Summary information and primary citation
- PDB-id
- 1lfu; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription
- Method
- NMR
- Summary
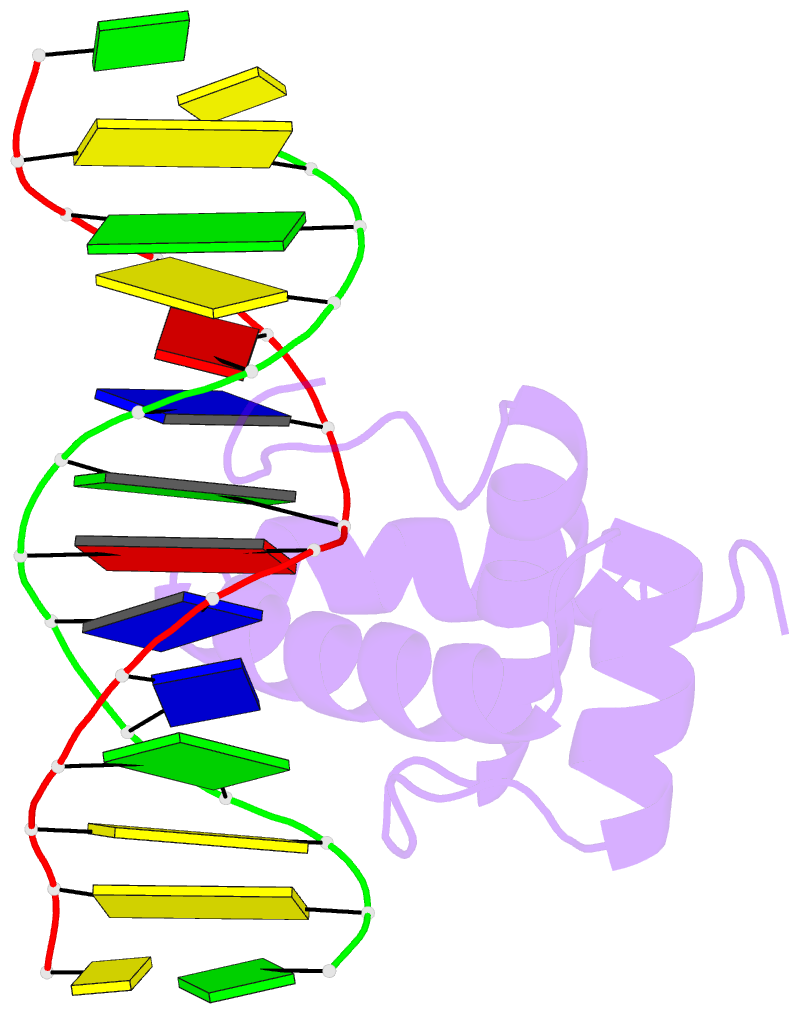
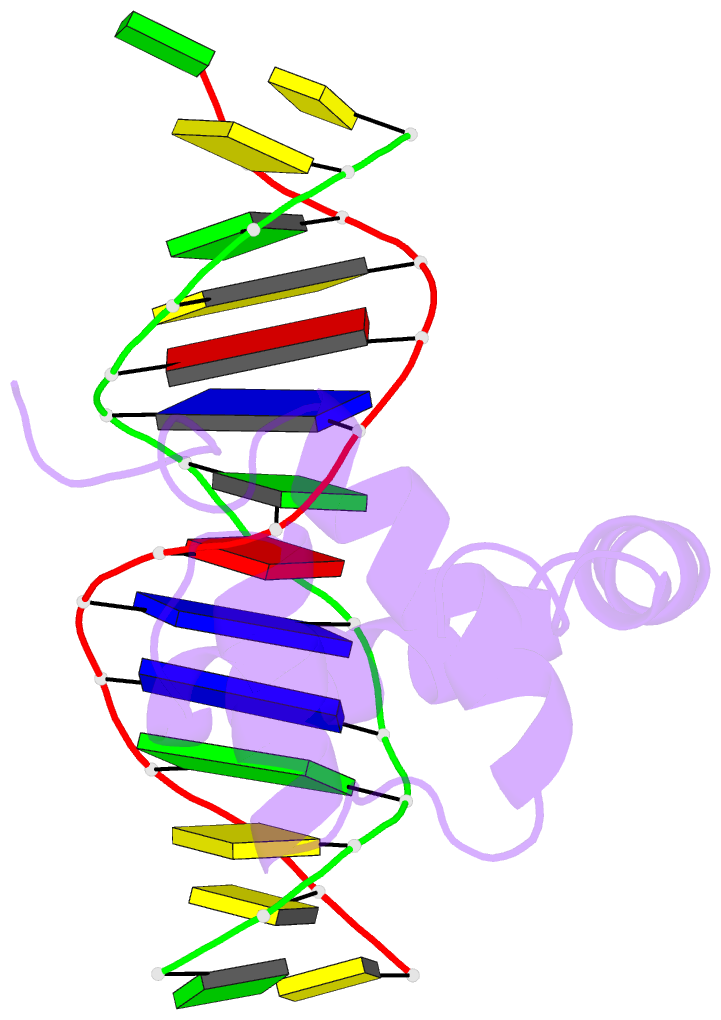
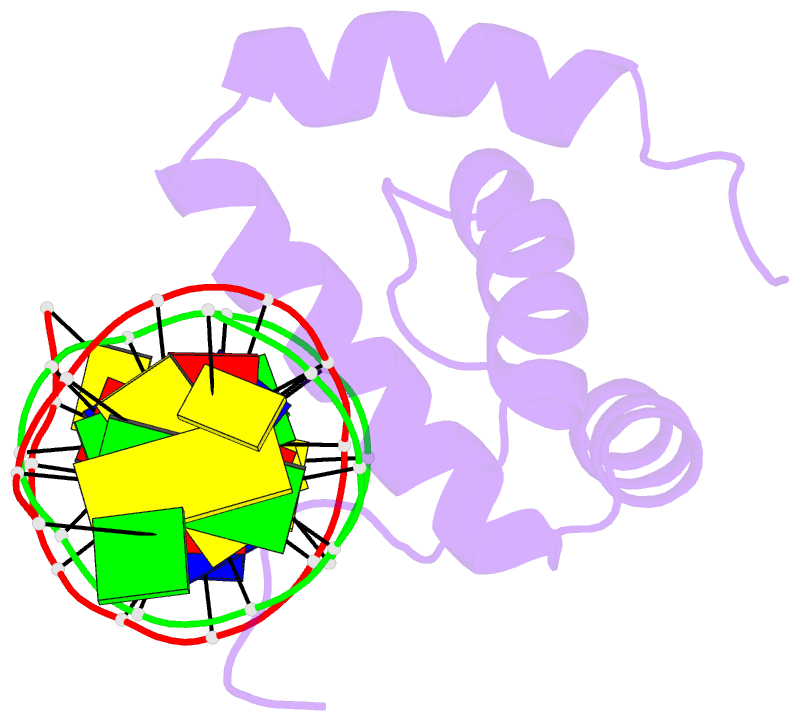
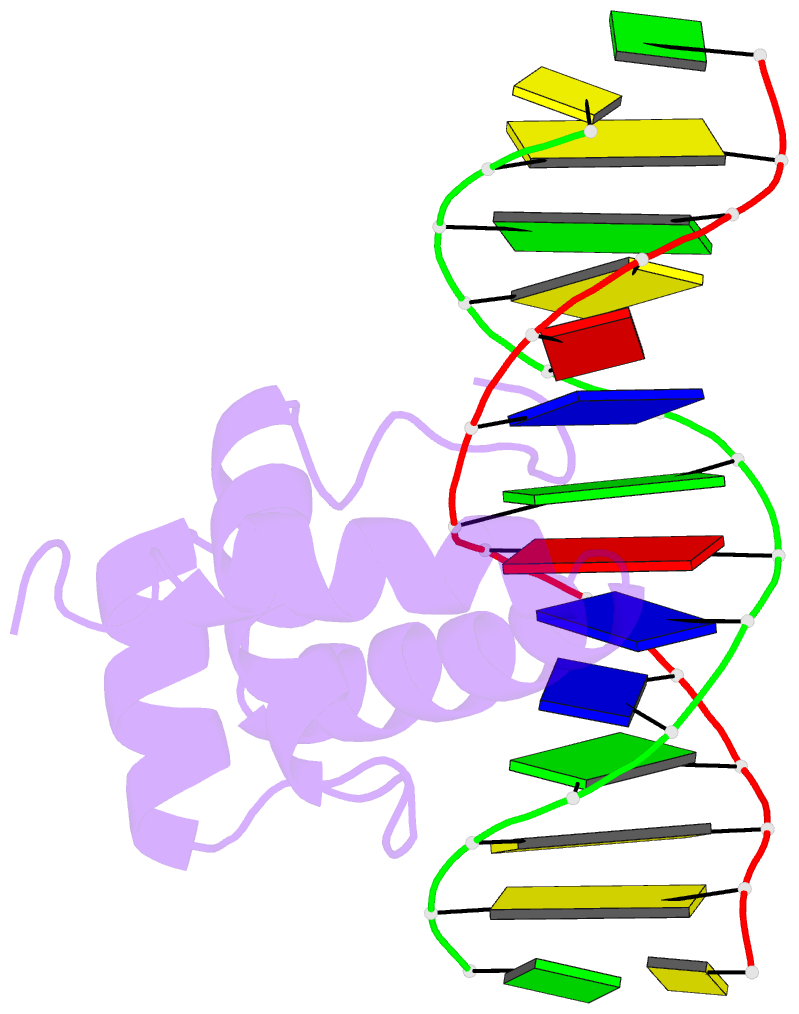
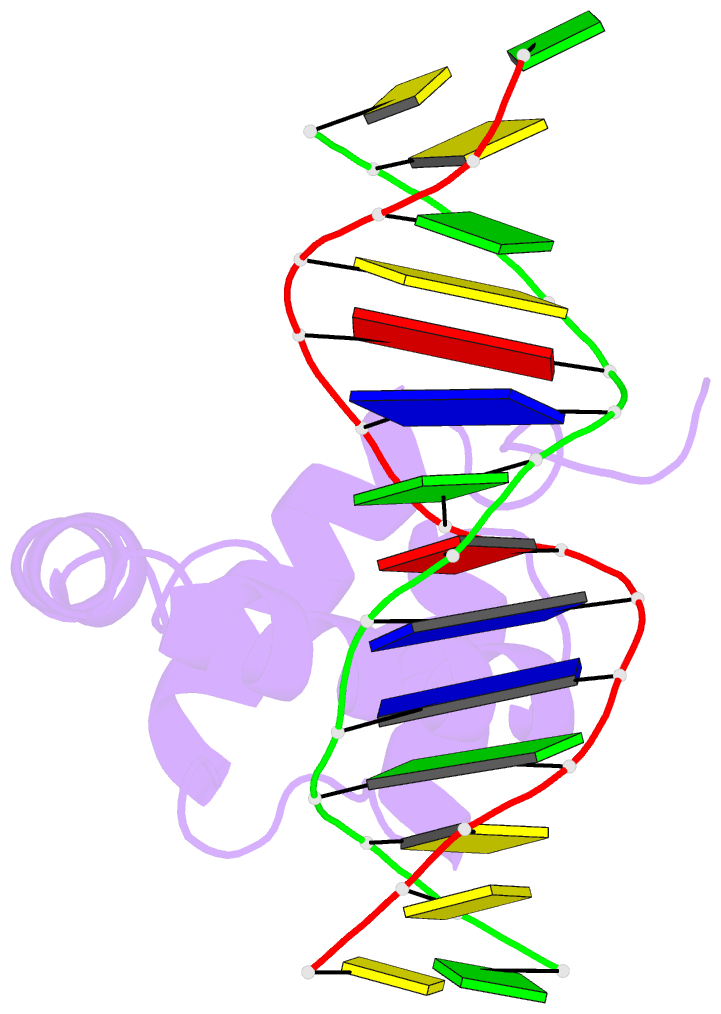
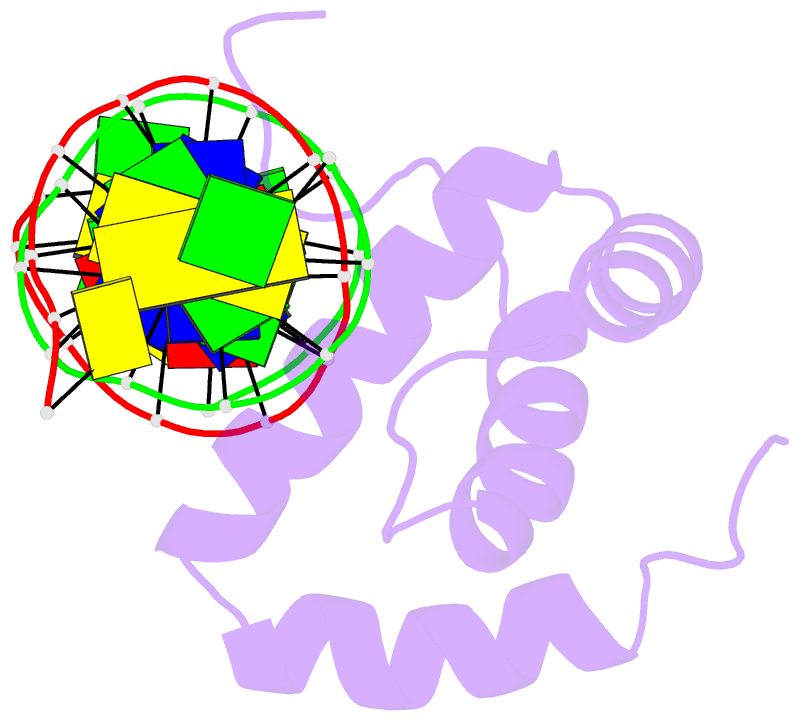
- NMR solution structure of the extended pbx homeodomain bound to DNA
- Reference
- Sprules T, Green N, Featherstone M, Gehring K (2003): "Lock and Key Binding of the HOX YPWM Peptide to the PBX Homeodomain." J.Biol.Chem., 278, 1053-1058. doi: 10.1074/jbc.M207504200.
- Abstract
- HOX homeodomain proteins bind short core DNA sequences to control very specific developmental processes. DNA binding affinity and sequence selectivity are increased by the formation of cooperative complexes with the PBX homeodomain protein. A conserved YPWM motif in the HOX protein is necessary for cooperative binding with PBX. We have determined the structure of a PBX homeodomain bound to a 14-mer DNA duplex. A relaxation-optimized procedure was developed to measure DNA residual dipolar couplings at natural abundance in the 20-kDa binary complex. When the PBX homeodomain binds to DNA, a fourth alpha-helix is formed in the homeodomain. This helix rigidifies the DNA recognition helix of PBX and forms a hydrophobic binding site for the HOX YPWM peptide. The HOX peptide itself shows some structure in solution and suggests that the interaction between PBX and HOX is an example of "lock and key" binding. The NMR structure explains the requirement of DNA for the PBX-HOX interaction and the increased affinity of DNA binding.