Summary information and primary citation
- PDB-id
- 7cy6; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase
- Method
- X-ray (2.1 Å)
- Summary
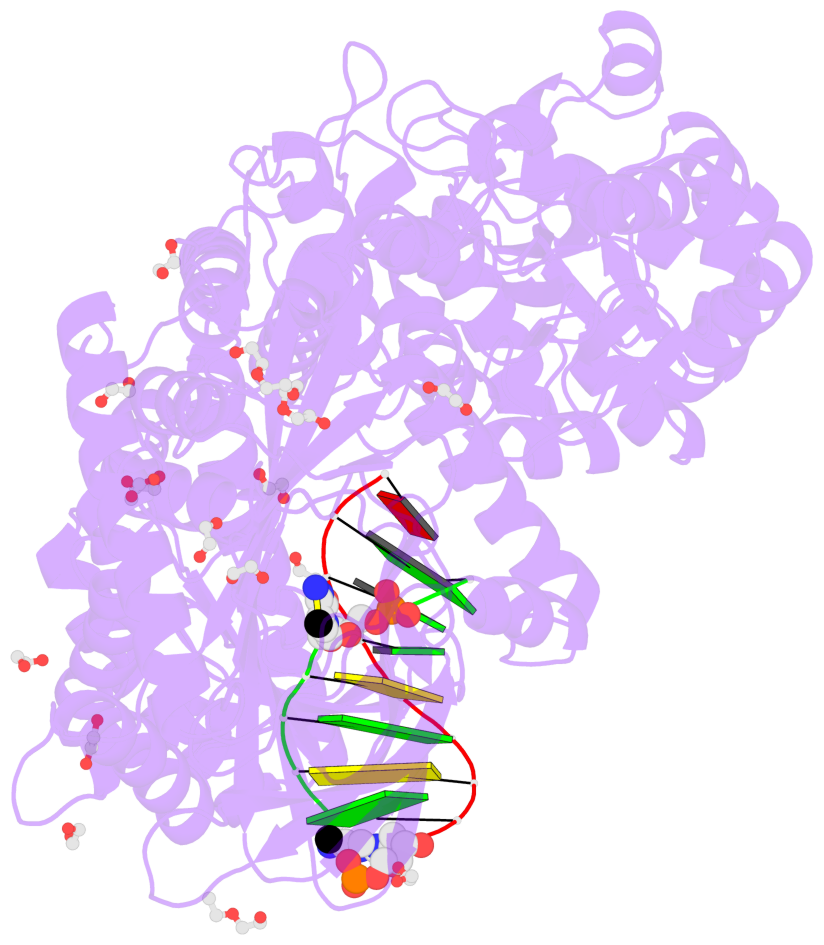
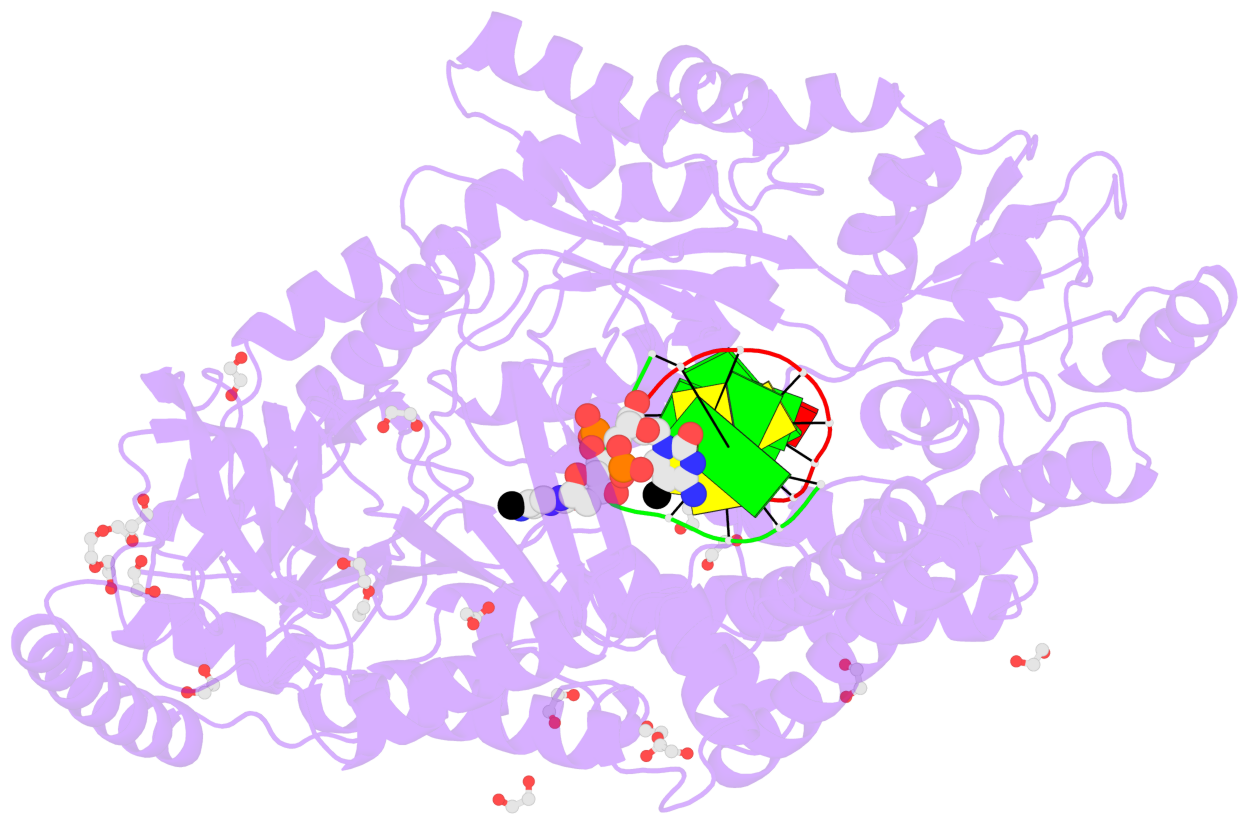
- Crystal structure of cmd1 in complex with 5mc-DNA
- Reference
- Li W, Zhang T, Sun M, Shi Y, Zhang XJ, Xu GL, Ding J (2021): "Molecular mechanism for vitamin C-derived C 5 -glyceryl-methylcytosine DNA modification catalyzed by algal TET homologue CMD1." Nat Commun, 12, 744. doi: 10.1038/s41467-021-21061-2.
- Abstract
- C5-glyceryl-methylcytosine (5gmC) is a novel DNA modification catalyzed by algal TET homologue CMD1 using vitamin C (VC) as co-substrate. Here, we report the structures of CMD1 in apo form and in complexes with VC or/and dsDNA. CMD1 exhibits comparable binding affinities for DNAs of different lengths, structures, and 5mC levels, and displays a moderate substrate preference for 5mCpG-containing DNA. CMD1 adopts the typical DSBH fold of Fe2+/2-OG-dependent dioxygenases. The lactone form of VC binds to the active site and mono-coordinates the Fe2+ in a manner different from 2-OG. The dsDNA binds to a positively charged cleft of CMD1 and the 5mC/C is inserted into the active site and recognized by CMD1 in a similar manner as the TET proteins. The functions of key residues are validated by mutagenesis and activity assay. Our structural and biochemical data together reveal the molecular mechanism for the VC-derived 5gmC DNA modification by CMD1.
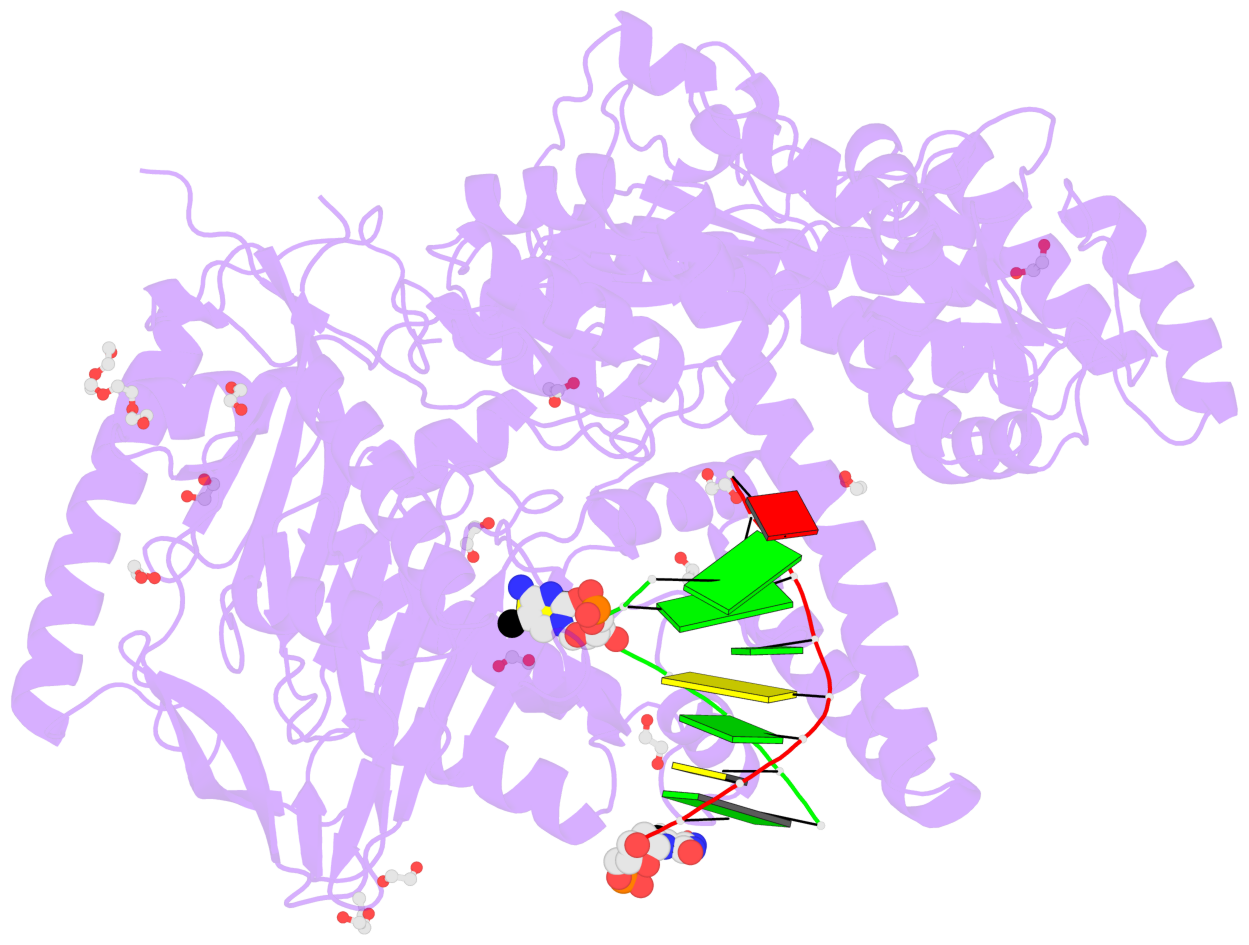
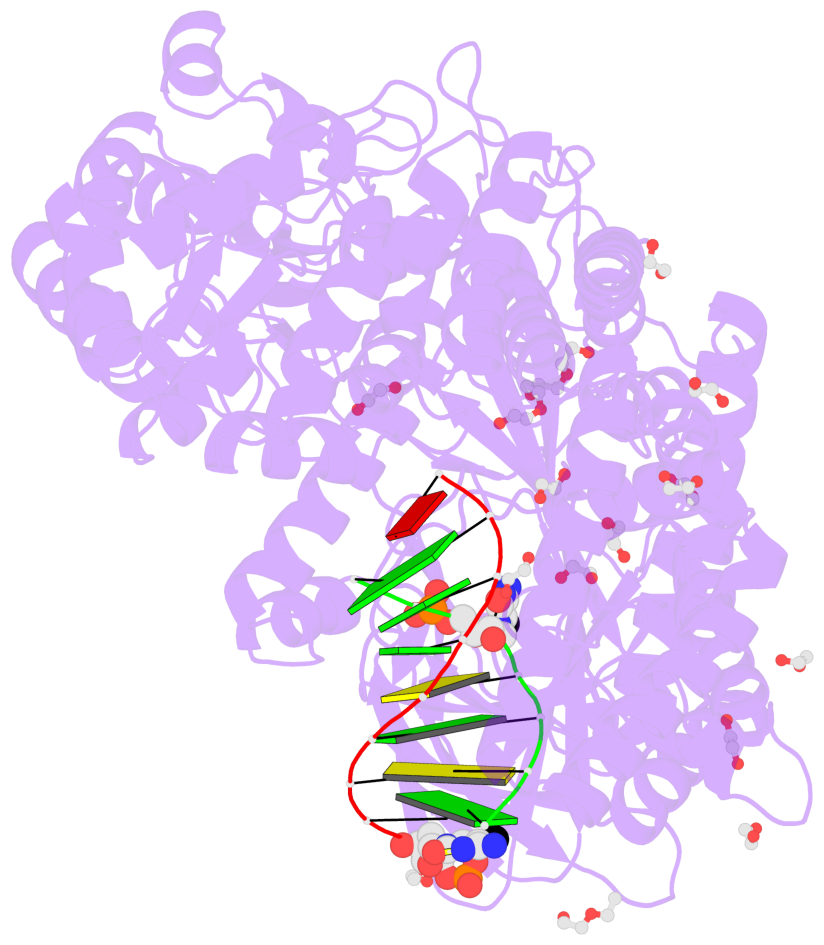
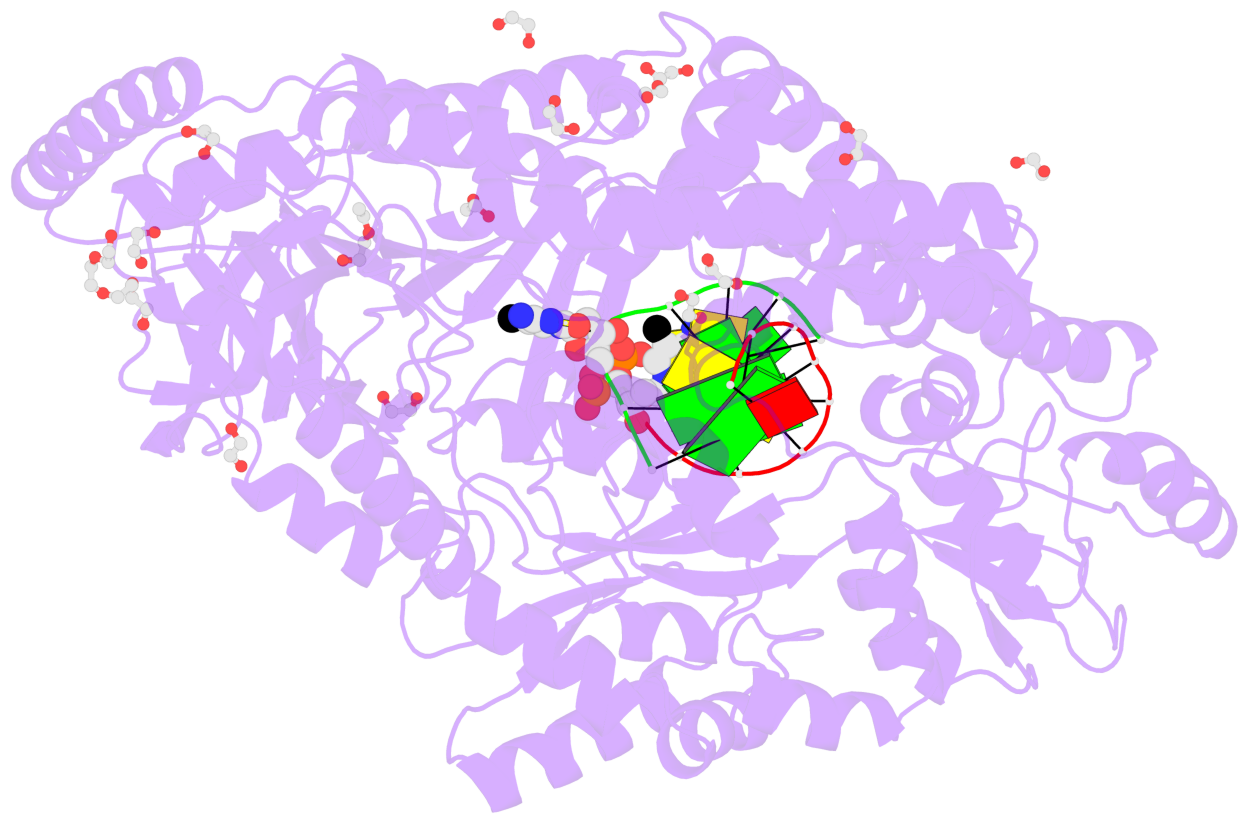
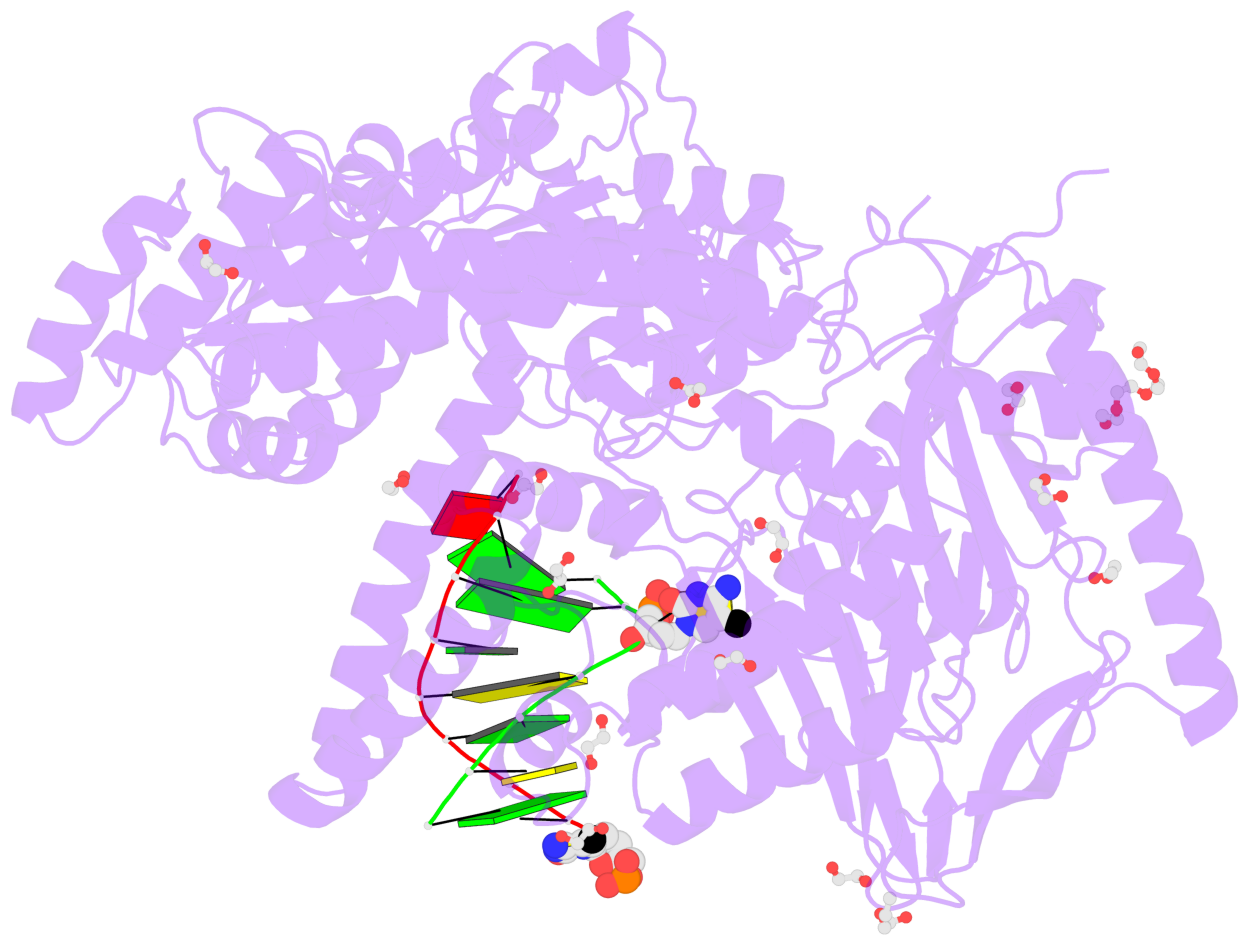
- The contacts include paired nucleotides (mostly a G in Watson-Crick G-C pairing), and
amino-acids within a 4.5-A distance cutoff to base atoms of 5mC.
- The structure is oriented in the base reference frame of 5mC, allowing for easy comparison
and direct superimposition between entries.
- The black sphere (•) denotes the 5-methyl carbon atom in 5mC.
No. 1 D.5CM3: stacking-with-A.ARG244 stacking-with-A.PHE416 not-WC-paired not-in-duplex |
 |
|