Summary information and primary citation
- PDB-id
- 2iso; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-DNA
- Method
- X-ray (2.1 Å)
- Summary
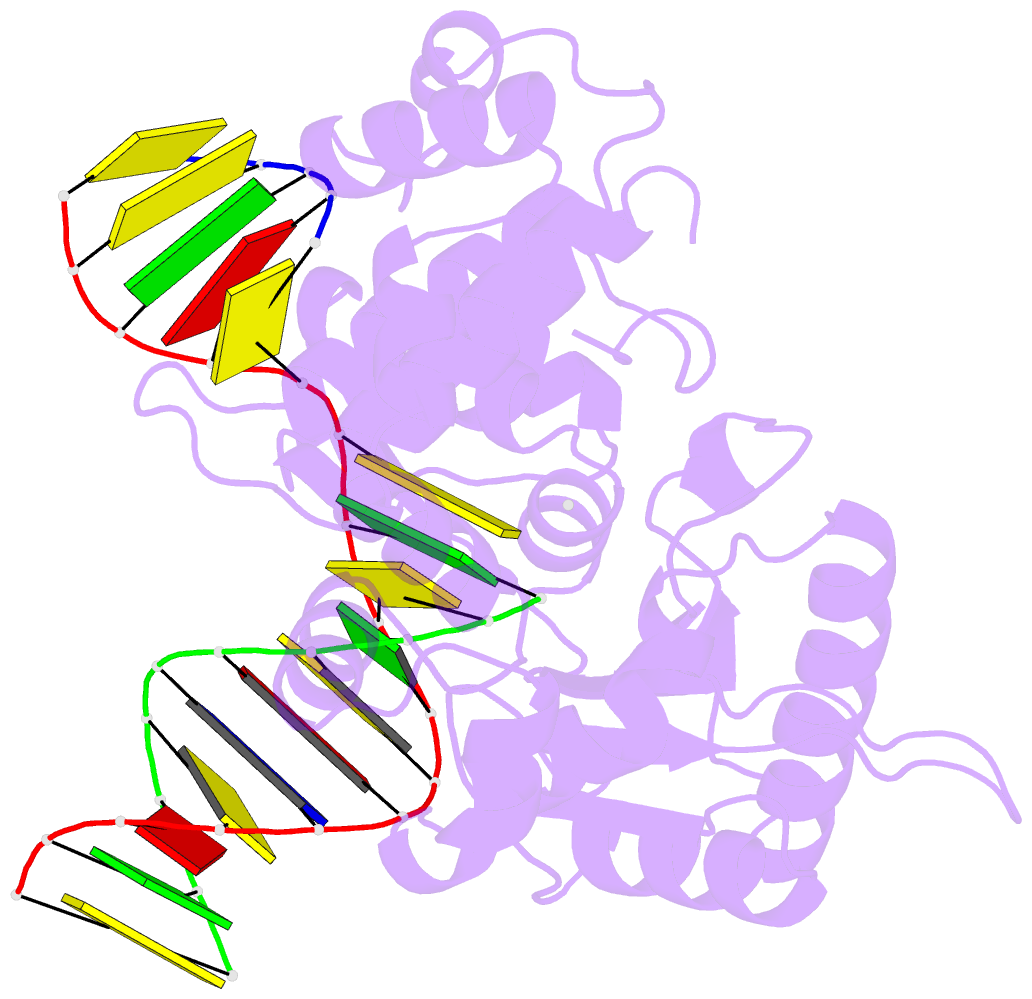
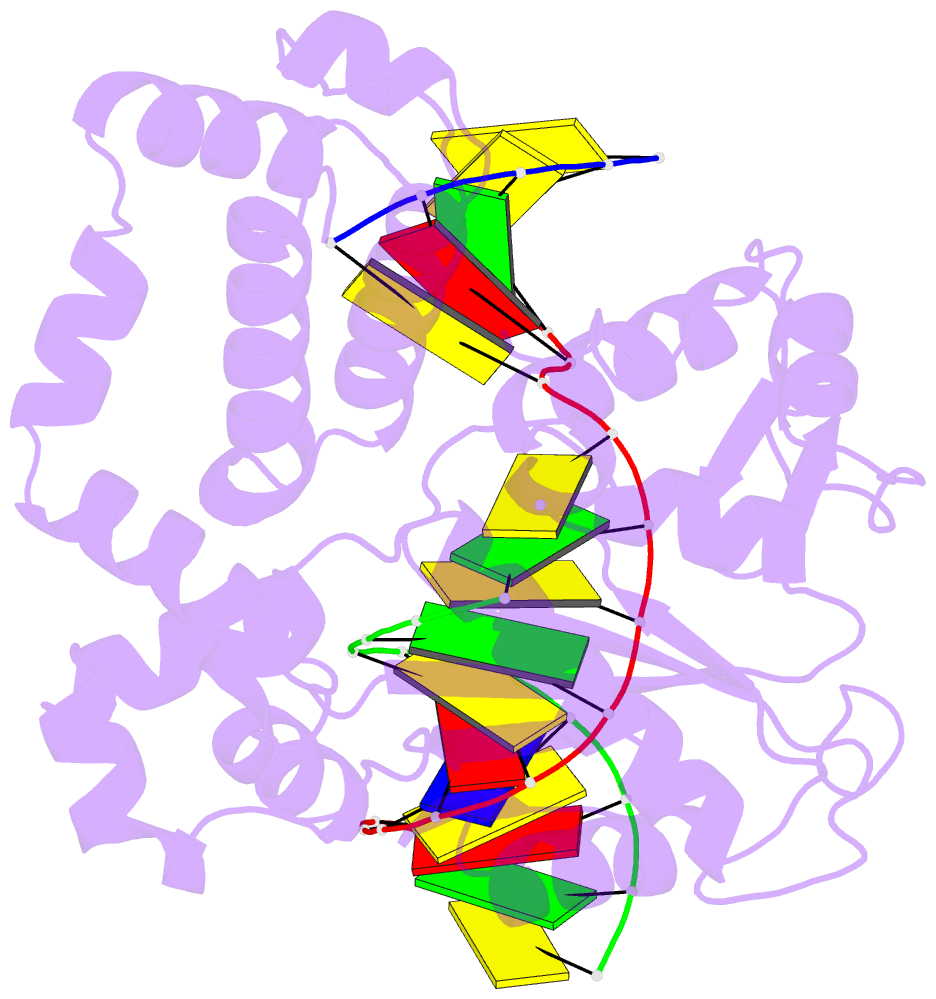
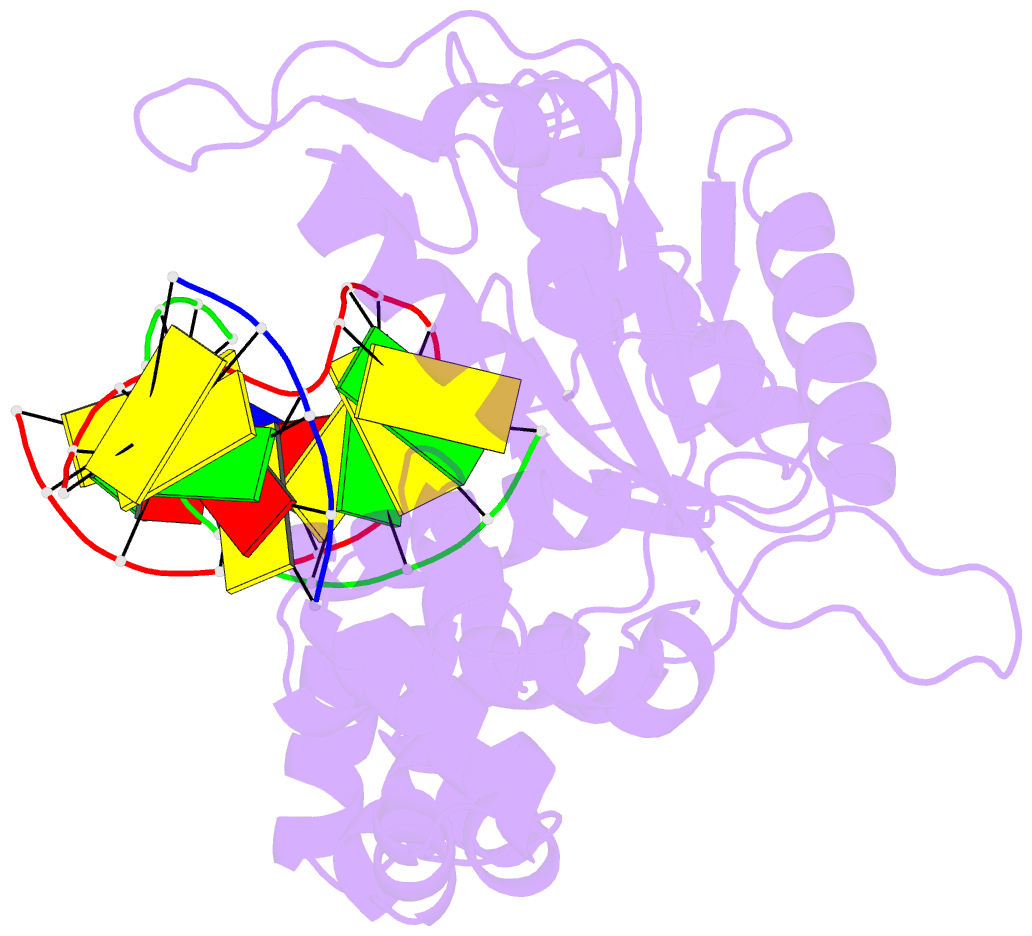
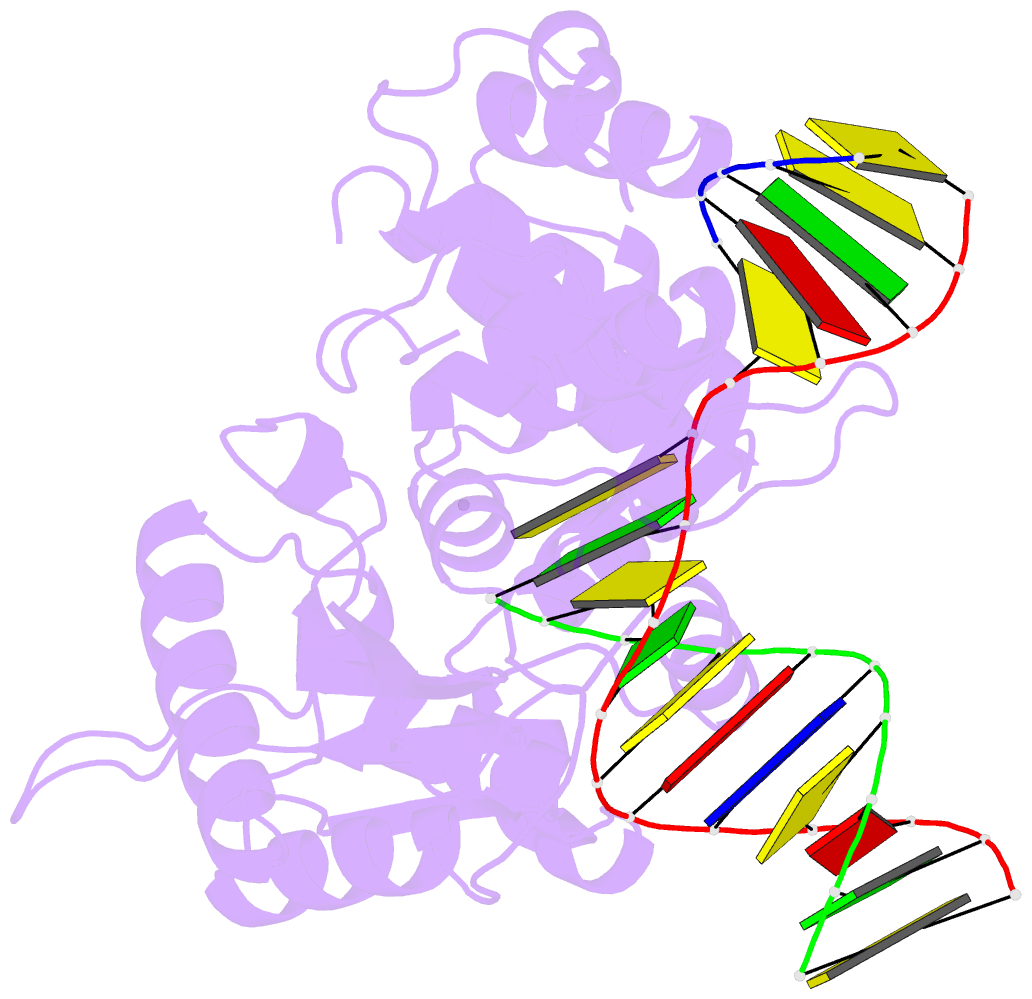
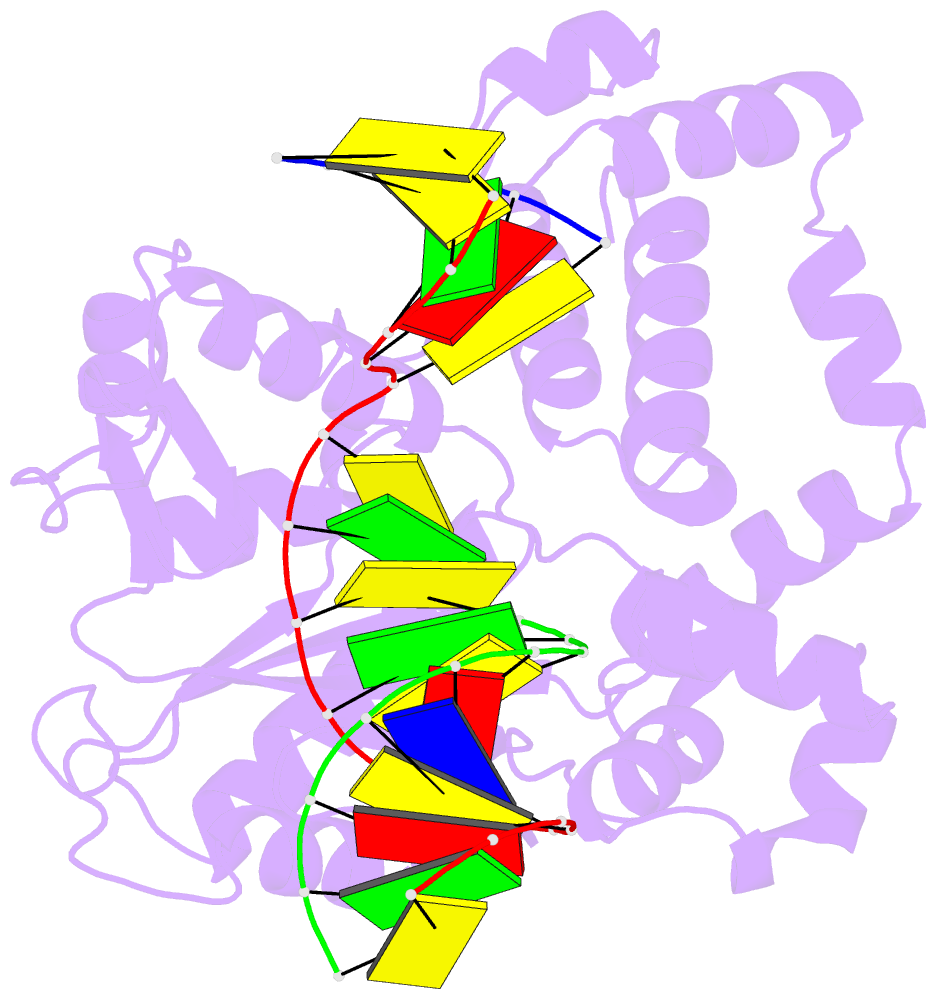
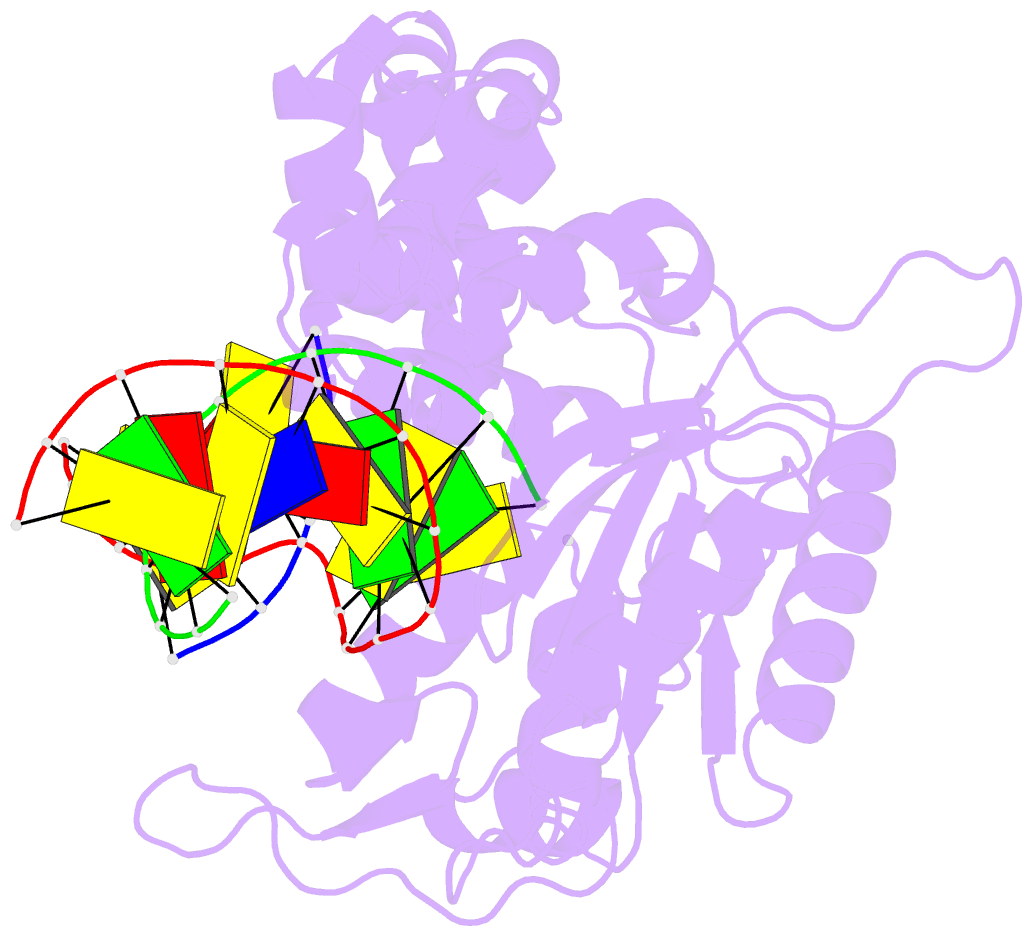
- Ternary complex of DNA polymerase beta with a dideoxy terminated primer and 2'-deoxyguanosine 5'-beta, gamma-difluoromethylene triphosphate
- Reference
- Sucato CA, Upton TG, Kashemirov BA, Batra VK, Martinek V, Xiang Y, Beard WA, Pedersen LC, Wilson SH, McKenna CE, Florian J, Warshel A, Goodman MF (2007): "Modifying the beta,gamma Leaving-Group Bridging Oxygen Alters Nucleotide Incorporation Efficiency, Fidelity, and the Catalytic Mechanism of DNA Polymerase beta." Biochemistry, 46, 461-471. doi: 10.1021/bi061517b.
- Abstract
- DNA polymerase catalysis and fidelity studies typically compare incorporation of "right" versus "wrong" nucleotide bases where the leaving group is pyrophosphate. Here we use dGTP analogues replacing the beta,gamma-bridging O with CH2, CHF, CF2, or CCl2 to explore leaving-group effects on the nucleotidyl transfer mechanism and fidelity of DNA polymerase (pol) beta. T.G mismatches occur with fidelities similar to dGTP with the exception of the CH2 analogue, which is incorporated with 5-fold higher fidelity. All analogues are observed to bind opposite template C with Kds between 1 and 4 microM, and structural evidence suggests that the analogues bind in essentially the native conformation, making them suitable substrates for probing linear free energy relationships (LFERs) in transient-kinetics experiments. Importantly, Brnsted correlations of log(kpol) versus leaving-group pKa for both right and wrong base incorporation reveal similar sensitivities (betalg approximately -0.8) followed by departures from linearity, suggesting that a chemical step rather than enzyme conformational change is rate-limiting for either process. The location of the breaks relative to pKas of CF2, O, and the sterically bulky CCl2-bridging compounds suggests a modification-induced change in the mechanism by stabilization of leaving-group elimination. The results are addressed theoretically in terms of the energetics of successive primer 3'-O addition (bond forming) and pyrophosphate analogue elimination (bond breaking) reaction energy barriers.