Summary information and primary citation
- PDB-id
- 2hof; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- recombination-DNA
- Method
- X-ray (2.4 Å)
- Summary
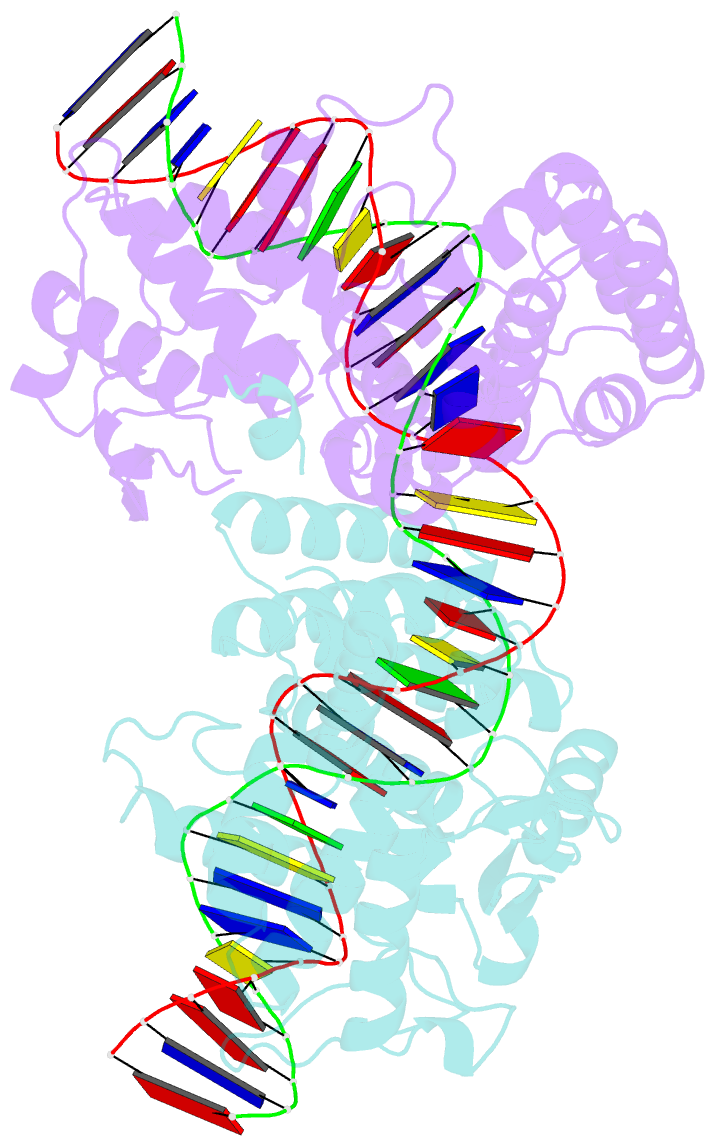
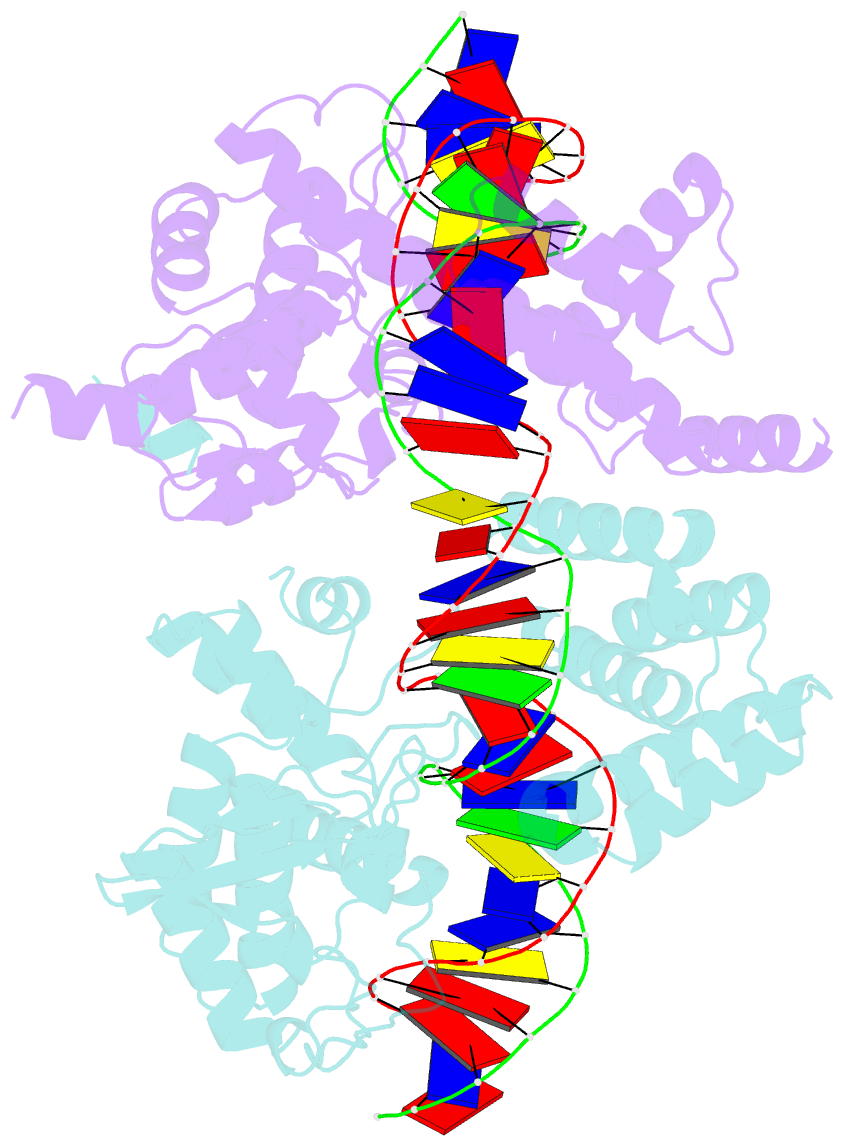
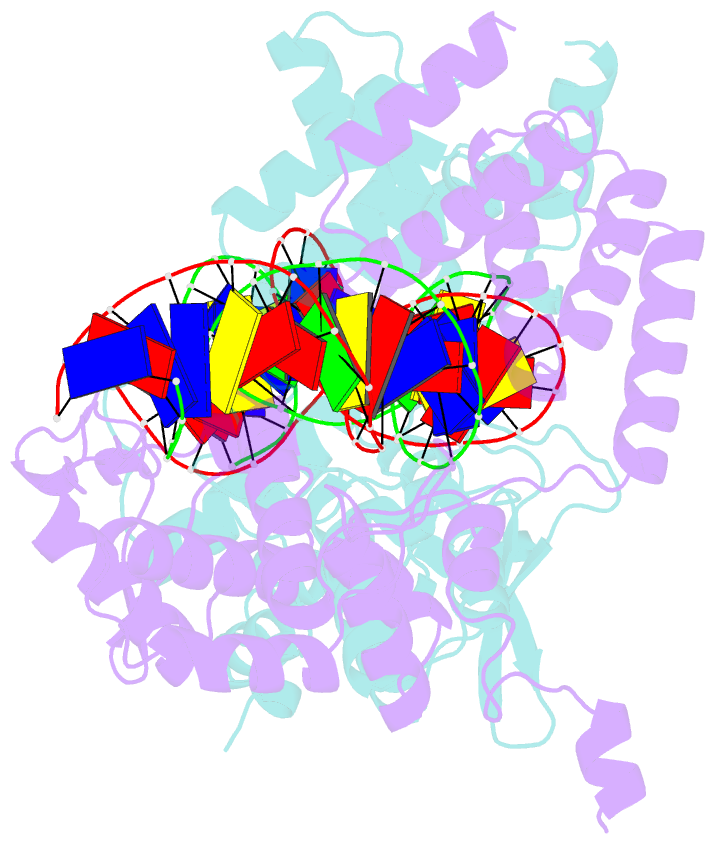
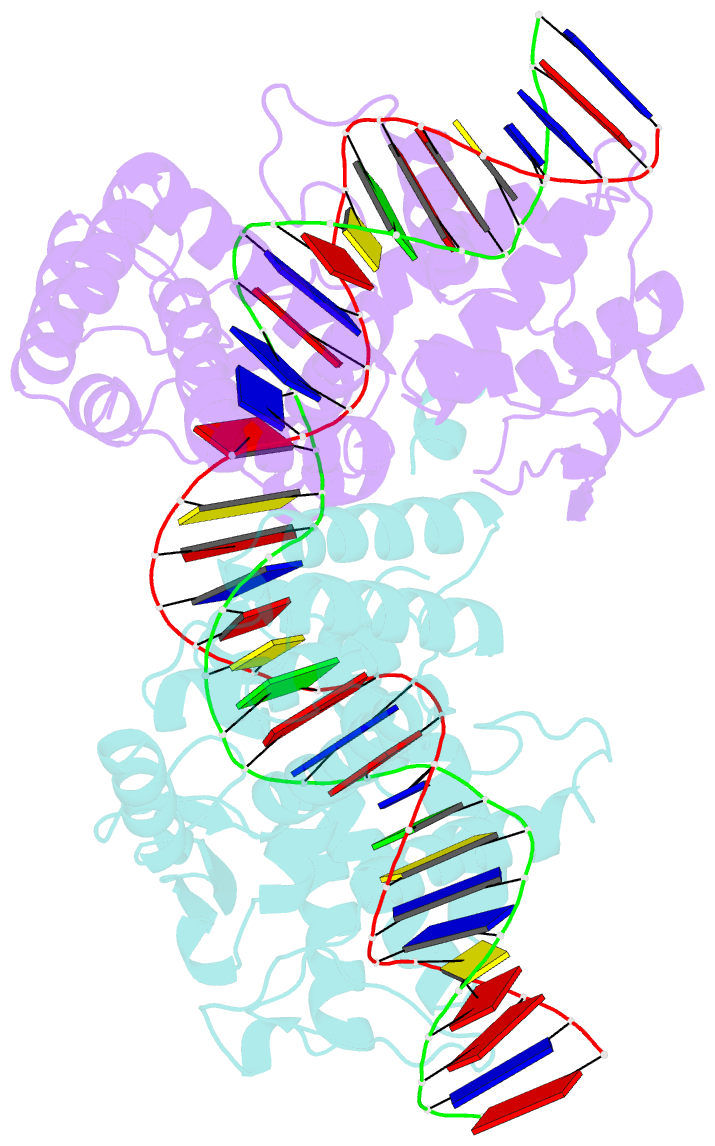
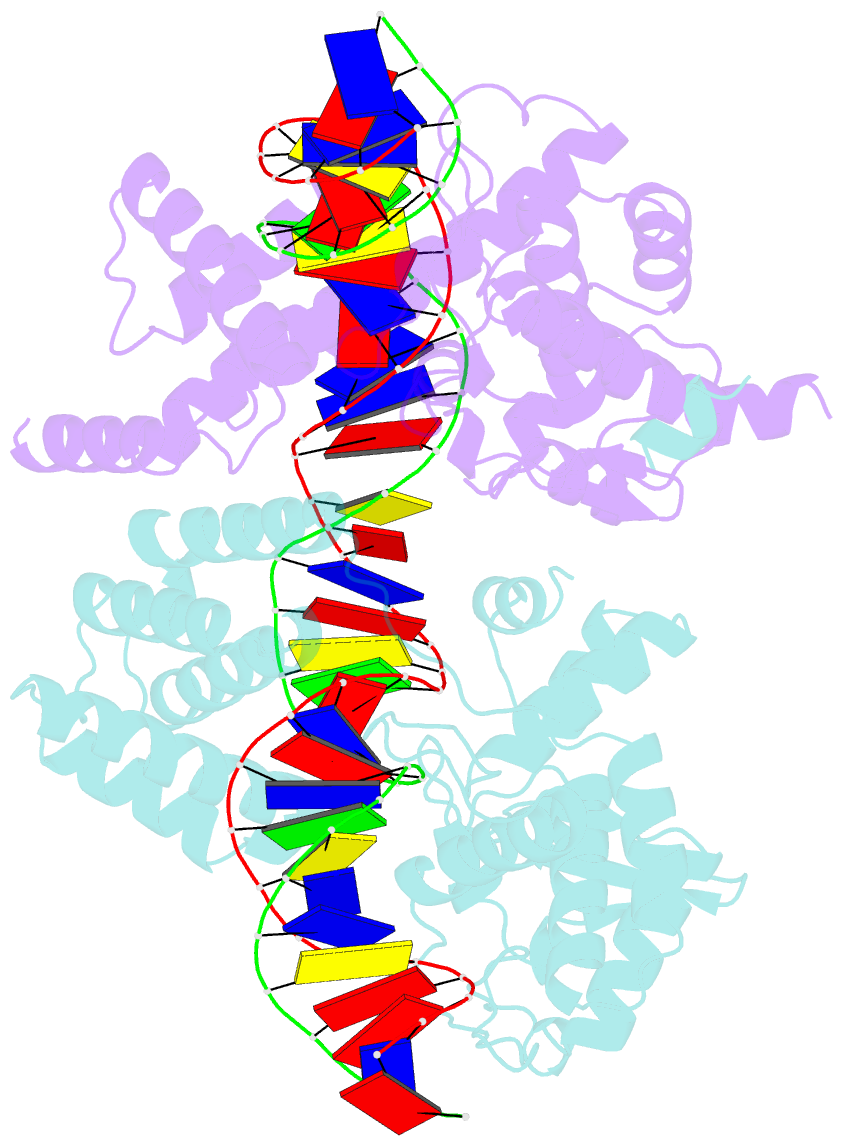
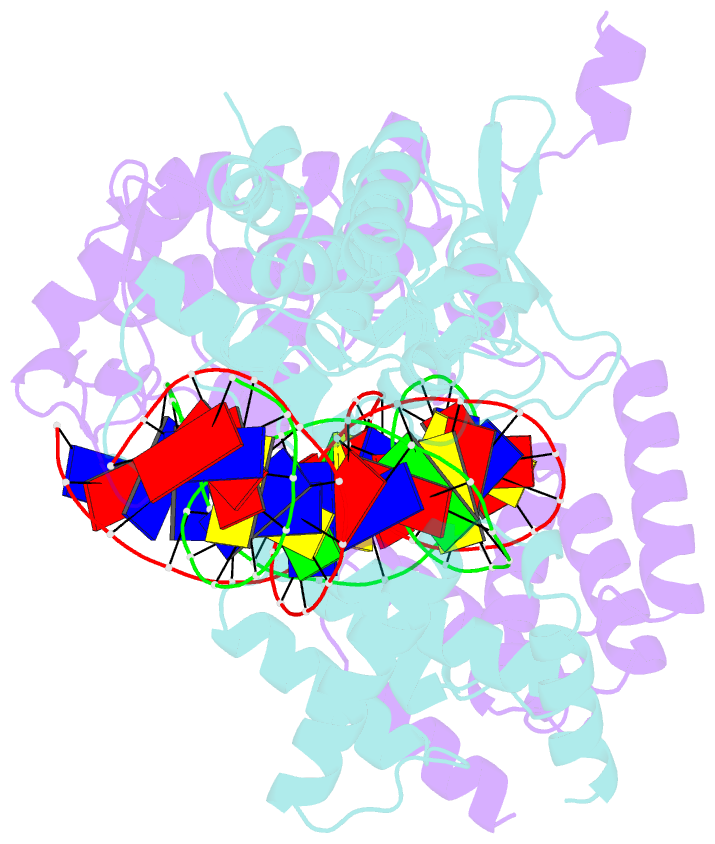
- Crystal structure of the pre-cleavage synaptic complex in the cre-loxp site-specific recombination
- Reference
- Ghosh K, Guo F, Van Duyne GD (2007): "Synapsis of loxP sites by Cre recombinase." J.Biol.Chem., 282, 24004-24016. doi: 10.1074/jbc.M703283200.
- Abstract
- Cre recombinase catalyzes site-specific recombination between 34-bp loxP sites in a variety of topological and cellular contexts. An obligatory step in the recombination reaction is the association, or synapsis, of Cre-bound loxP sites to form a tetrameric protein assembly that is competent for strand exchange. Using analytical ultracentrifugation and electrophoresis approaches, we have studied the energetics of Cre-mediated synapsis of loxP sites. We found that synapsis occurs with a high affinity (Kd = 10 nM) and is pH-dependent but does not require divalent cations. Surprisingly, the catalytically inactive Cre K201A mutant is fully competent for synapsis of loxP sites, yet the inactive Y324F and R173K mutants are defective for synapsis. These findings have allowed us to determine the first crystal structures of a pre-cleavage Cre-loxP synaptic complex in a configuration representing the starting point in the recombination pathway. When combined with a quantitative analysis of synapsis using loxP mutants, the structures explain how the central 8 bp of the loxP site are able to dictate the order of strand exchange in the Cre system.