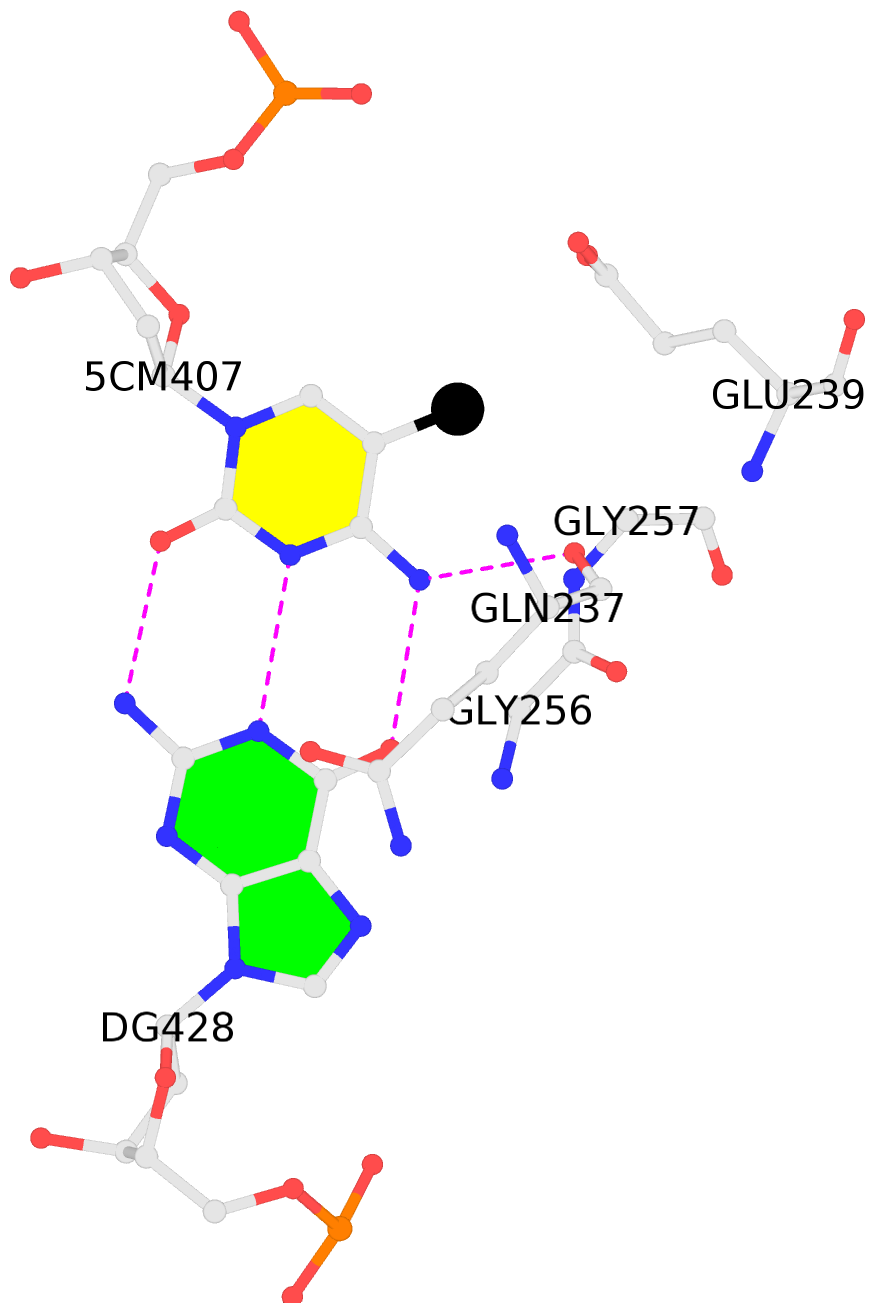
- The contacts include paired nucleotides (mostly a G in Watson-Crick G-C pairing), and
amino-acids within a 4.5-A distance cutoff to base atoms of 5mC.
- The structure is oriented in the base reference frame of 5mC, allowing for easy comparison
and direct superimposition between entries.
- The black sphere (•) denotes the 5-methyl carbon atom in 5mC.
No. 1 C.5CM407: hydrophobic-with-A.GLY256 hydrophobic-with-A.GLY257 is-WC-paired is-in-duplex [+]:GcC/GGC |
 |
|