Summary information and primary citation
- PDB-id
- 2j0q; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydrolase
- Method
- X-ray (3.2 Å)
- Summary
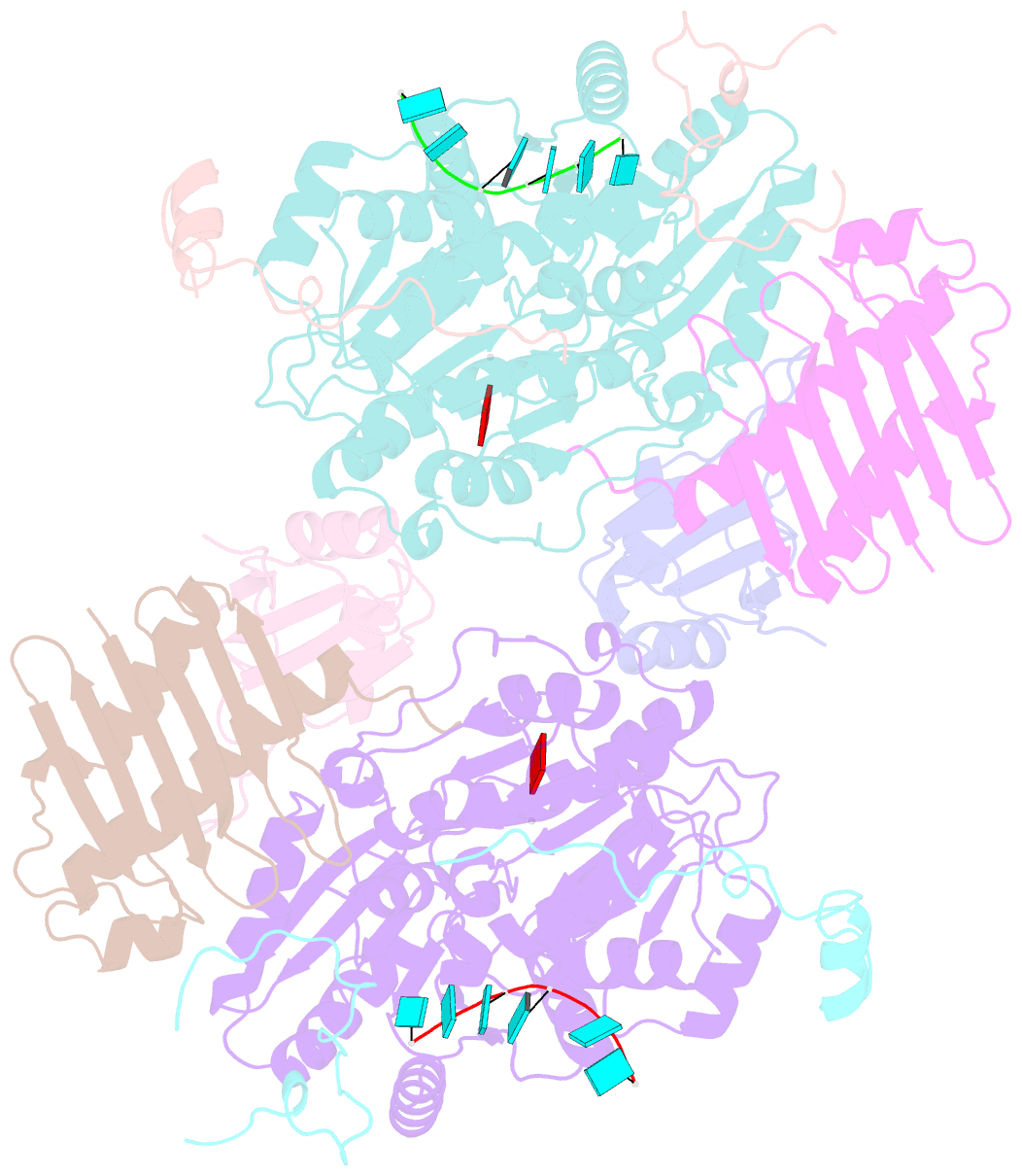
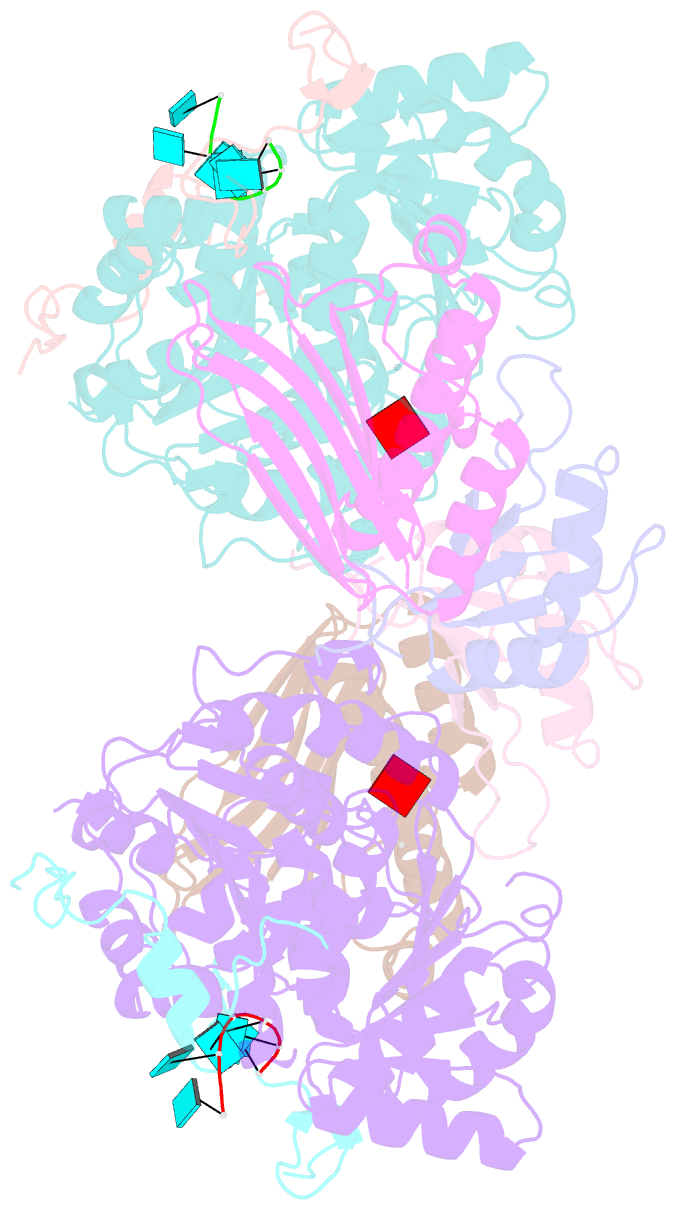
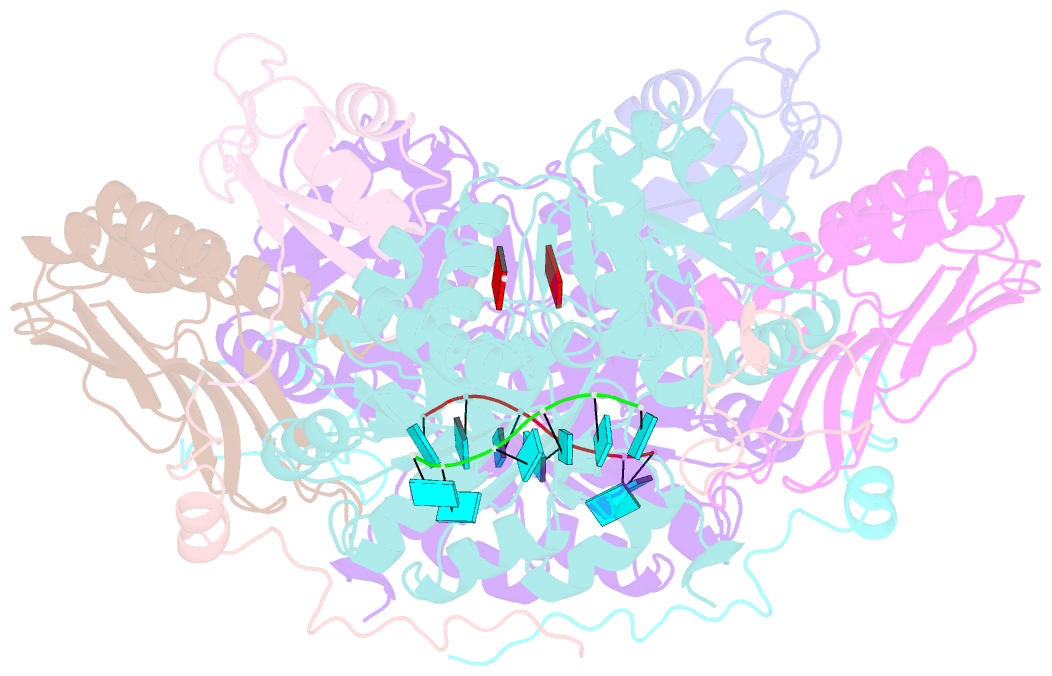
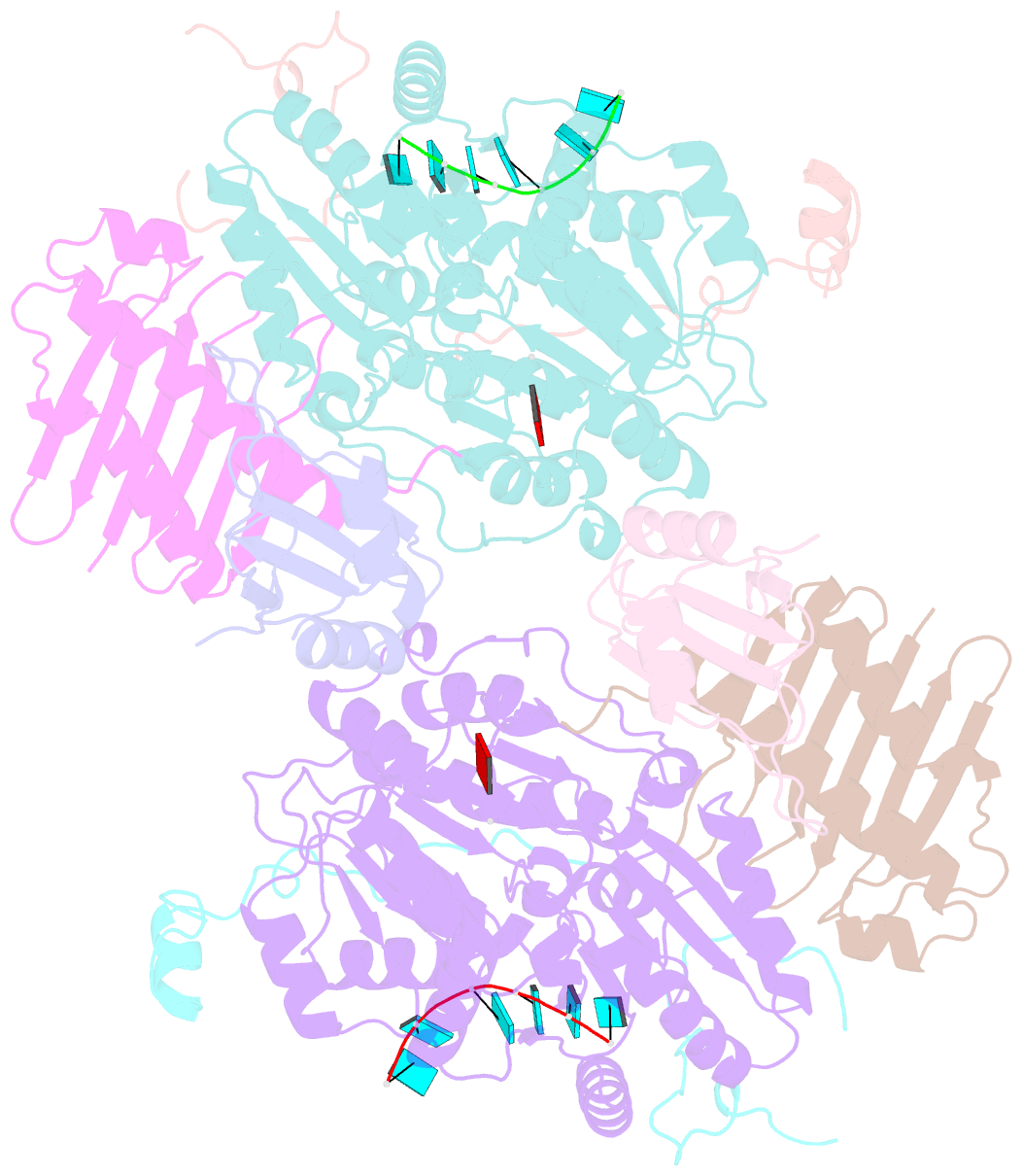
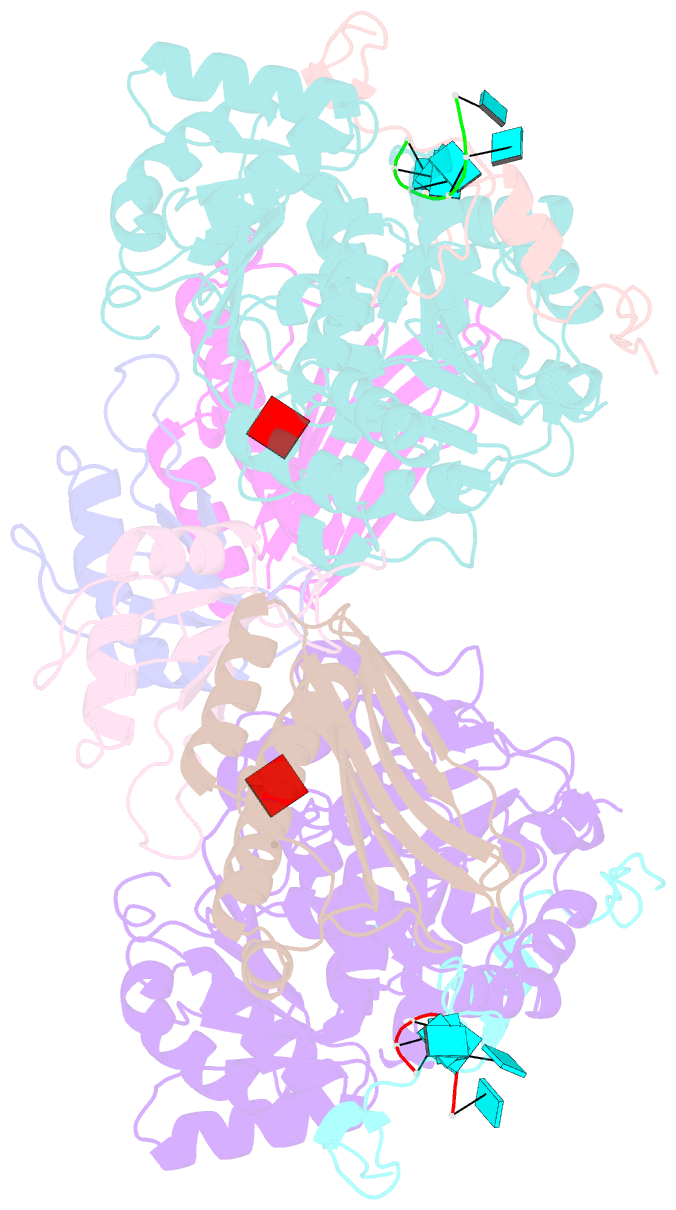
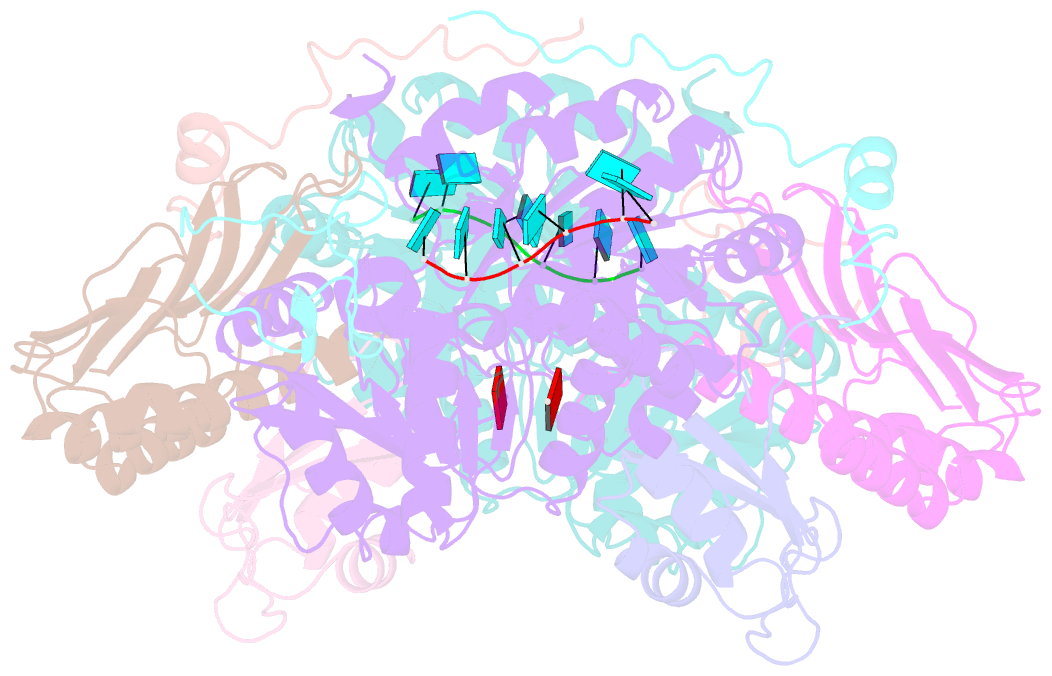
- The crystal structure of the exon junction complex at 3.2 Å resolution
- Reference
- Bono F, Ebert J, Lorentzen E, Conti E (2006): "The Crystal Structure of the Exon Junction Complex Reveals How It Maintains a Stable Grip on Mrna." Cell(Cambridge,Mass.), 126, 713. doi: 10.1016/J.CELL.2006.08.006.
- Abstract
- The exon junction complex (EJC) plays a major role in posttranscriptional regulation of mRNA in metazoa. The EJC is deposited onto mRNA during splicing and is transported to the cytoplasm where it influences translation, surveillance, and localization of the spliced mRNA. The complex is formed by the association of four proteins (eIF4AIII, Barentsz [Btz], Mago, and Y14), mRNA, and ATP. The 2.2 A resolution structure of the EJC reveals how it stably locks onto mRNA. The DEAD-box protein eIF4AIII encloses an ATP molecule and provides the binding sites for six ribonucleotides. Btz wraps around eIF4AIII and stacks against the 5' nucleotide. An intertwined network of interactions anchors Mago-Y14 and Btz at the interface between the two domains of eIF4AIII, effectively stabilizing the ATP bound state. Comparison with the structure of the eIF4AIII-Btz subcomplex that we have also determined reveals that large conformational changes are required upon EJC assembly and disassembly.