Summary information and primary citation
- PDB-id
- 1zvv; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription-DNA
- Method
- X-ray (2.98 Å)
- Summary
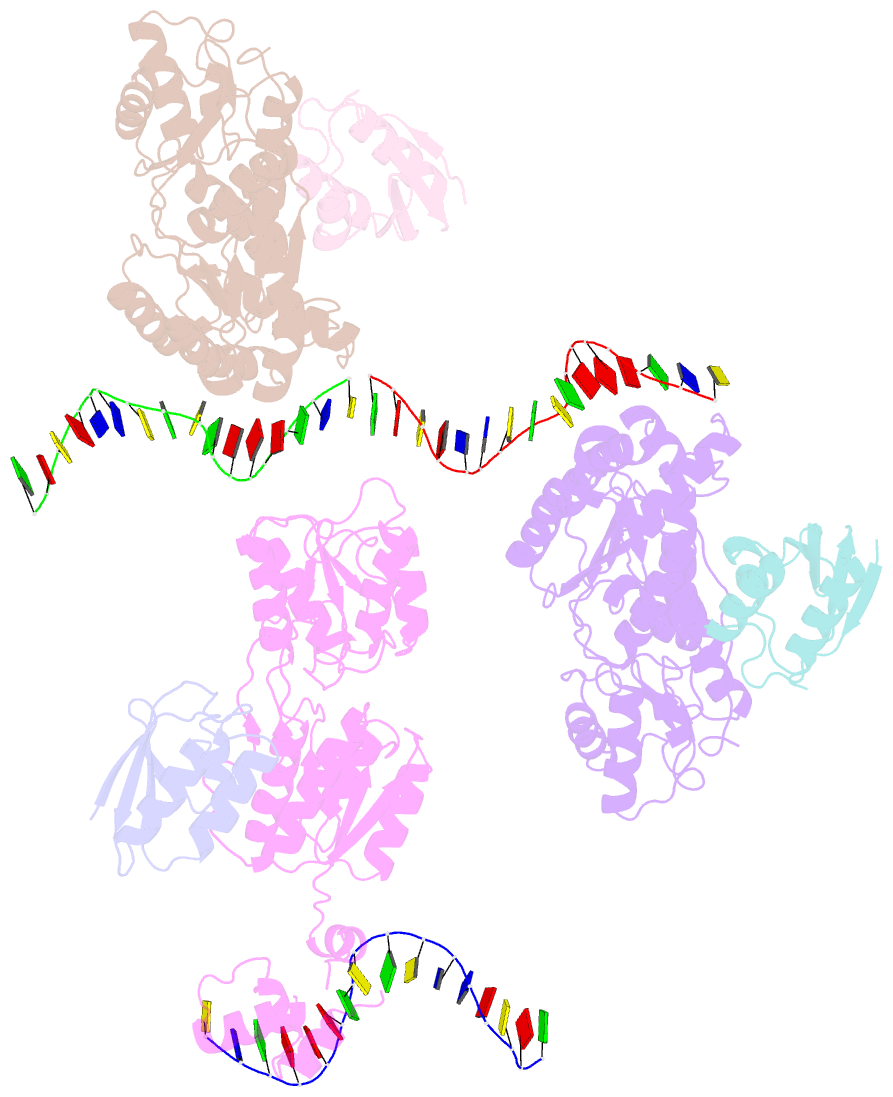
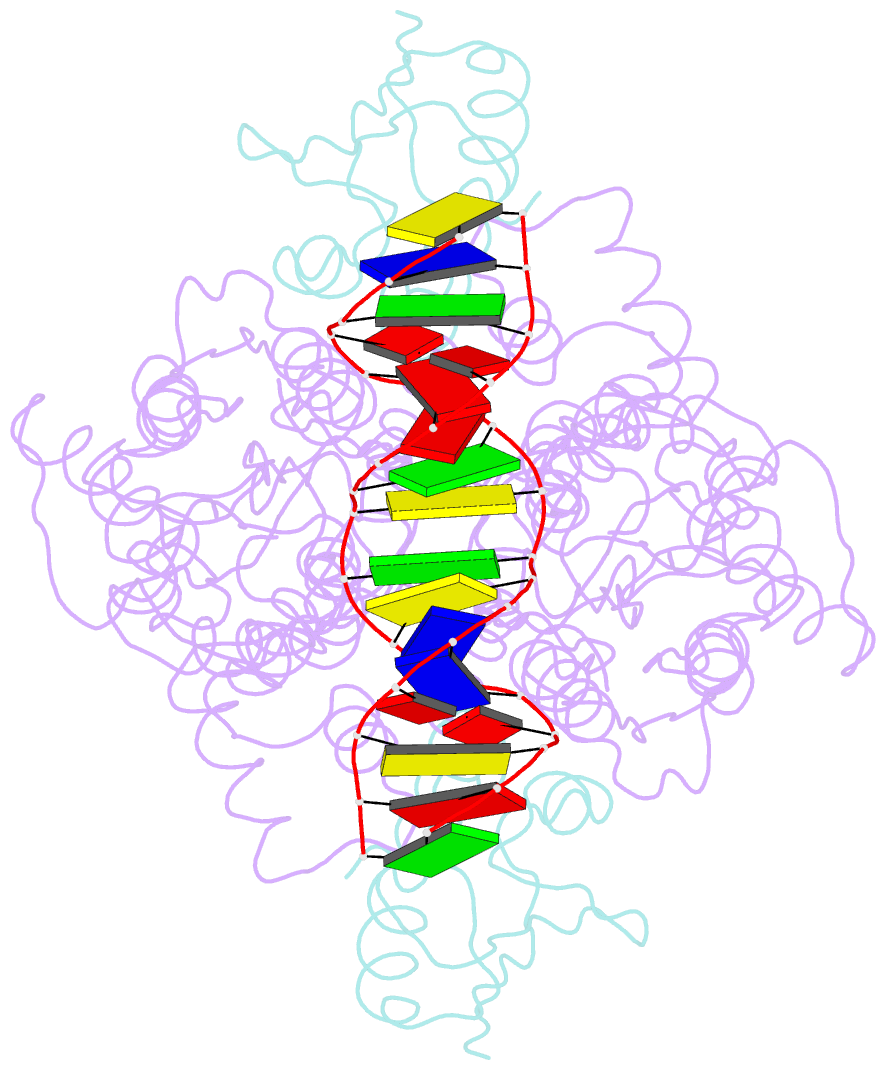
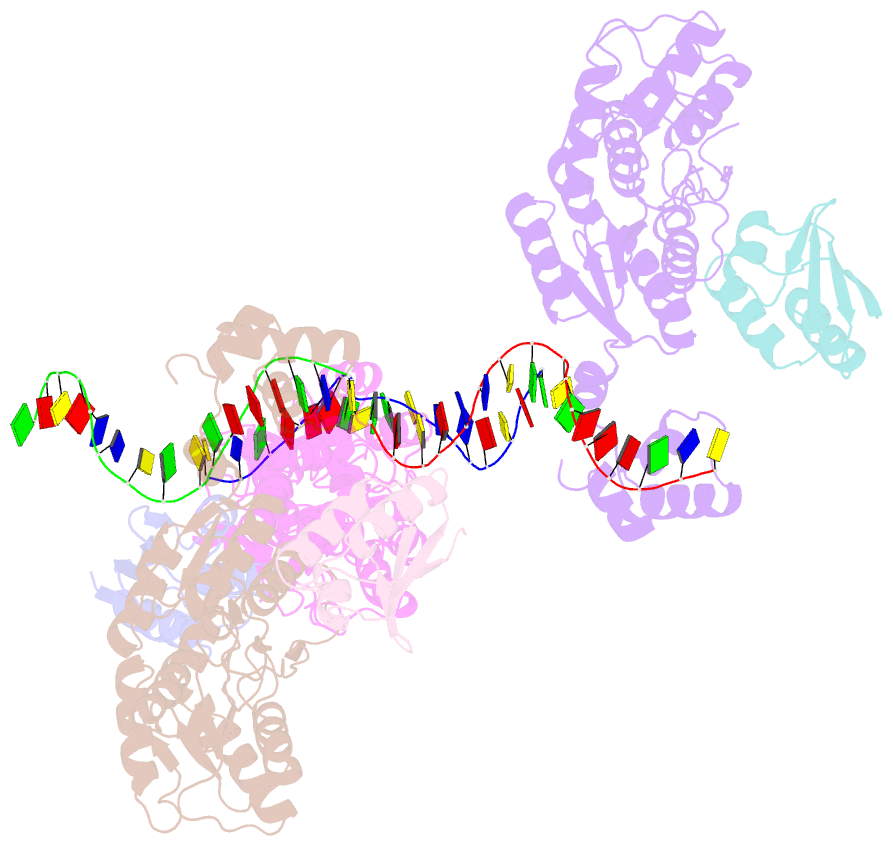
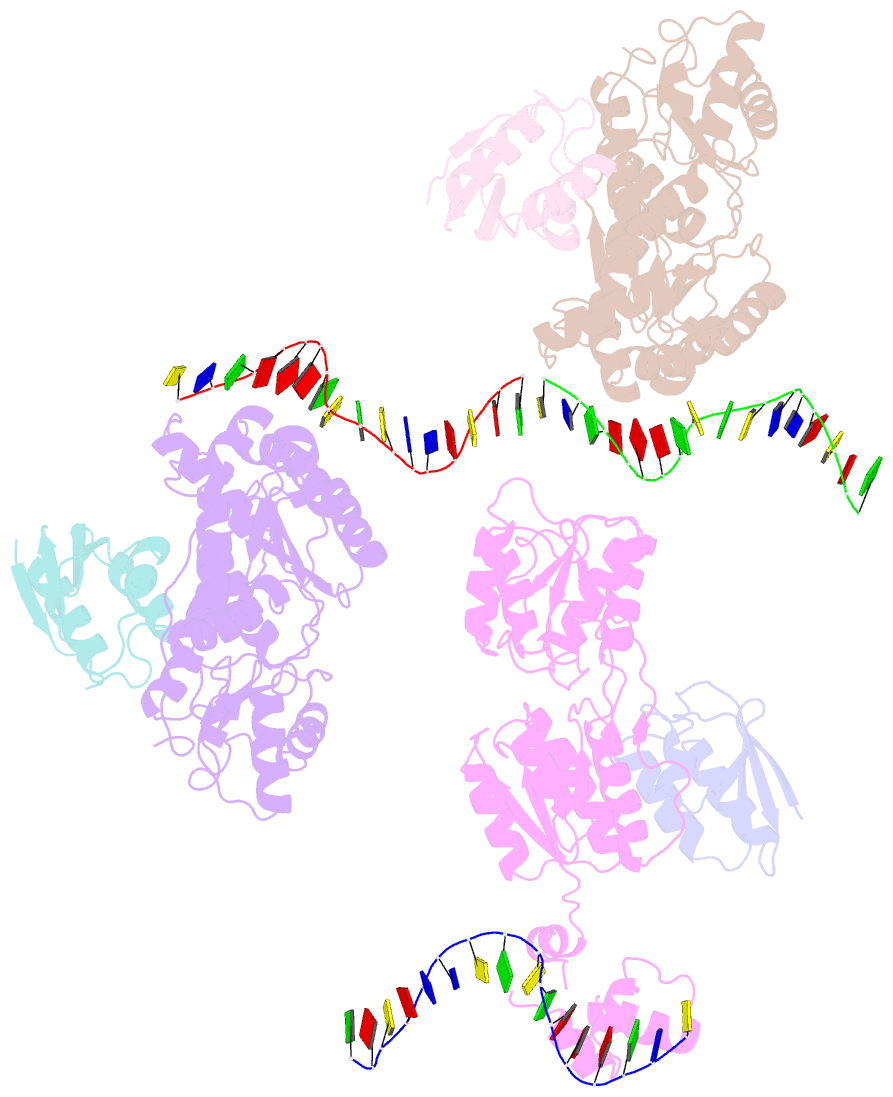
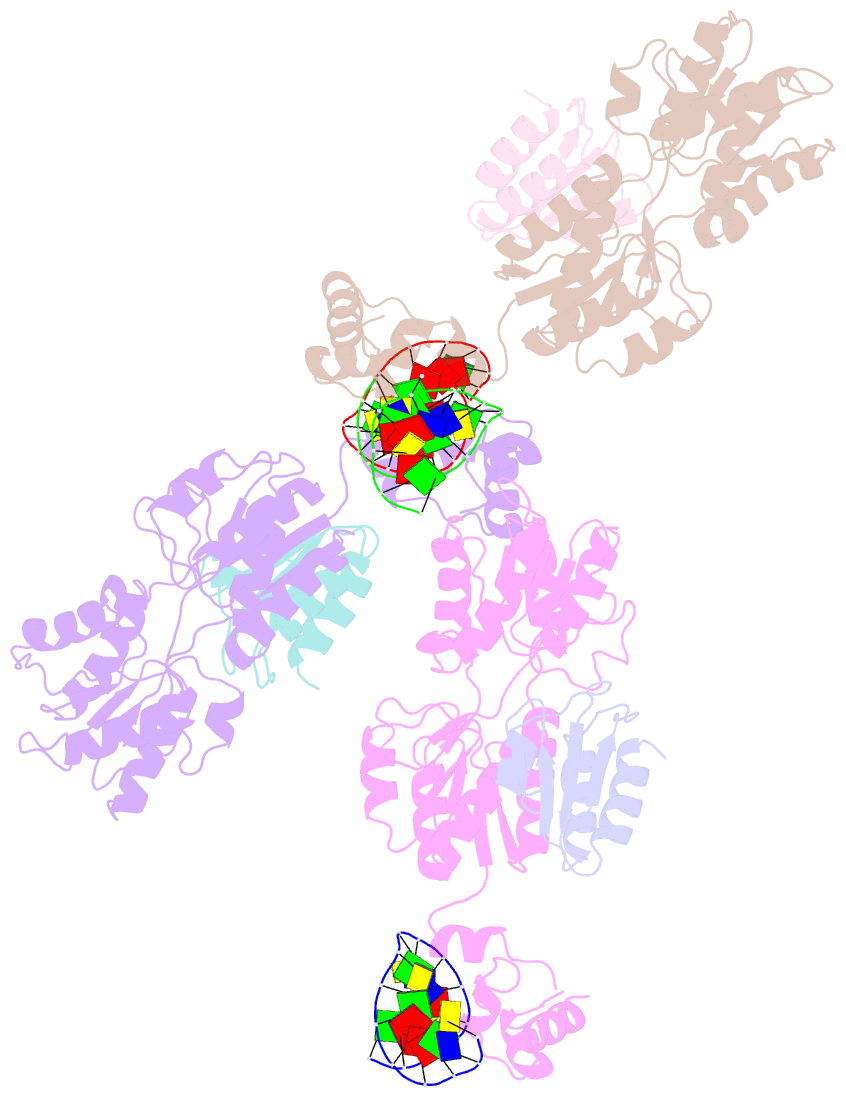
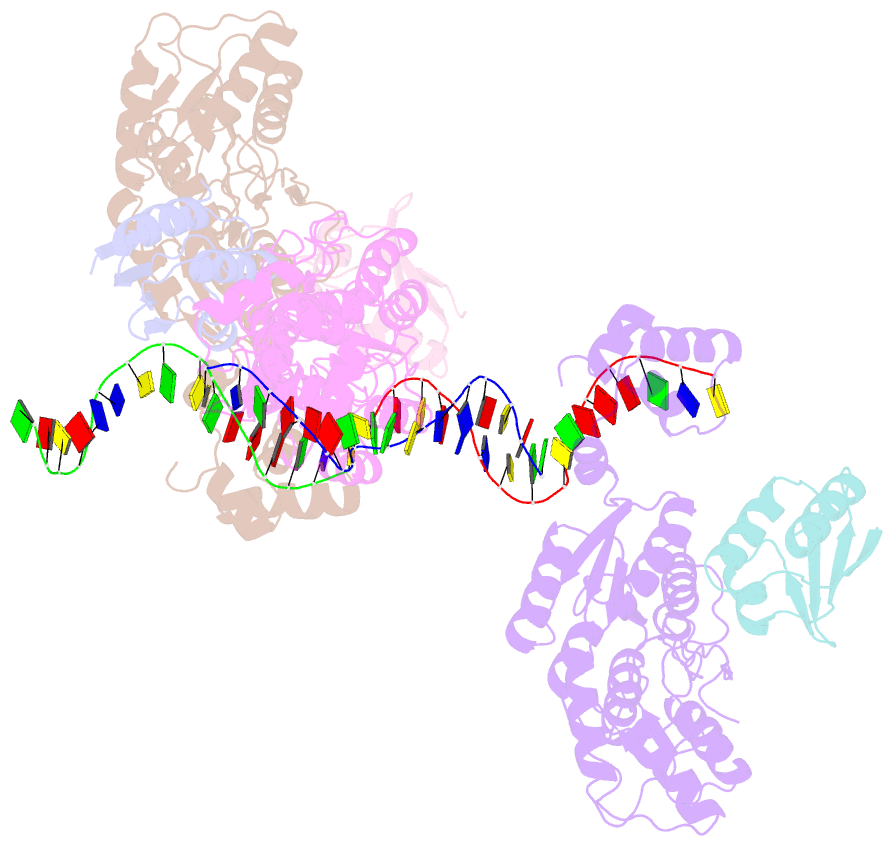
- Crystal structure of a ccpa-crh-DNA complex
- Reference
- Schumacher MA, Seidel G, Hillen W, Brennan RG (2006): "Phosphoprotein Crh-Ser46-P displays altered binding to CcpA to effect carbon catabolite regulation." J.Biol.Chem., 281, 6793-6800. doi: 10.1074/jbc.M509977200.
- Abstract
- In Gram-positive bacteria, the catabolite control protein A (CcpA) functions as the master transcriptional regulator of carbon catabolite repression/regulation (CCR). To effect CCR, CcpA binds a phosphoprotein, either HPr-Ser46-P or Crh-Ser46-P. Although Crh and histidine-containing protein (HPr) are structurally homologous, CcpA binds Crh-Ser46-P more weakly than HPr-Ser46-P. Moreover, Crh can form domain-swapped dimers, which have been hypothesized to be functionally relevant in CCR. To understand the molecular mechanism of Crh-Ser46-P regulation of CCR, we determined the structure of a CcpA-(Crh-Ser46-P)-DNA complex. The structure reveals that Crh-Ser46-P does not bind CcpA as a dimer but rather interacts with CcpA as a monomer in a manner similar to that of HPr-Ser46-P. The reduced affinity of Crh-Ser46-P for CcpA as compared with that of HPr-Ser46 P is explained by weaker Crh-Ser46-P interactions in its contact region I to CcpA, which causes this region to shift away from CcpA. Nonetheless, the interface between CcpA and helix alpha 2 of the second contact region (contact region II) of Crh-Ser46-P is maintained. This latter finding demonstrates that this contact region is necessary and sufficient to throw the allosteric switch to activate cre binding by CcpA.