Summary information and primary citation
- PDB-id
- 2jei; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase
- Method
- X-ray (2.39 Å)
- Summary
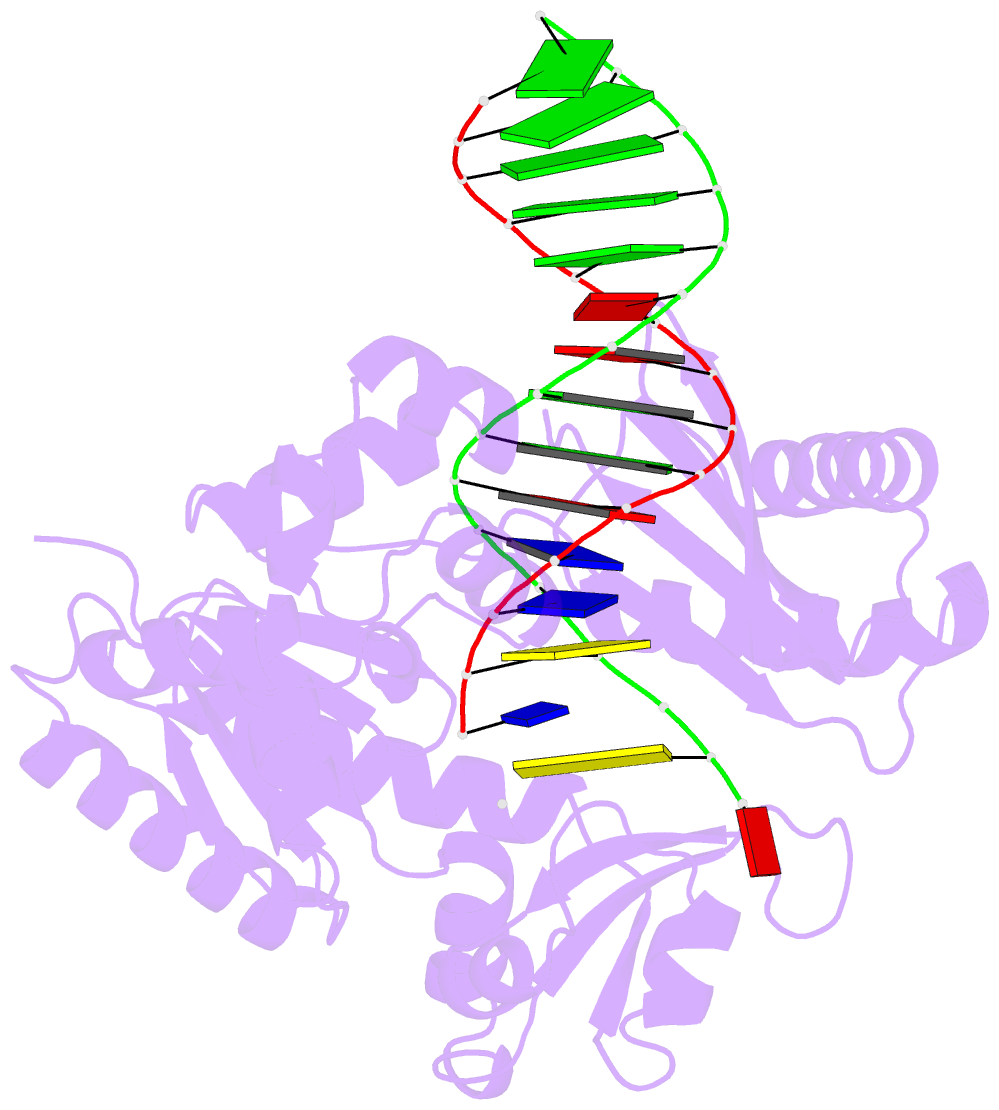
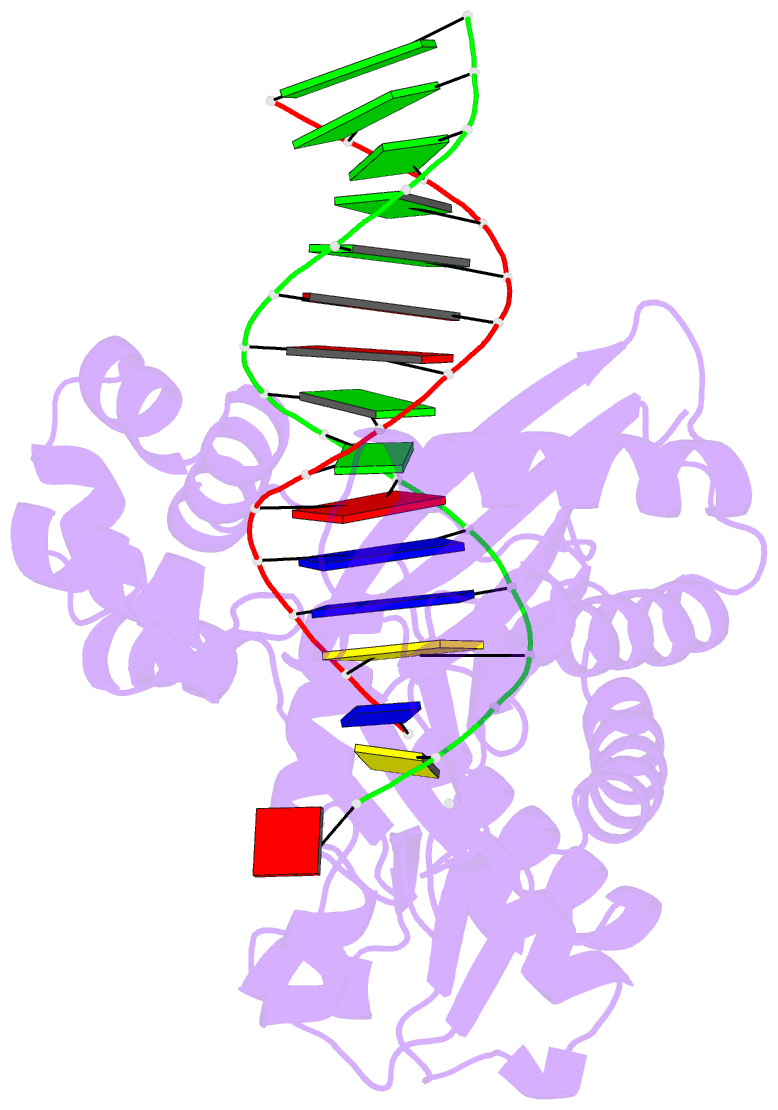
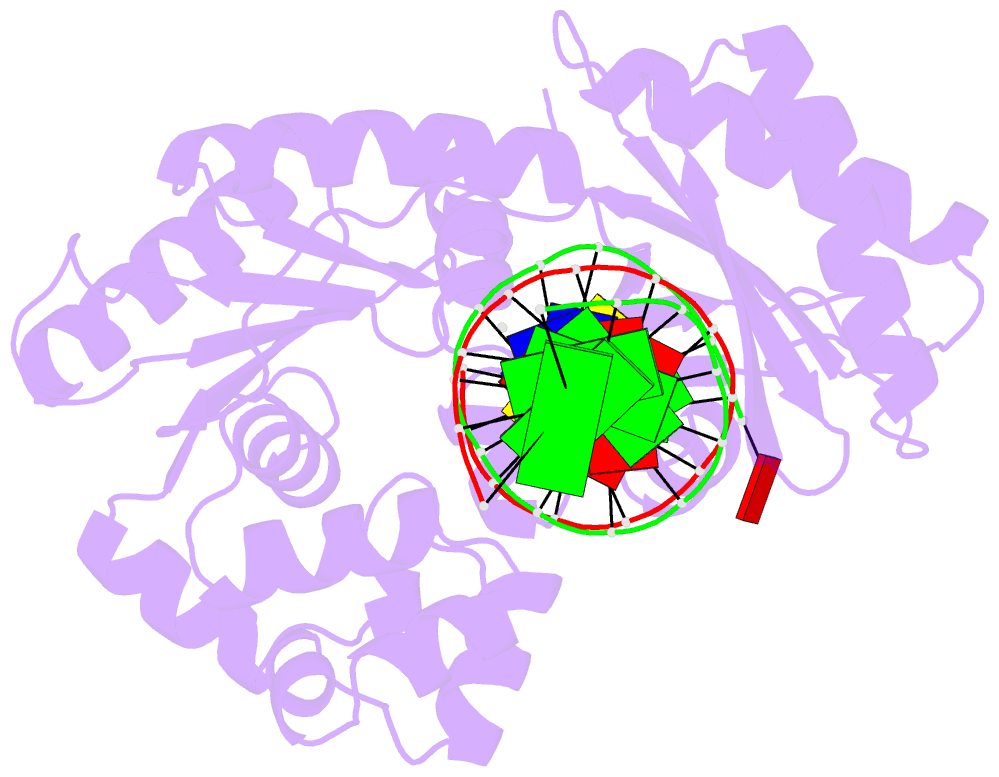
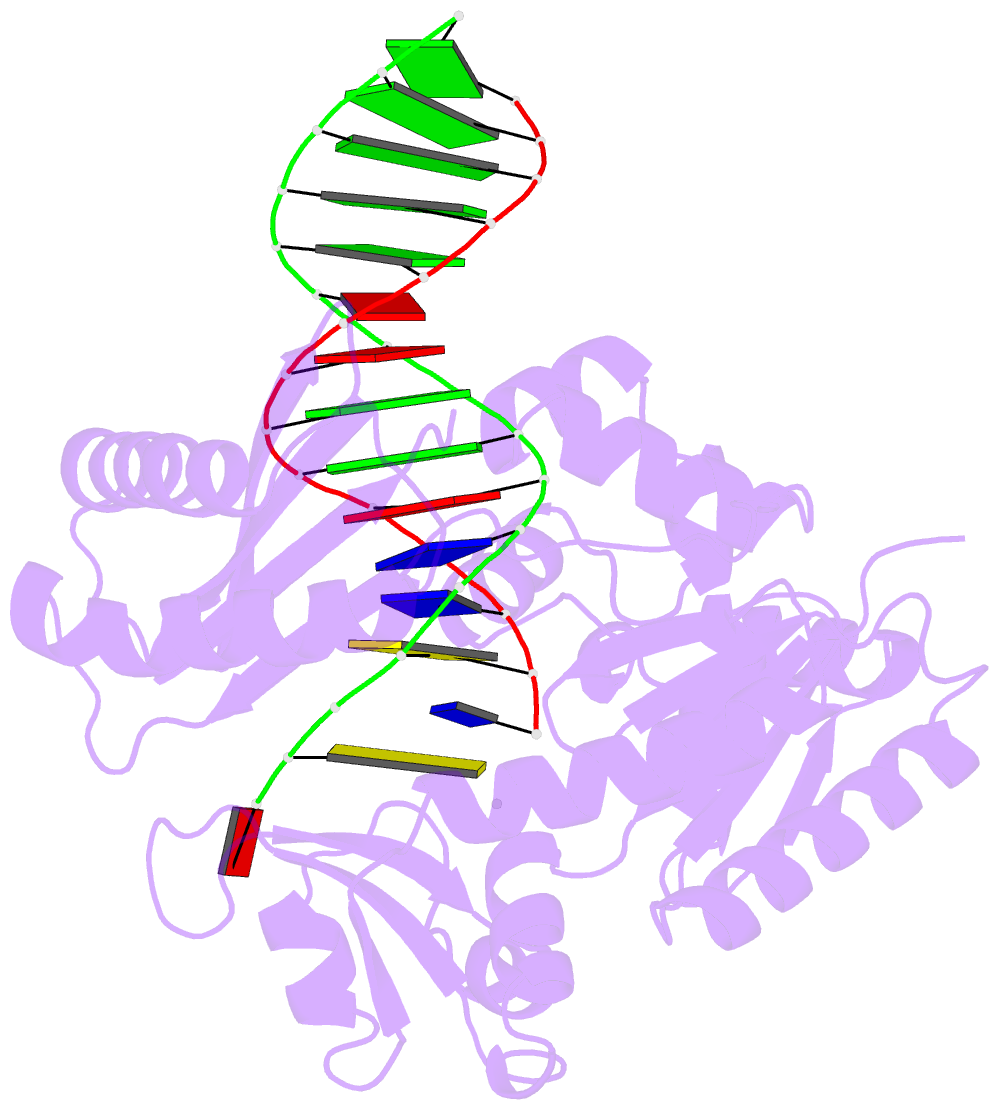
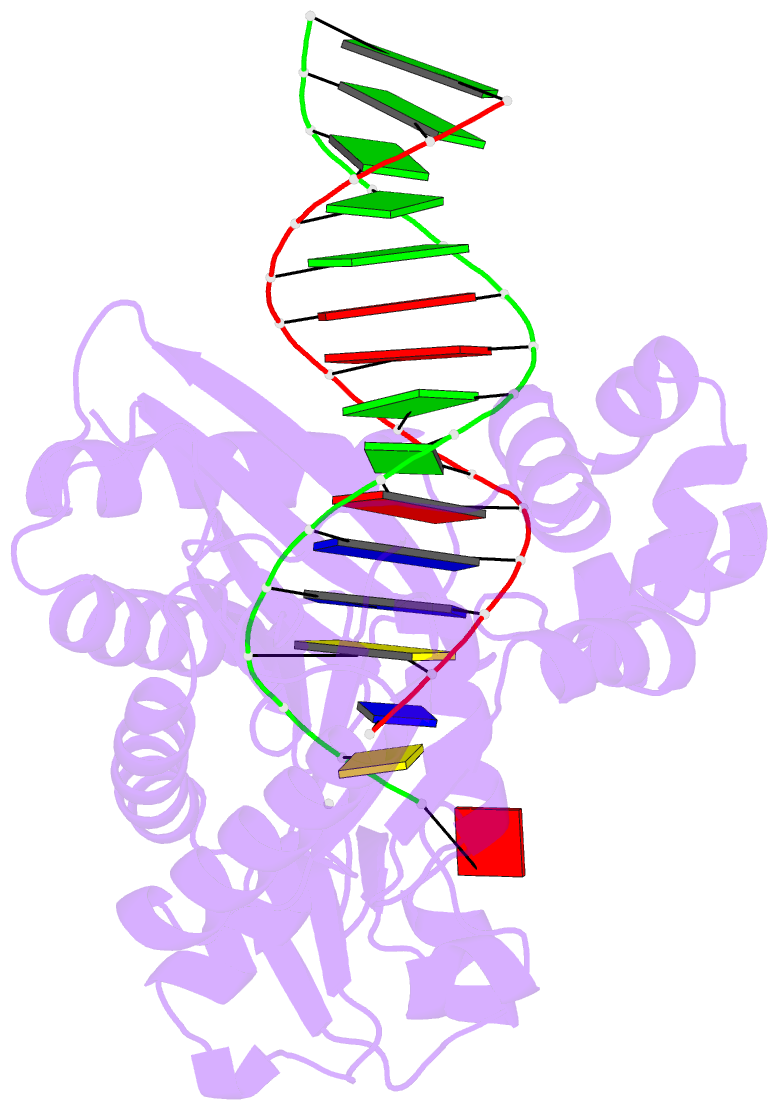
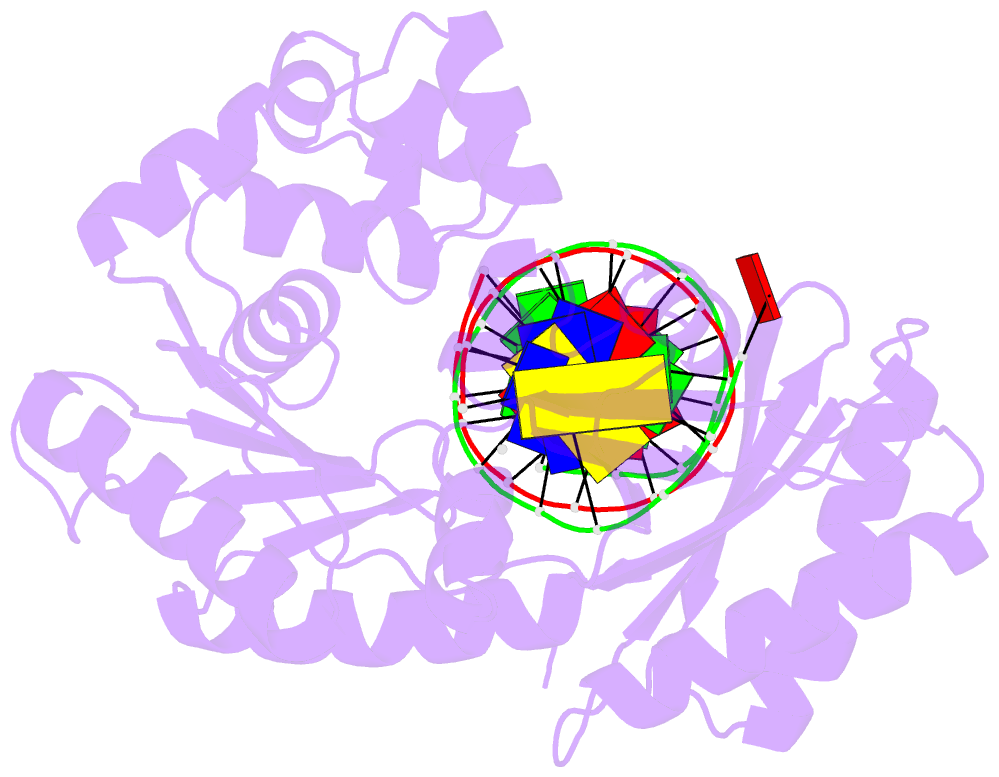
- The molecular basis of selectivity of nucleoside triphosphate incorporation opposite o6-benzylguanine by sulfolobus solfataricus DNA polymerase iv: steady-state and pre-steady-state kinetics and x- ray crystallography of correct and incorrect pairing
- Reference
- Eoff RL, Angel KC, Egli M, Guengerich FP (2007): "Molecular Basis of Selectivity of Nucleoside Triphosphate Incorporation Opposite O6-Benzylguanine by Sulfolobus Solfataricus DNA Polymerase Dpo4: Steady-State and Pre-Steady-State Kinetics and X-Ray Crystallography of Correct and Incorrect Pairing." J.Biol.Chem., 282, 13573. doi: 10.1074/JBC.M700656200.
- Abstract
- Previous work has shown that Sulfolobus solfataricus DNA polymerase Dpo4-catalyzed bypass of O(6)-methylguanine (O(6)-MeG) proceeds largely in an accurate but inefficient manner with a "wobble" base pairing between C and O(6)-MeG (Eoff, R. L., Irimia, A., Egli, M., and Guengerich, F. P. (2007) J. Biol. Chem. 282, 1456-1467). We considered here the bulky lesion O(6)-benzylguanine (O(6)-BzG) in DNA and catalysis by Dpo4. Mass spectrometry analysis of polymerization products revealed that the enzyme bypasses and extends across from O(6)-BzG, with C the major product ( approximately 70%) and some T and A ( approximately 15% each) incorporated opposite the lesion. Steady-state kinetic parameters indicated that Dpo4 was 7-, 5-, and 27-fold more efficient at C incorporation opposite O(6)-BzG than T, A, or G, respectively. In transient state kinetic analysis, the catalytic efficiency was decreased 62-fold for C incorporation opposite O(6)-BzG relative to unmodified DNA. Crystal structures reveal wobble pairing between C and O(6)-BzG. Pseudo-"Watson-Crick" pairing was observed between T and O(6)-BzG. Two other structures illustrate a possible mechanism for the accommodation of a +1 frameshift in the Dpo4 active site. The overall effect of O(6)-BzG is to decrease the efficiency of bypass by roughly an order of magnitude in every case except correct bypass, where the effect is not as pronounced. By comparison, Dpo4 is more accurate but no more efficient than model replicative polymerases, such as bacteriophage T7(-) DNA polymerase and human immunodeficiency virus-1 reverse transcriptase in the polymerization past O(6)-MeG and O(6)-BzG.