Summary information and primary citation
- PDB-id
- 1wvl; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- structural protein-DNA
- Method
- X-ray (2.6 Å)
- Summary
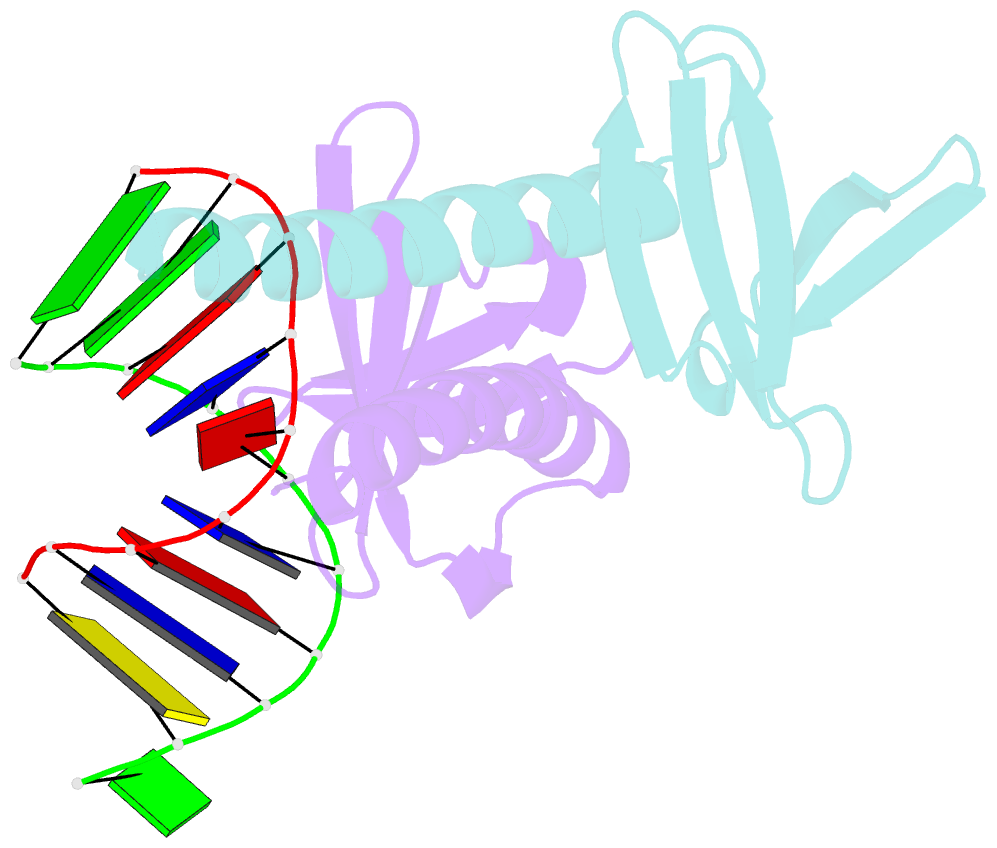
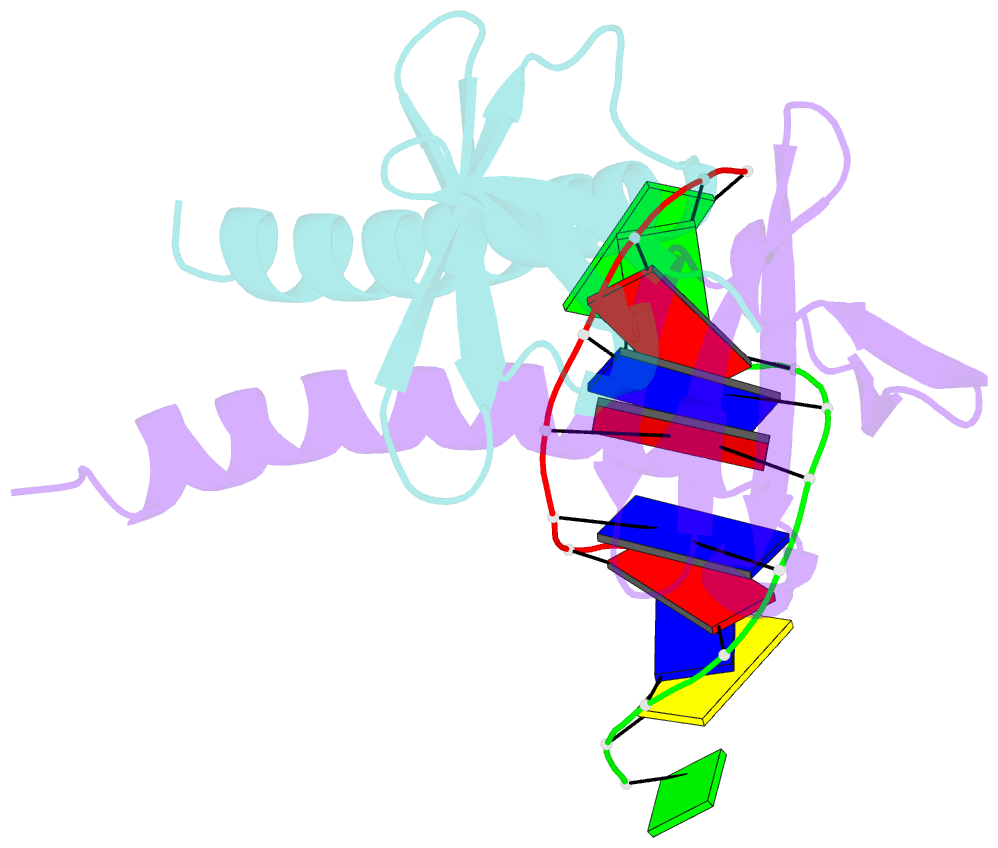
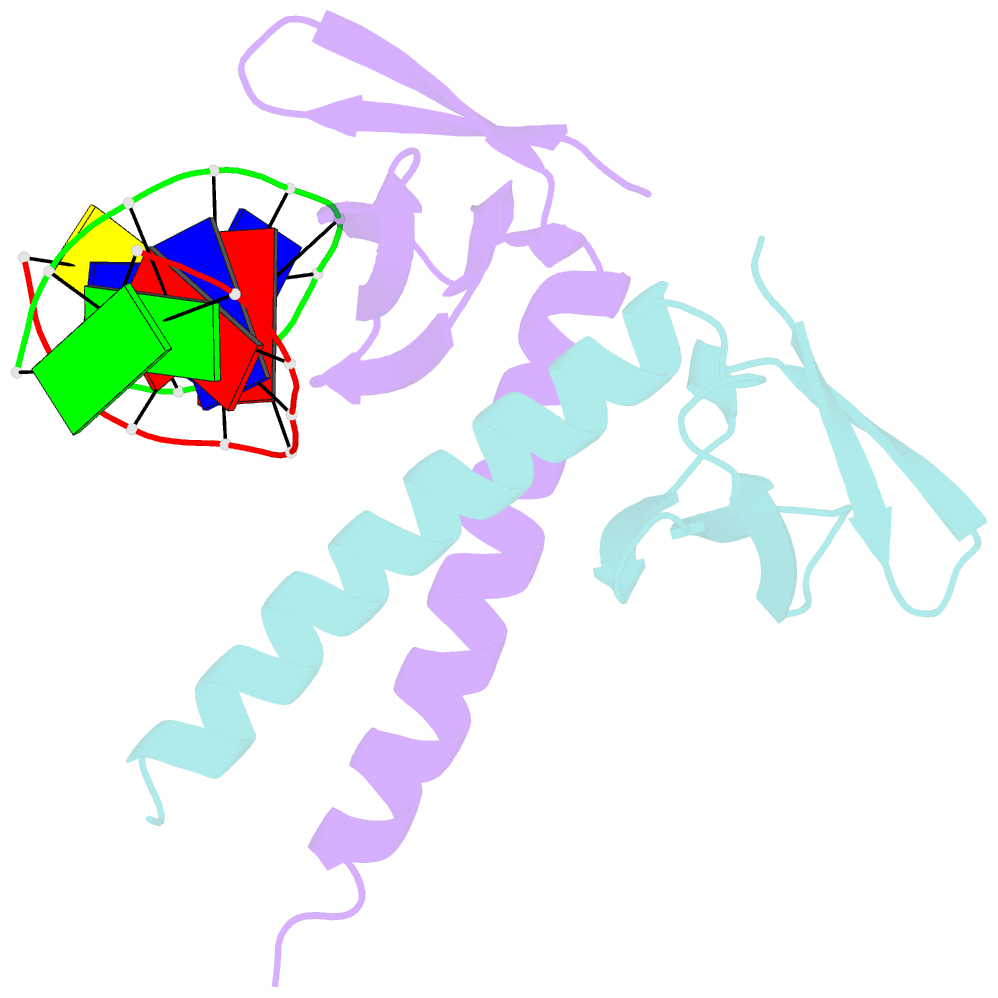
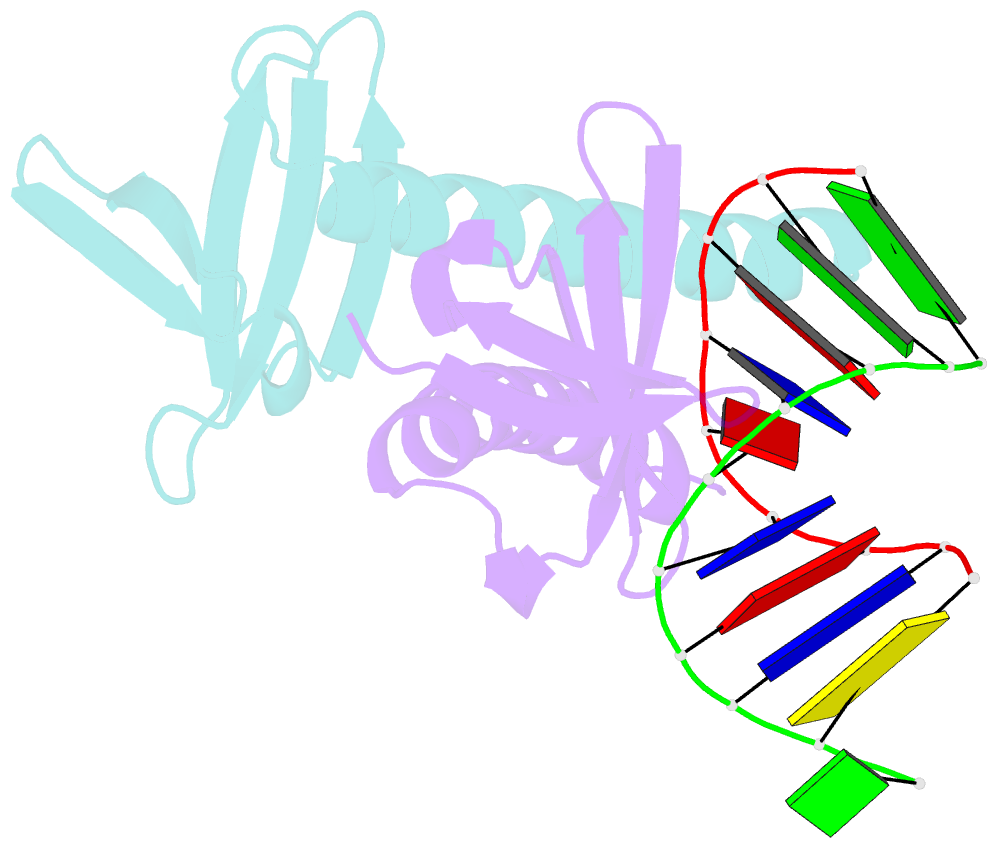
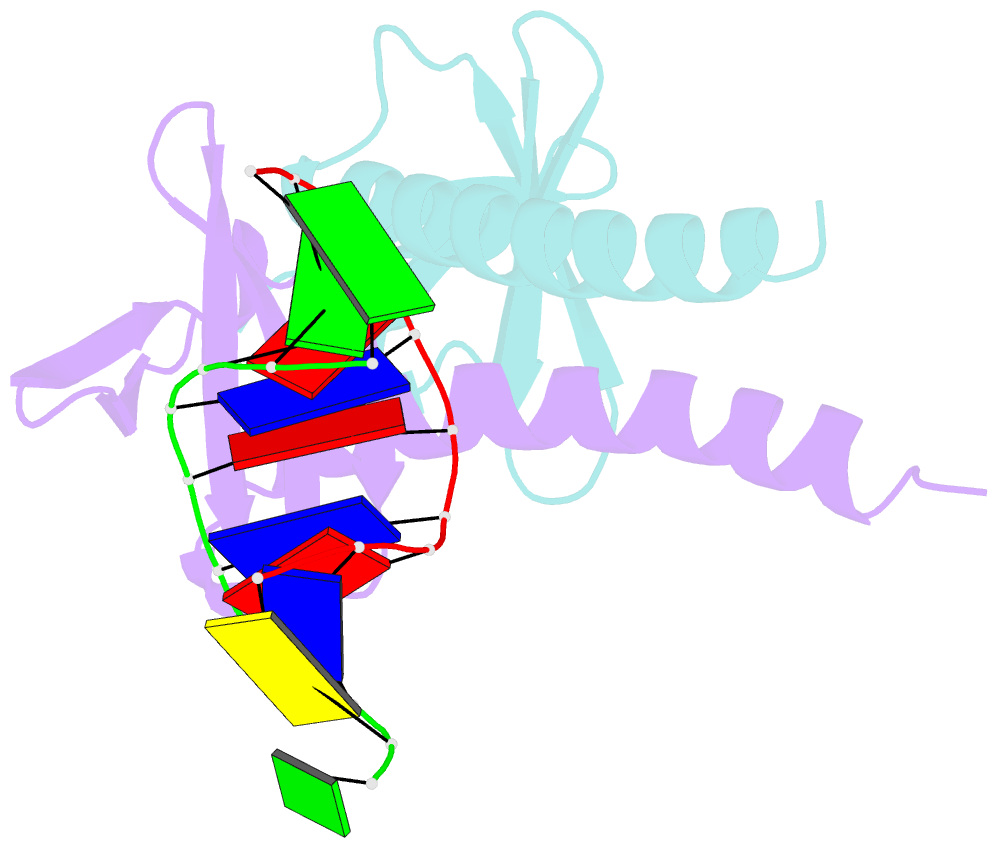
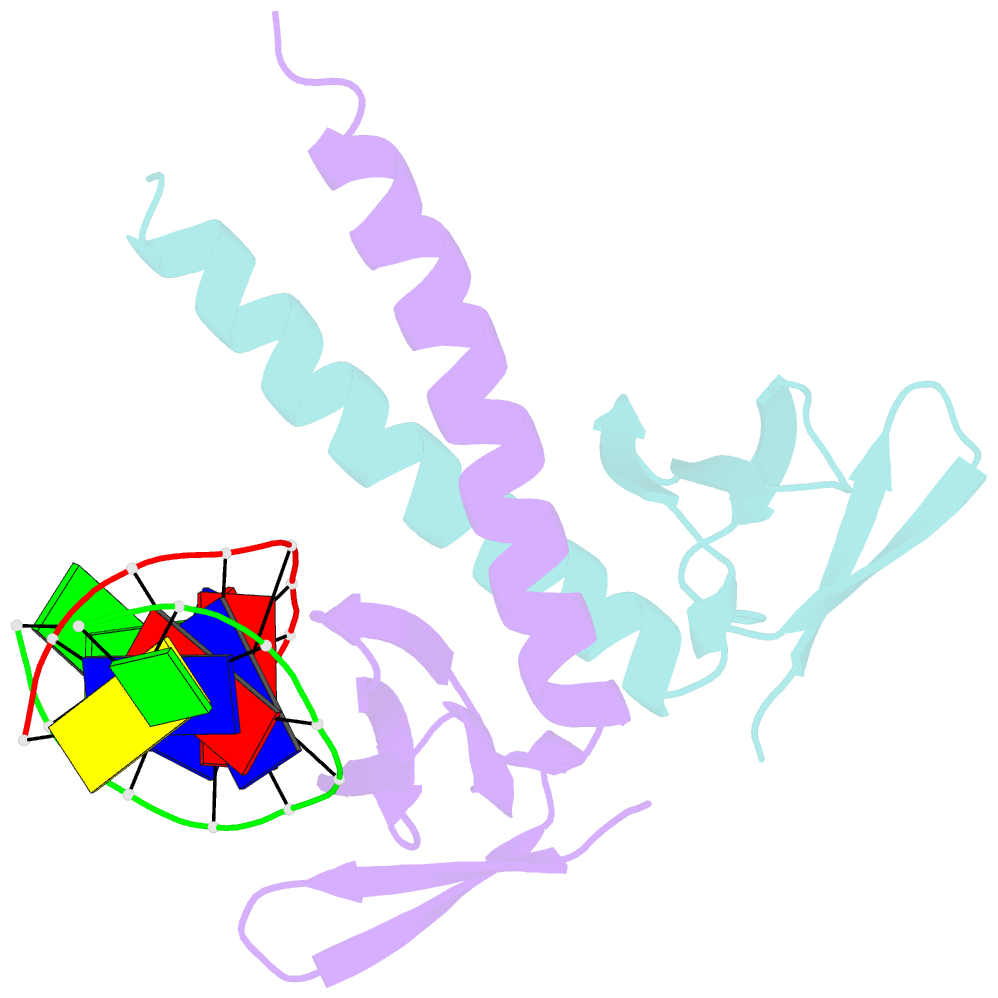
- Crystal structure of multimeric DNA-binding protein sac7d-gcn4 with DNA decamer
- Reference
- Wu SW, Ko TP, Chou CC, Wang AH (2005): "Design and characterization of a multimeric DNA binding protein using Sac7d and GCN4 as templates." Proteins, 60, 617-628. doi: 10.1002/prot.20524.
- Abstract
- The protein Sac7d belongs to a class of small chromosomal proteins from the hyperthermophilic archaeon Sulfolobus acidocaldarius. Sac7d is extremely stable to heat, acid, and chemical agents. This protein is a monomer and it binds DNA without any particular sequence preference, while inducing a sharp kink in the DNA. By appending a leucine-zipper-like helical peptide derived from the yeast transcriptional activator GCN4 to the C-terminal end of Sac7d, the modified monomers (denoted S7dLZ) are expected to interact with each other via hydrophobic force to form a parallel dimer. The recombinant S7dLZ was expressed in Escherichia coli and purified by heating and ion-exchange chromatography. The formation of dimer was detected by gel-filtration chromatography and chemical cross-link. The results of surface plasmon resonance and circular dichroism experiments showed that the DNA-binding capacity was retained. Furthermore, X-ray diffraction analysis of single crystals of S7dLZ in complex with DNA decamer CCTATATAGG showed that the leucine-zipper segments of S7dLZ were associated into an antiparallel four-helix bundle. There are two DNA fragments bound to each S7dLZ tetramer in the crystal. This model works as a successful template that endows protein a new function without losing original properties.