Summary information and primary citation
- PDB-id
- 7fef; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein-DNA
- Method
- X-ray (2.39 Å)
- Summary
- Crystal structure of atmbd6 with DNA
- Reference
- Wu Z, Chen S, Zhou M, Jia L, Li Z, Zhang X, Min J, Liu K (2022): "Family-wide Characterization of Methylated DNA Binding Ability of Arabidopsis MBDs." J.Mol.Biol., 434, 167404. doi: 10.1016/j.jmb.2021.167404.
- Abstract
- 13 MBD-containing genes (AtMBD1-13) have been identified in Arabidopsis thaliana so far, however, their DNA binding ability is still controversial. Here, we systematically measured the DNA binding affinities of these MBDs by ITC and EMSA binding assays, except for those of pseudogenes AtMBD3 and AtMBD13, and found that only AtMBD6 and AtMBD7 function as methylated DNA readers. We also found that the MBD of AtMBD5 exhibits very weak binding to methylated DNA compared to that of AtMBD6. To further investigate the structural basis of AtMBDs in binding to methylated DNA, we determined the complex structure of the AtMBD6 MBD with a 12mer mCG DNA and the apo structure of the AtMBD5 MBD. Structural analysis coupled with mutagenesis studies indicated that, in addition to the conserved arginine fingers contributing to the DNA binding specificity, the residues located in the loop1 and α1 are also essential for the methylated DNA binding of these MBDs in Arabidopsis thaliana, which explains why AtMBD5 MBD and the other AtMBDs display very weak or no binding to methylated DNA. Thus, our study here systematically demonstrates the DNA binding ability of the MBDs in Arabidopsis thaliana, which also provides a general guideline in understanding the DNA binding ability of the MBDs in other plants as a whole.
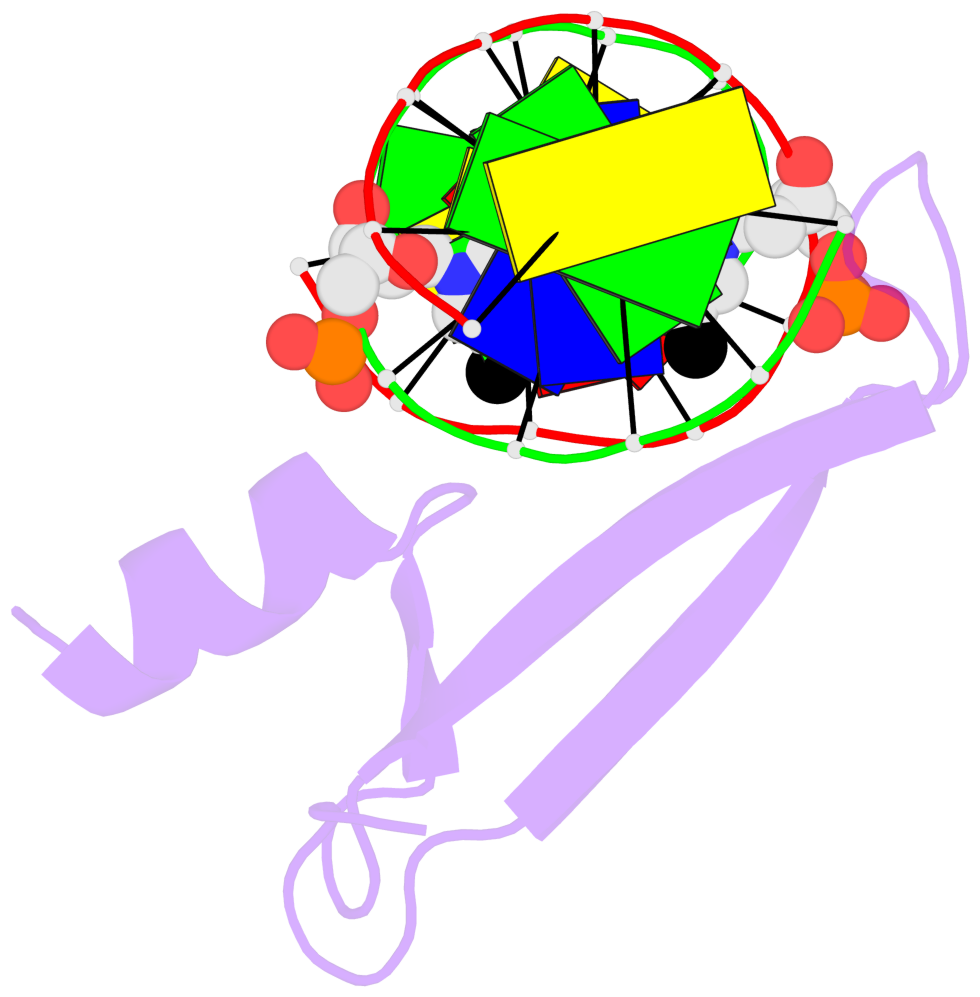
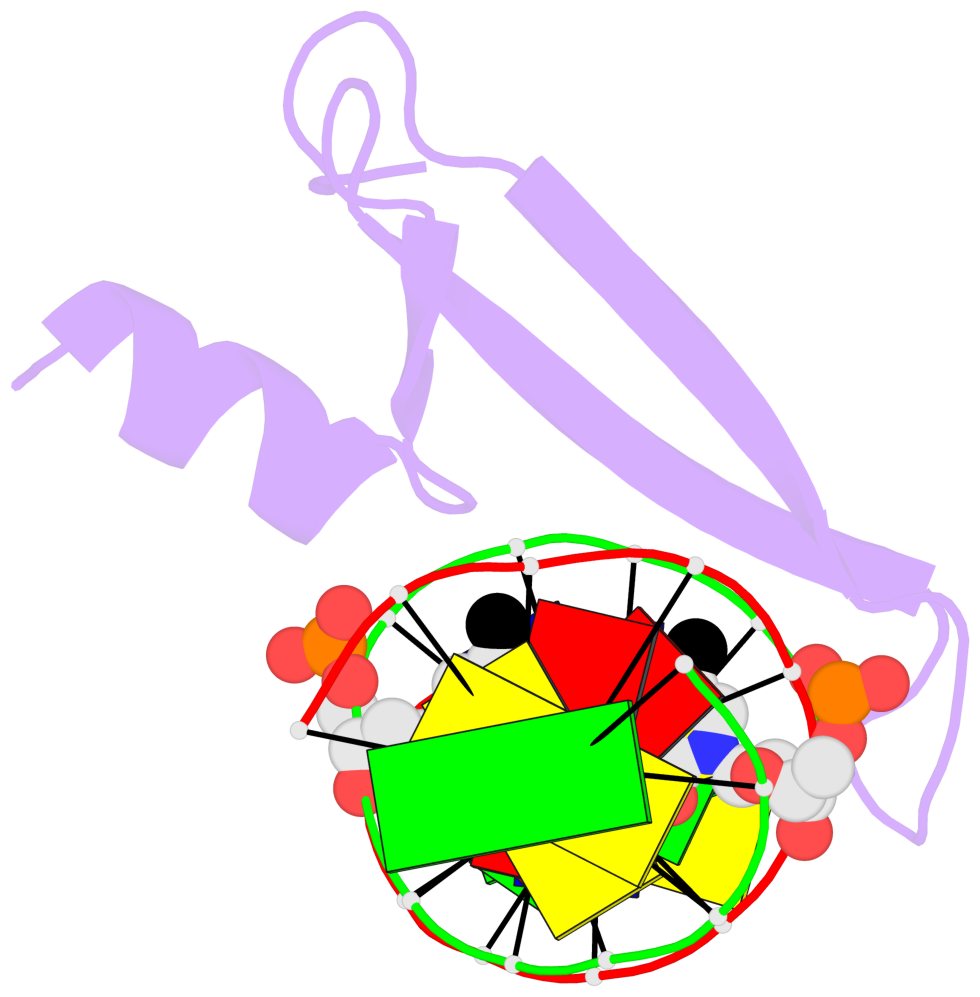
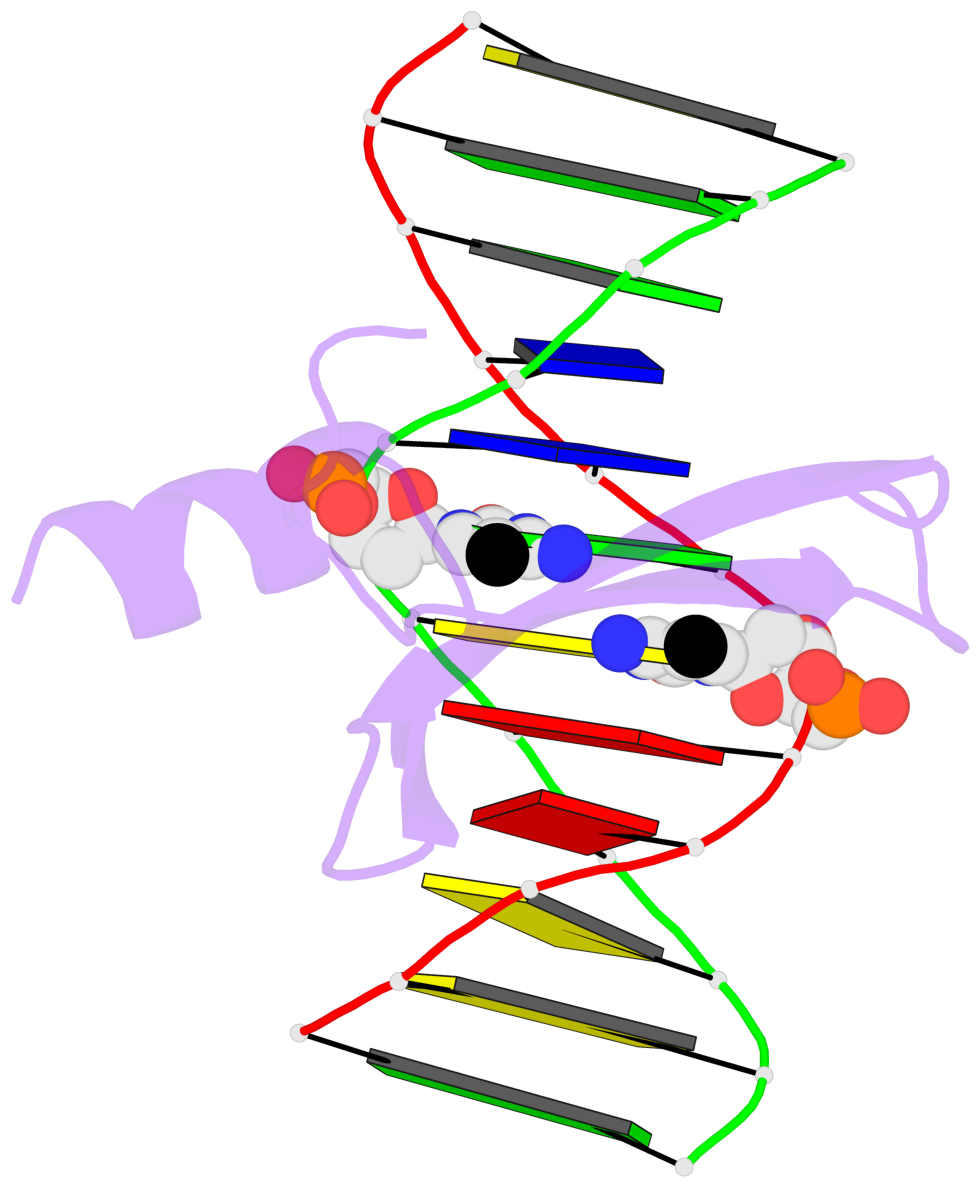
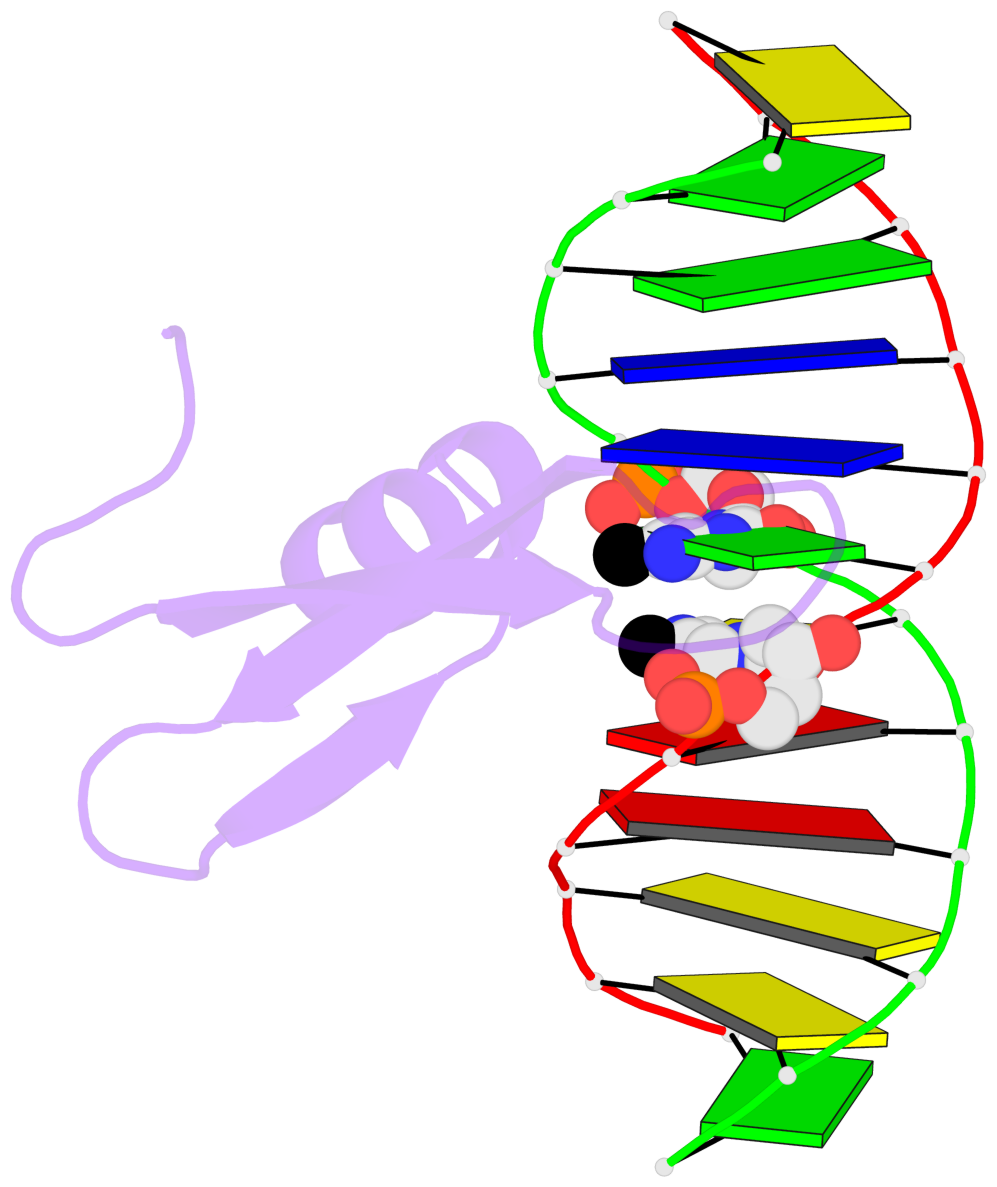
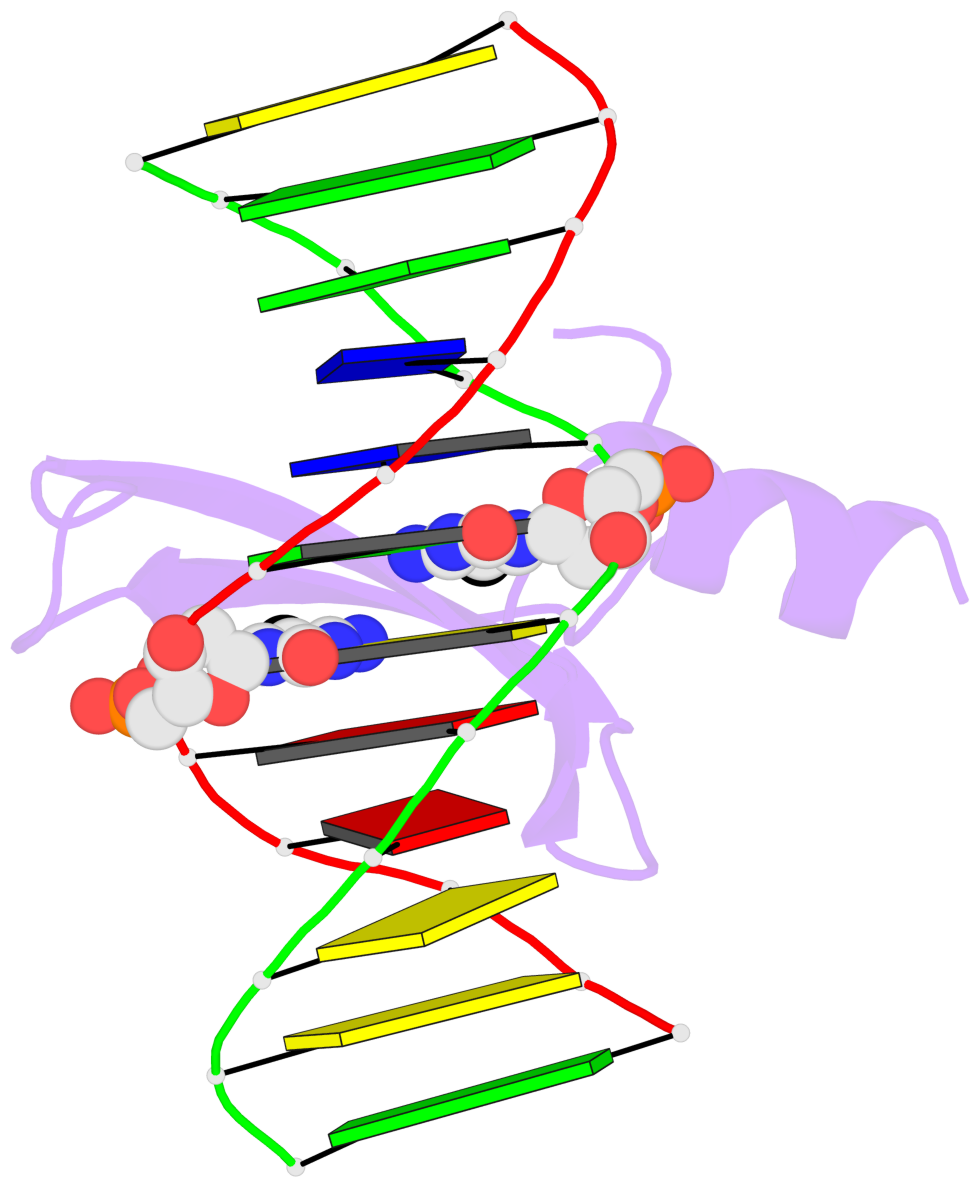
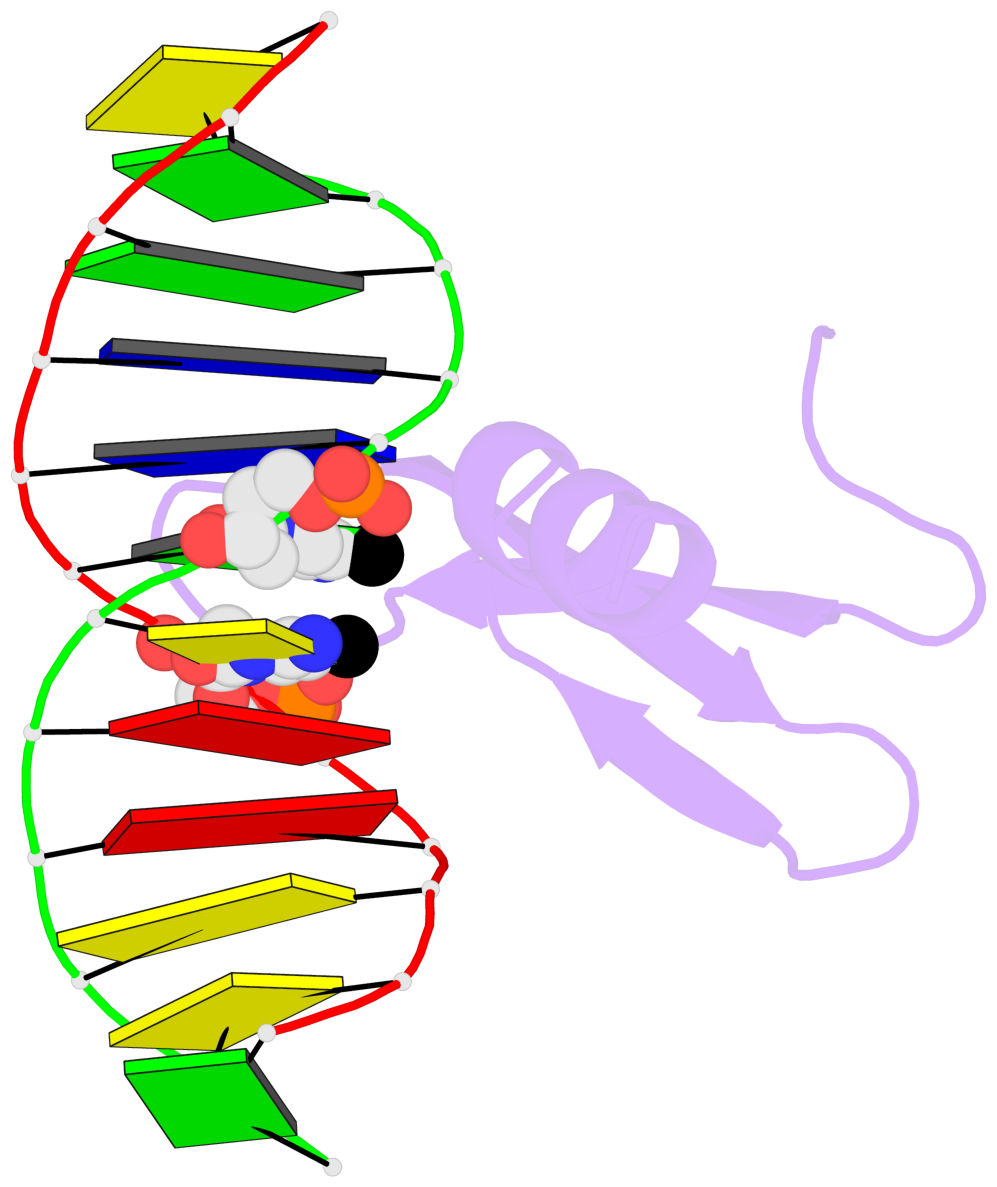
- The contacts include paired nucleotides (mostly a G in Watson-Crick G-C pairing), and
amino-acids within a 4.5-A distance cutoff to base atoms of 5mC.
- The structure is oriented in the base reference frame of 5mC, allowing for easy comparison
and direct superimposition between entries.
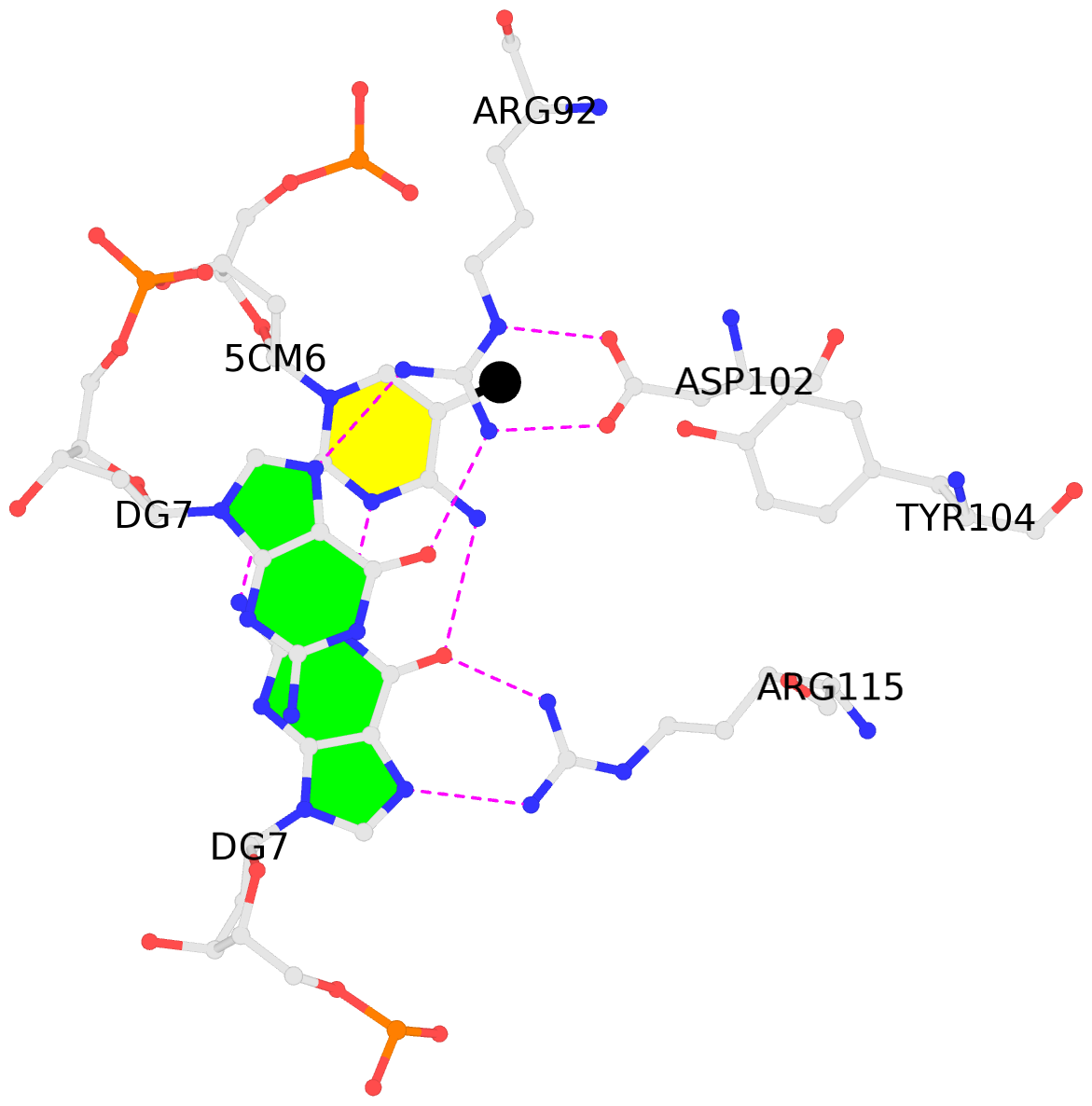
- The black sphere (•) denotes the 5-methyl carbon atom in 5mC.
No. 1 E.5CM6: stacking-with-A.ARG92 is-WC-paired is-in-duplex [+]:AcG/cGT |
 |
|
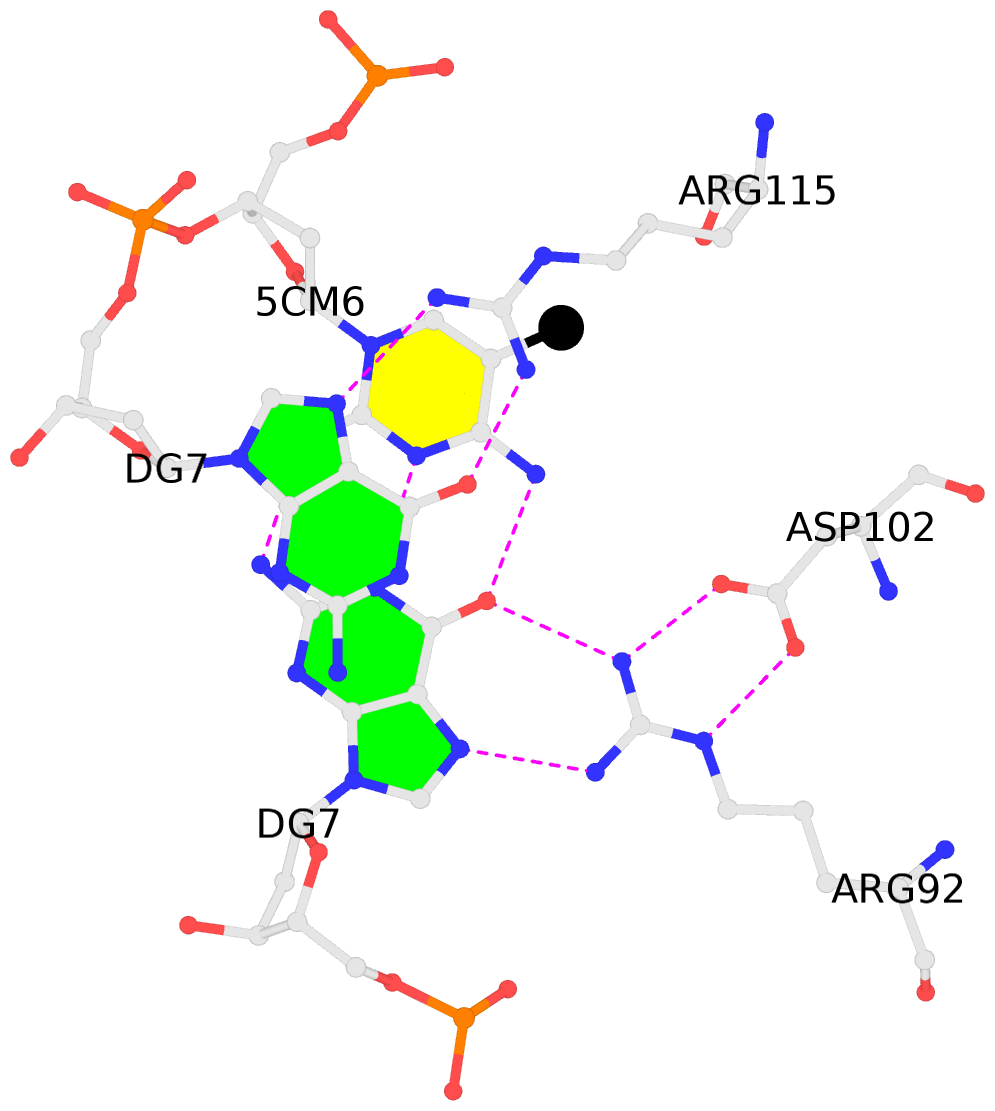
No. 2 F.5CM6: stacking-with-A.ARG115 is-WC-paired is-in-duplex [-]:cGT/AcG |
 |
|