Summary information and primary citation
- PDB-id
- 1exy; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein-RNA
- Method
- NMR
- Summary
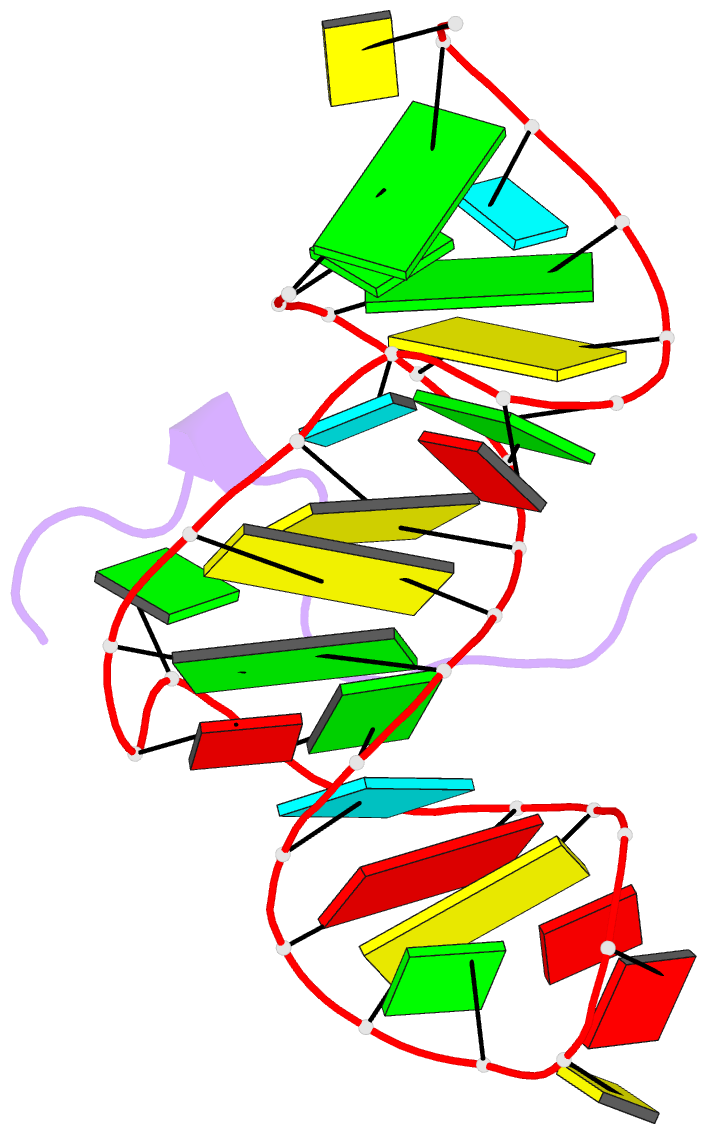
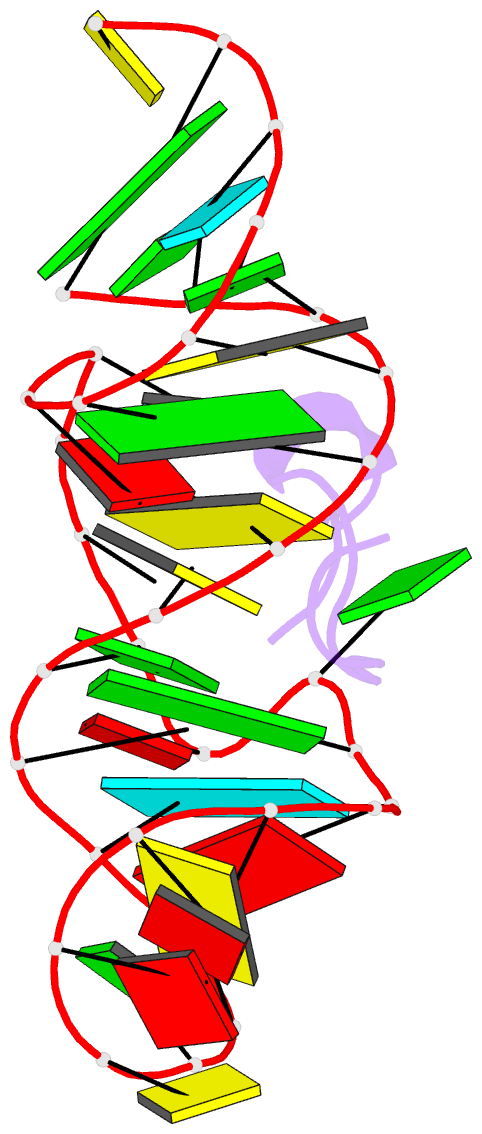
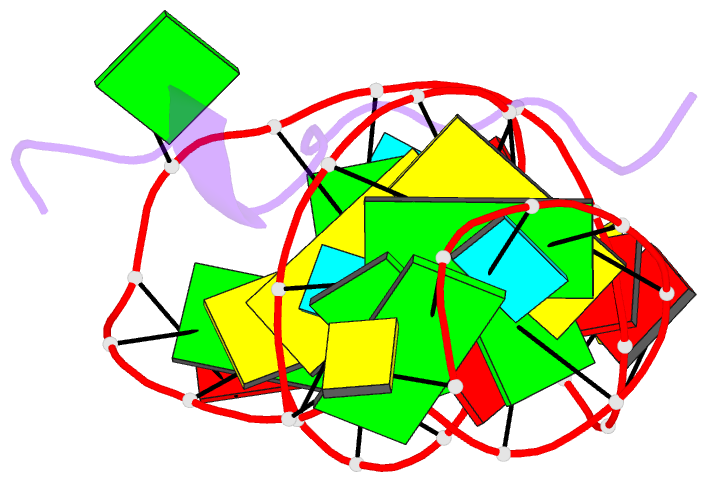
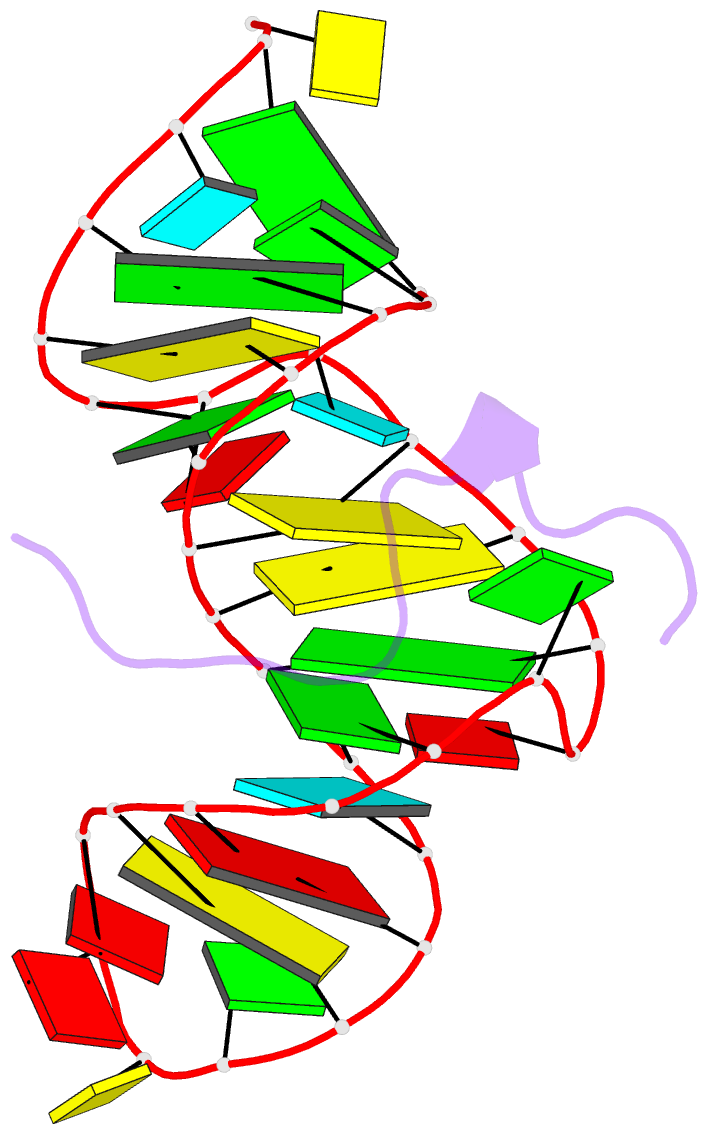
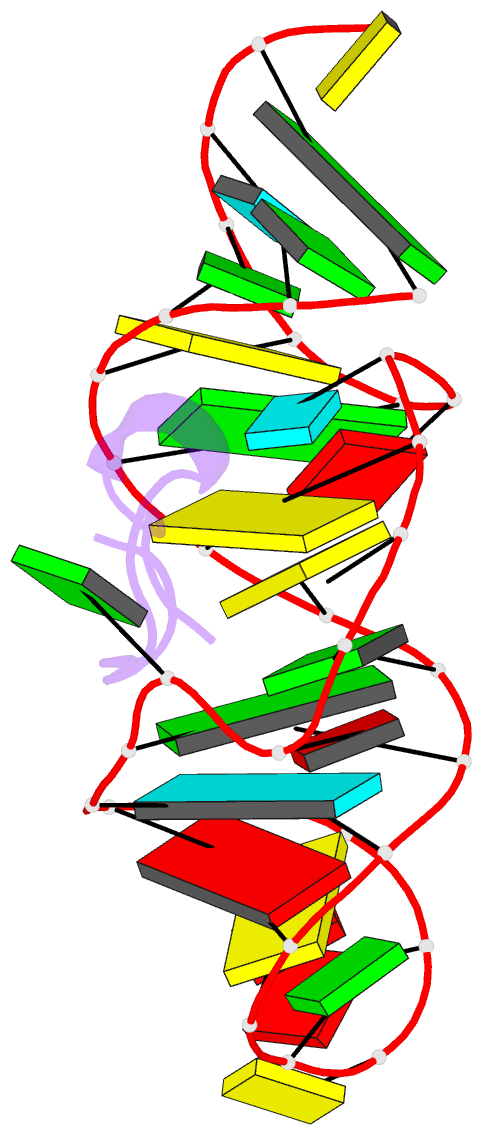
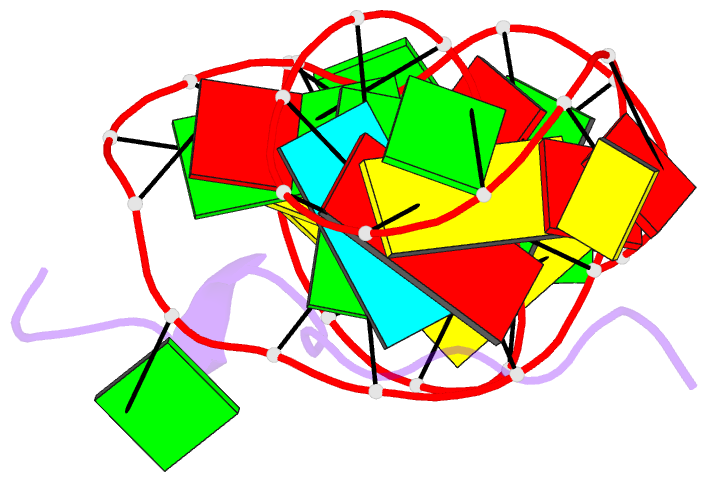
- Solution structure of htlv-1 peptide bound to its RNA aptamer target
- Reference
- Jiang F, Gorin A, Hu W, Majumdar A, Baskerville S, Xu W, Ellington A, Patel DJ (1999): "Anchoring an extended HTLV-1 Rex peptide within an RNA major groove containing junctional base triples." Structure Fold.Des., 7, 1461-1472. doi: 10.1016/S0969-2126(00)88337-9.
- Abstract
- Background: The Rex protein of the human T cell leukemia virus type 1 (HTLV-1) belongs to a family of proteins that use arginine-rich motifs (ARMs) to recognize their RNA targets. Previously, an in vitro selected RNA aptamer sequence was identified that mediates mRNA transport in vivo when placed in the primary binding site on stem-loop IID of the Rex response element. We present the solution structure of the HTLV-1 arginine-rich Rex peptide bound to its RNA aptamer target determined by multidimensional heteronuclear NMR spectroscopy.
Results: The Rex peptide in a predominantly extended conformation threads through a channel formed by the shallow and widened RNA major groove and a looped out guanine. The RNA aptamer contains three stems separated by a pair of two-base bulges, and adopts an unanticipated fold in which both junctional sites are anchored through base triple formation. Binding specificity is associated with intermolecular hydrogen bonding between guanidinium groups of three non-adjacent arginines and the guanine base edges of three adjacent G.C pairs.
Conclusions: The extended S-shaped conformation of the Rex peptide, together with previous demonstrations of a beta-hairpin conformation for the bovine immunodeficiency virus (BIV) Tat peptide and an alpha-helical conformation for the human immunodeficiency virus (HIV) Rev peptide in complex with their respective RNA targets, expands our understanding of the strategies employed by ARMs for adaptive recognition and highlights the importance of RNA tertiary structure in accommodating minimalist elements of protein secondary structure.