Summary information and primary citation
- PDB-id
- 2h1o; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- gene regulation-DNA complex
- Method
- X-ray (3.0 Å)
- Summary
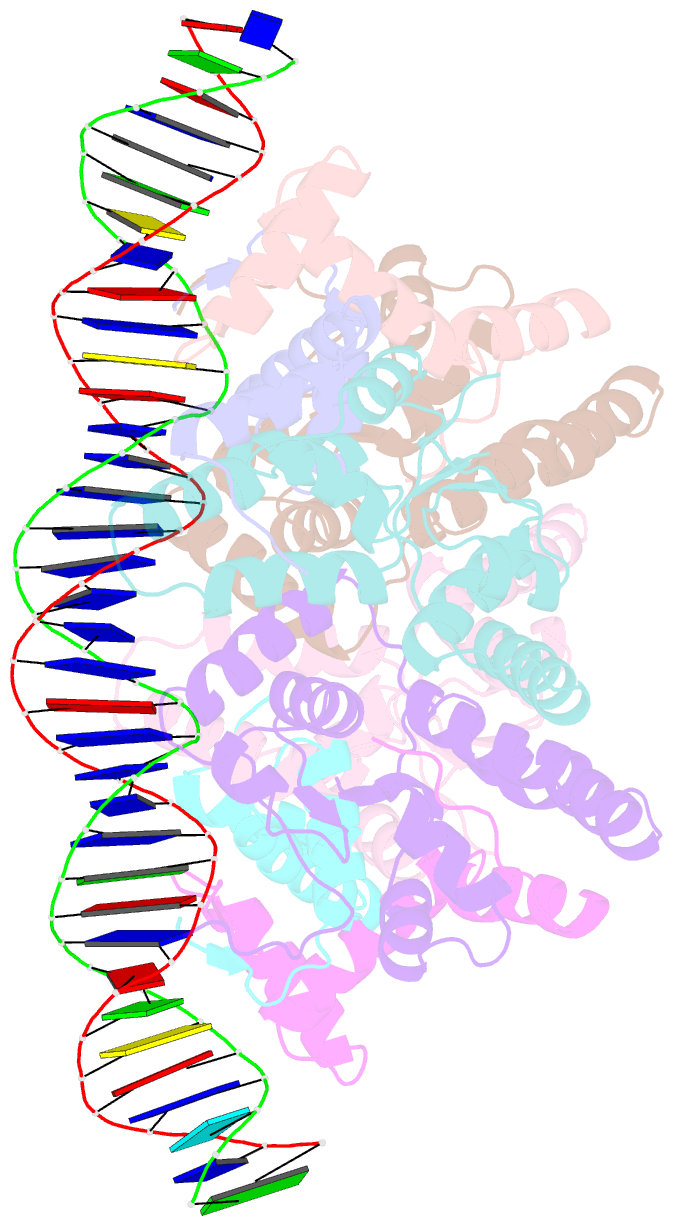
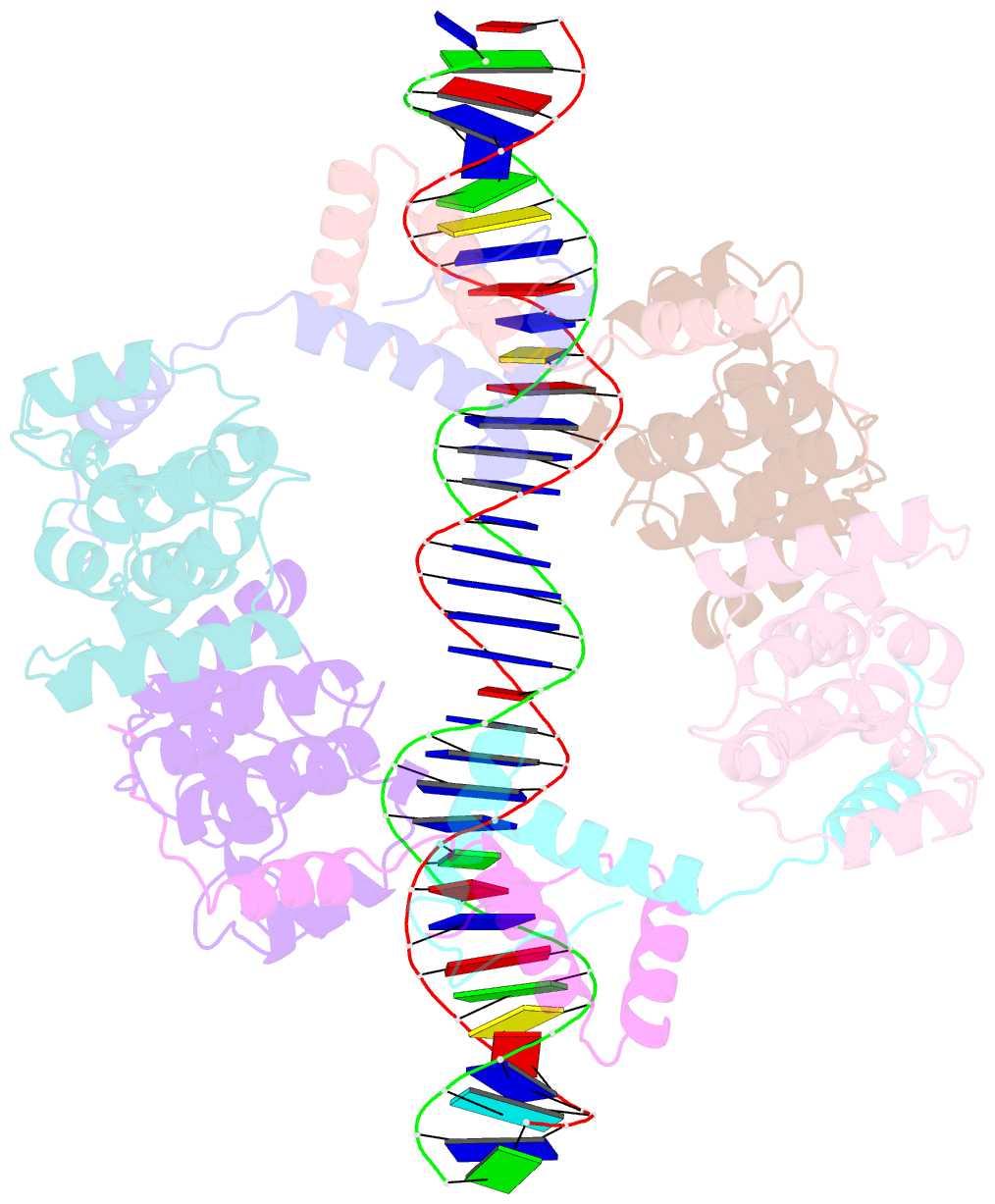
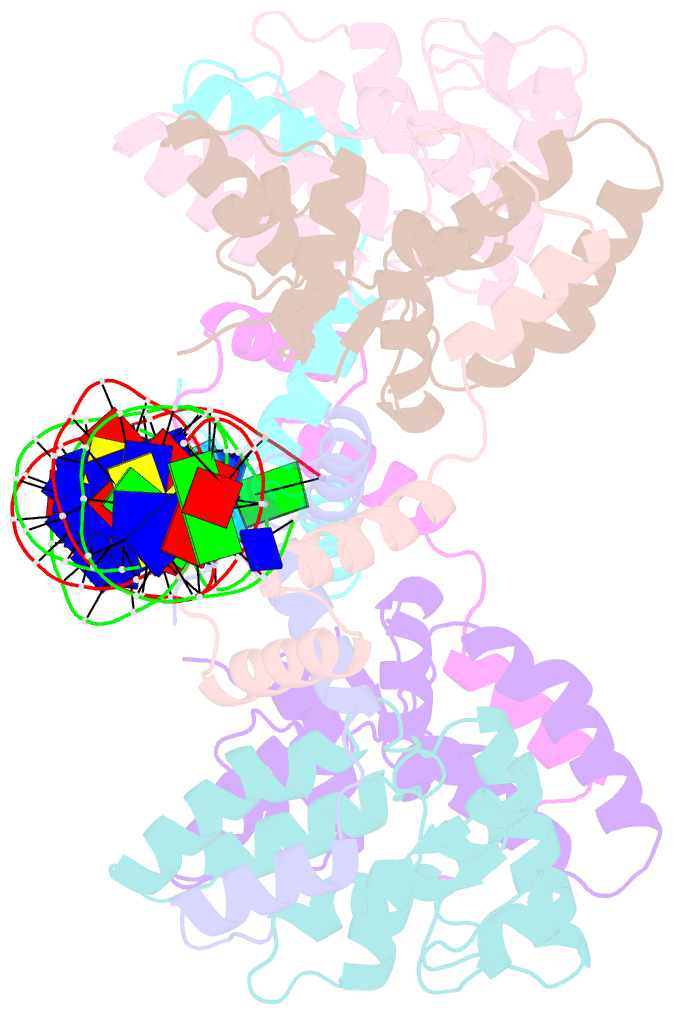
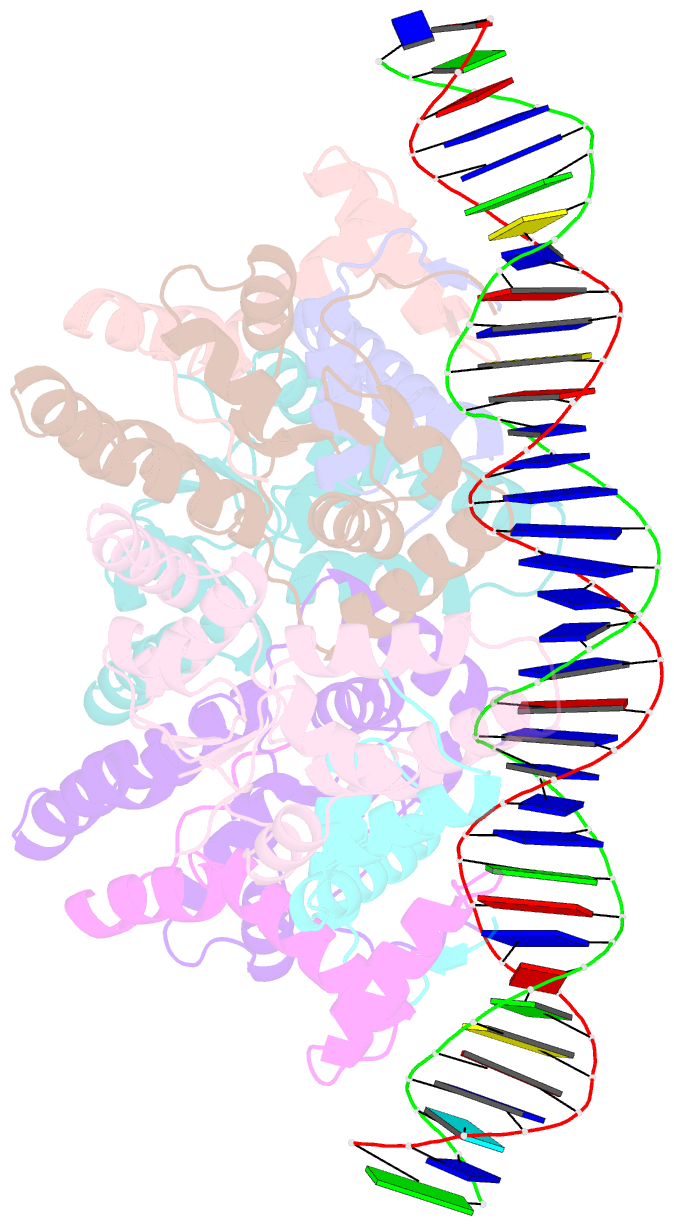
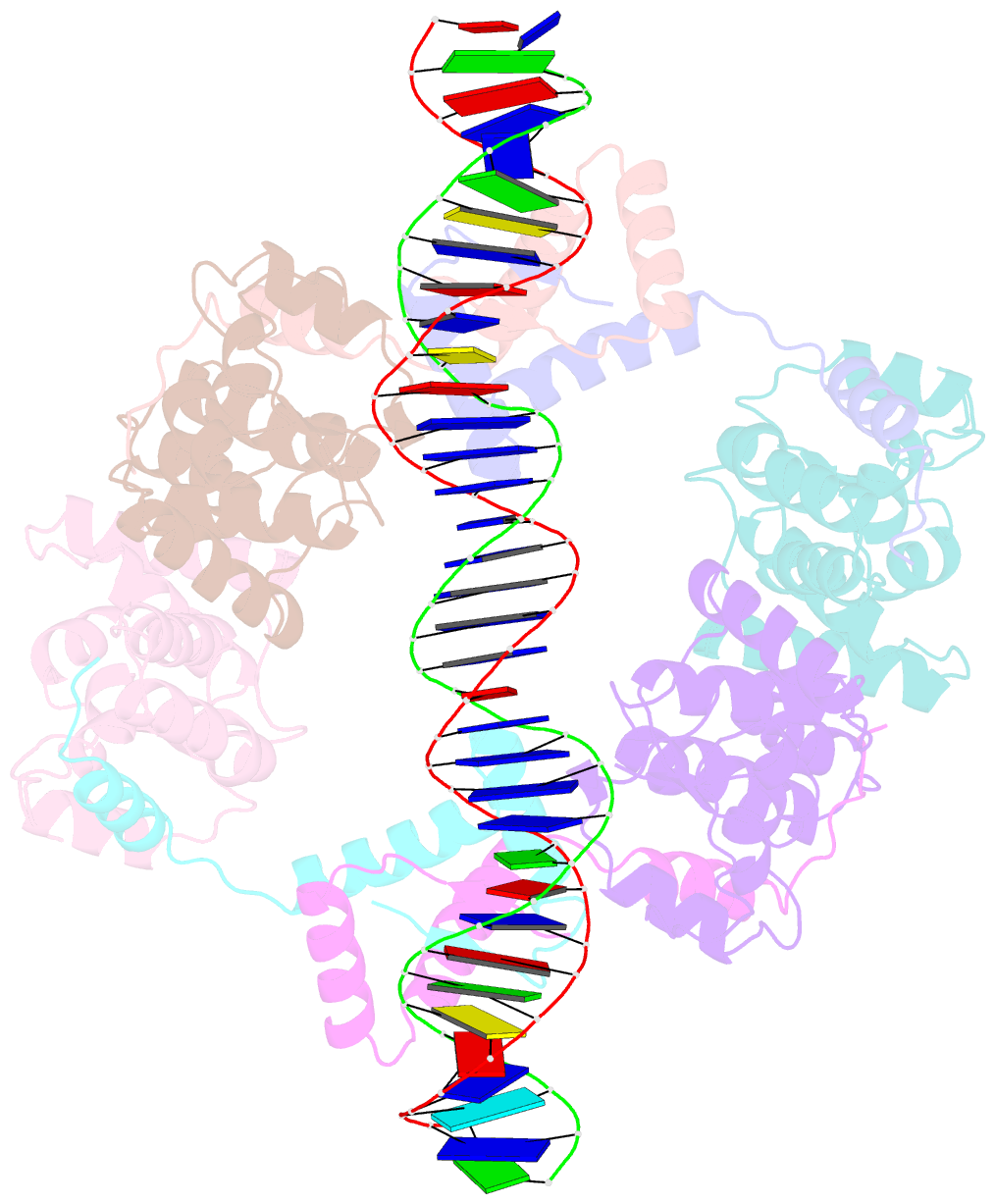
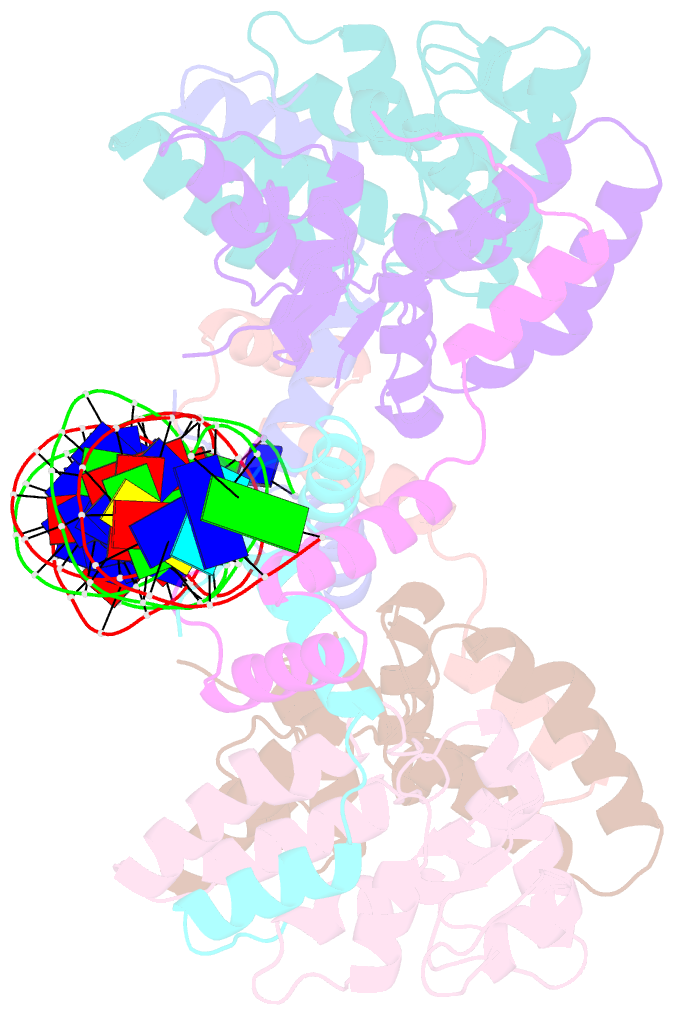
- Structure of fitab bound to ir36 DNA fragment
- Reference
- Mattison K, Wilbur JS, So M, Brennan RG (2006): "Structure of FitAB from Neisseria gonorrhoeae bound to DNA reveals a tetramer of toxin-antitoxin heterodimers containing pin domains and ribbon-helix-helix motifs." J.Biol.Chem., 281, 37942-37951. doi: 10.1074/jbc.M605198200.
- Abstract
- Neisseria gonorrhoeae is a sexually transmitted pathogen that initiates infections in humans by adhering to the mucosal epithelium of the urogenital tract. The bacterium then enters the apical region of the cell and traffics across the cell to exit into the subepithelial matrix. Mutations in the fast intracellular trafficking (fitAB) locus cause the bacteria to transit a polarized epithelial monolayer more quickly than the wild-type parent and to replicate within cells at an accelerated rate. Here, we describe the crystal structure of the toxin-antitoxin heterodimer, FitAB, bound to a high affinity 36-bp DNA fragment from the fitAB promoter. FitA, the antitoxin, binds DNA through its ribbon-helix-helix motif and is tethered to FitB, the toxin, to form a heterodimer by the insertion of a four turn alpha-helix into an extensive FitB hydrophobic pocket. FitB is composed of a PIN (PilT N terminus) domain, with a central, twisted, 5-stranded parallel beta-sheet that is open on one side and flanked by five alpha-helices. FitB in the context of the FitAB complex does not display nuclease activity against tested PIN substrates. The FitAB complex points to the mechanism by which antitoxins with RHH motifs can block the activity of toxins with PIN domains. Interactions between two FitB molecules result in the formation of a tetramer of FitAB heterodimers, which binds to the 36-bp DNA fragment and provides an explanation for how FitB enhances the DNA binding affinity of FitA.