Summary information and primary citation
- PDB-id
- 1nk7; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-DNA
- Method
- X-ray (1.9 Å)
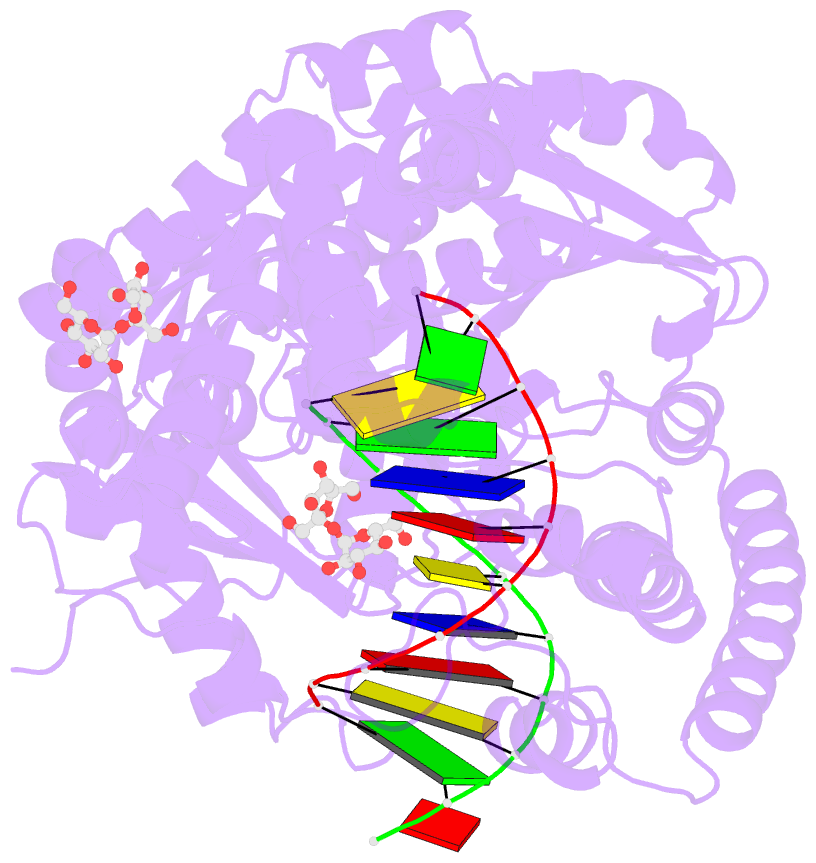
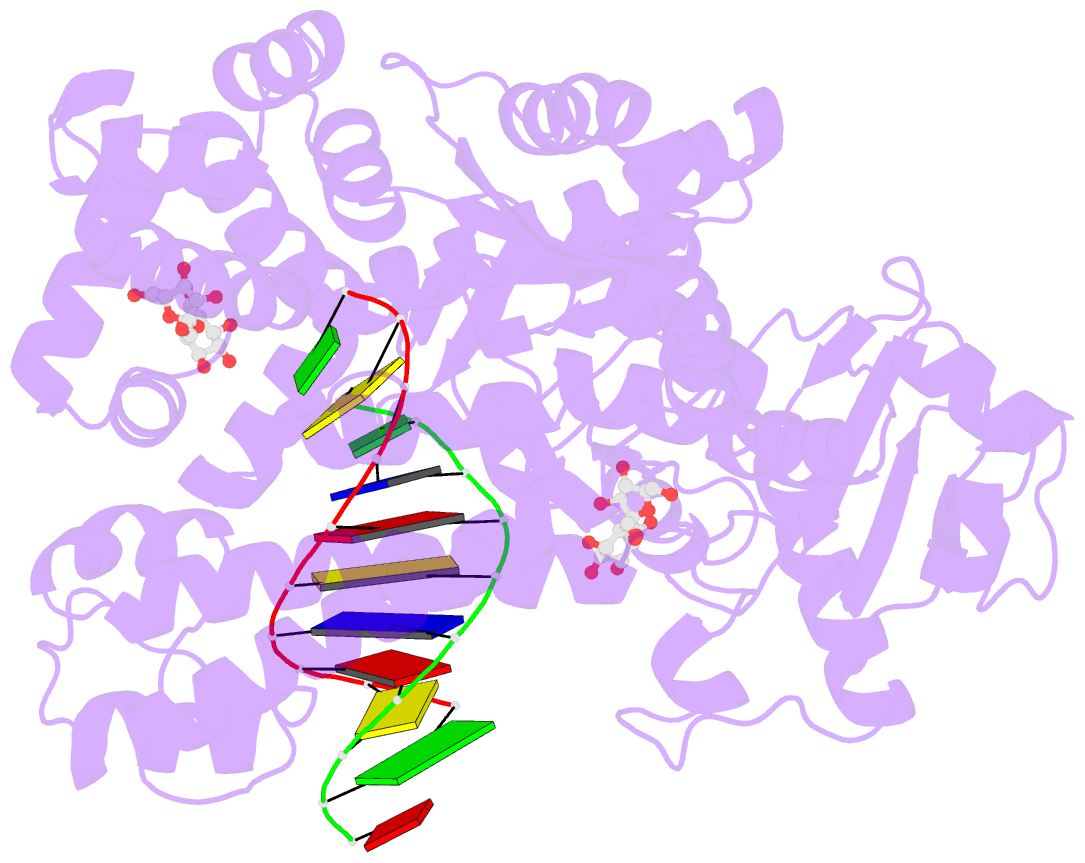
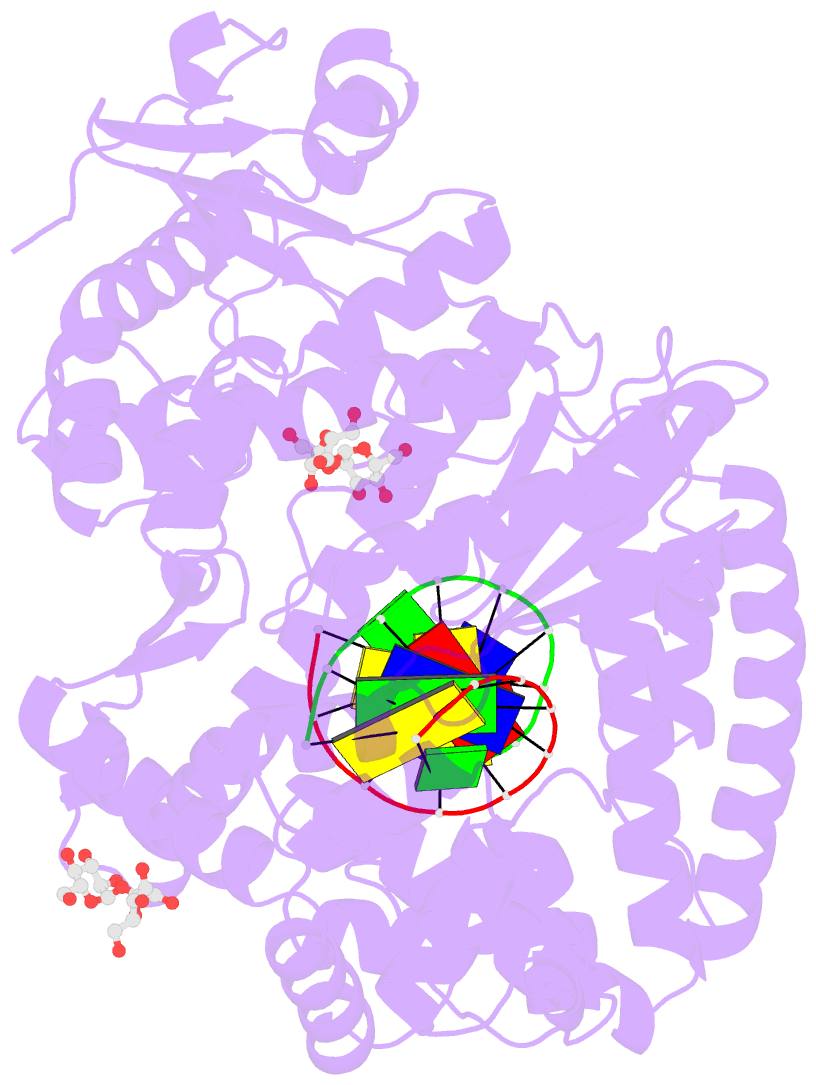
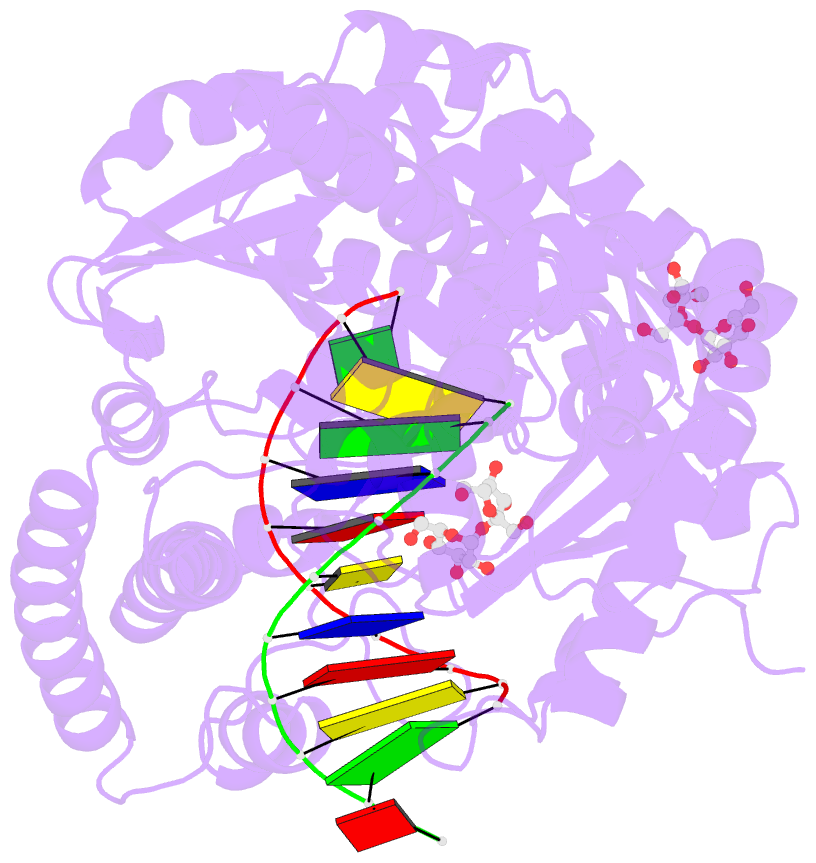
- Summary
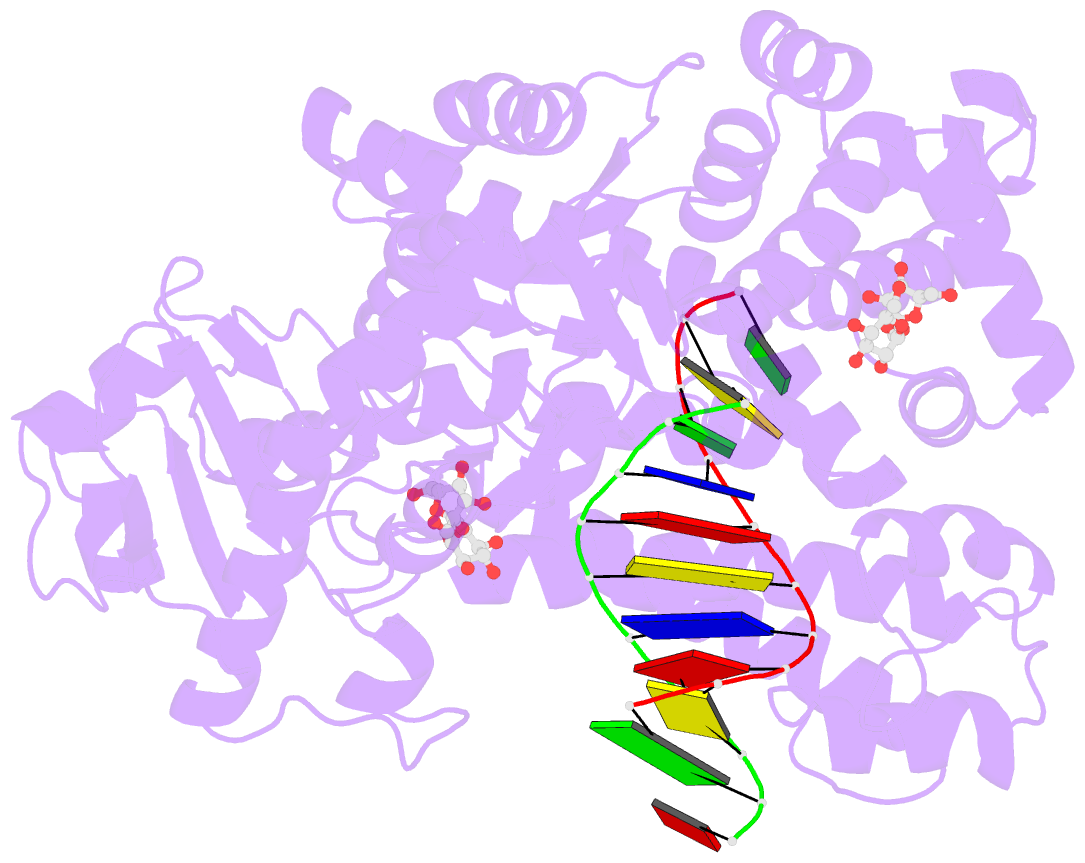
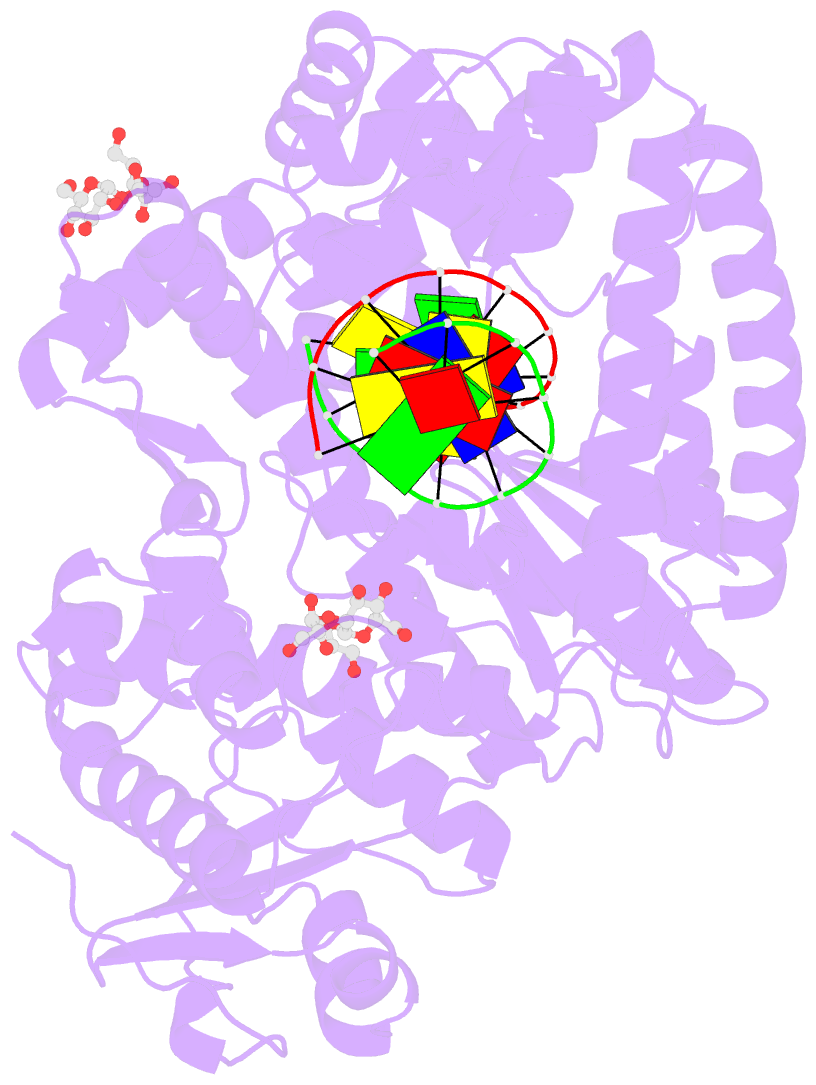
- Guanine-adenine mismatch at the polymerase active site
- Reference
- Johnson SJ, Beese LS (2004): "Structures of mismatch replication errors observed in a DNA polymerase." Cell(Cambridge,Mass.), 116, 803-816. doi: 10.1016/S0092-8674(04)00252-1.
- Abstract
- Accurate DNA replication is essential for genomic stability. One mechanism by which high-fidelity DNA polymerases maintain replication accuracy involves stalling of the polymerase in response to covalent incorporation of mismatched base pairs, thereby favoring subsequent mismatch excision. Some polymerases retain a "short-term memory" of replication errors, responding to mismatches up to four base pairs in from the primer terminus. Here we a present a structural characterization of all 12 possible mismatches captured at the growing primer terminus in the active site of a polymerase. Our observations suggest four mechanisms that lead to mismatch-induced stalling of the polymerase. Furthermore, we have observed the effects of extending a mismatch up to six base pairs from the primer terminus and find that long-range distortions in the DNA transmit the presence of the mismatch back to the enzyme active site, suggesting the structural basis for the short-term memory of replication errors.