Summary information and primary citation
- PDB-id
- 2pjp; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- translation-RNA
- Method
- X-ray (2.3 Å)
- Summary
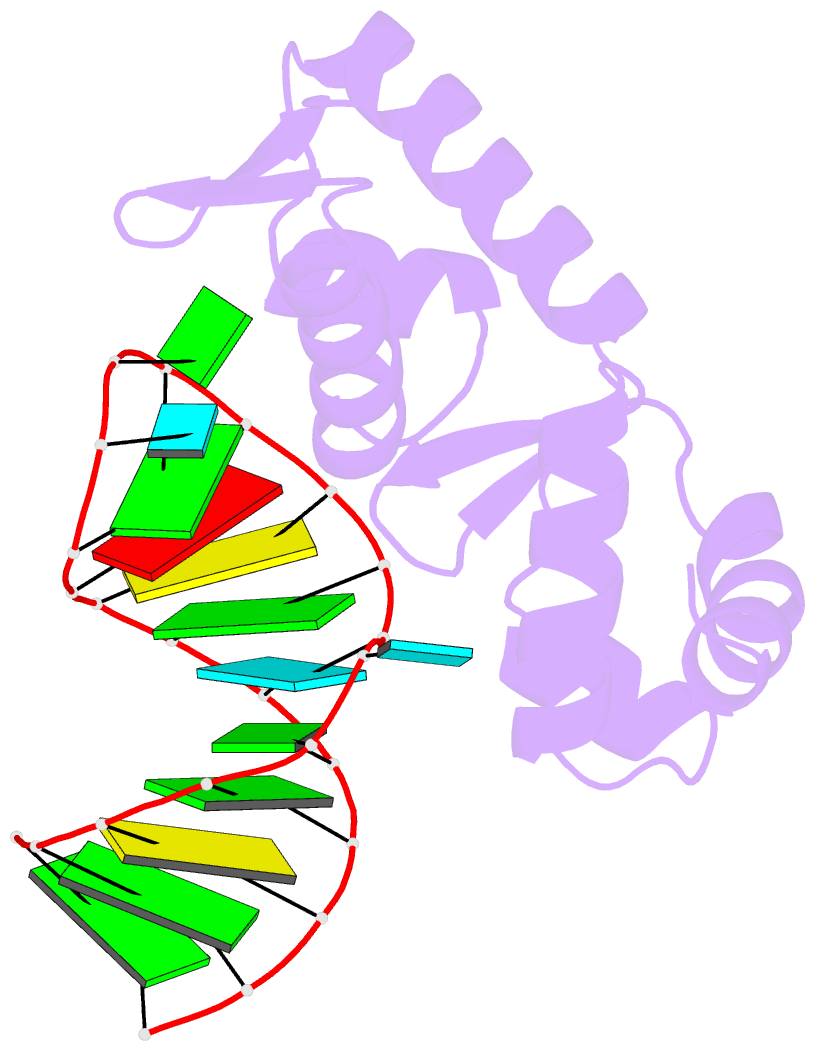
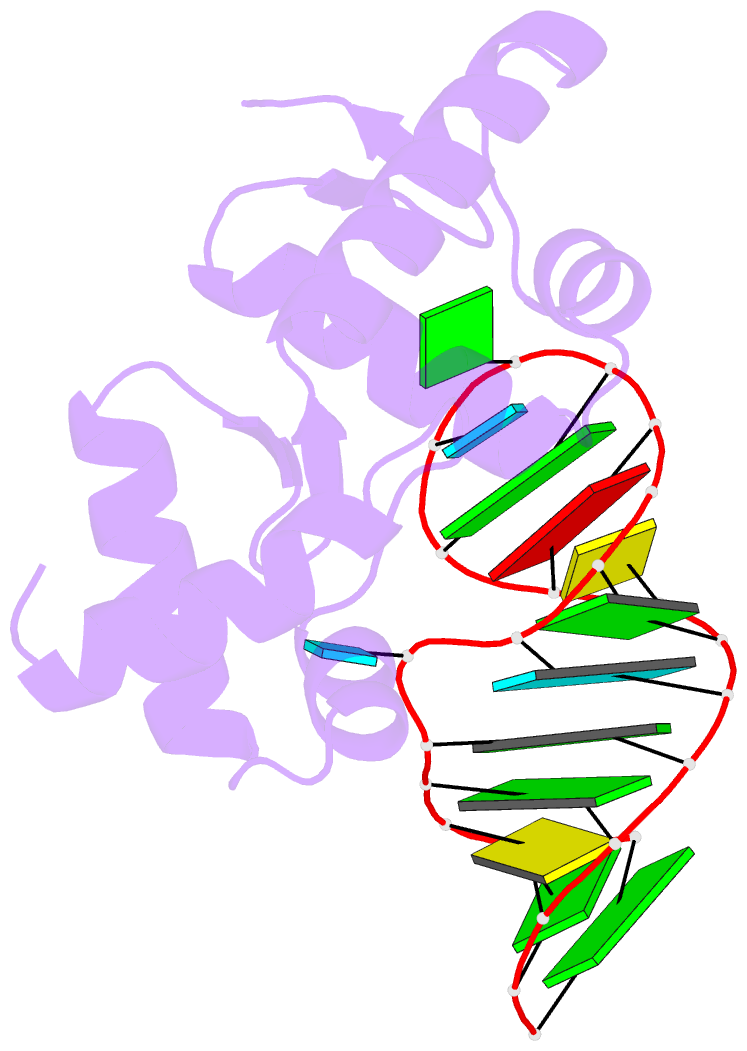
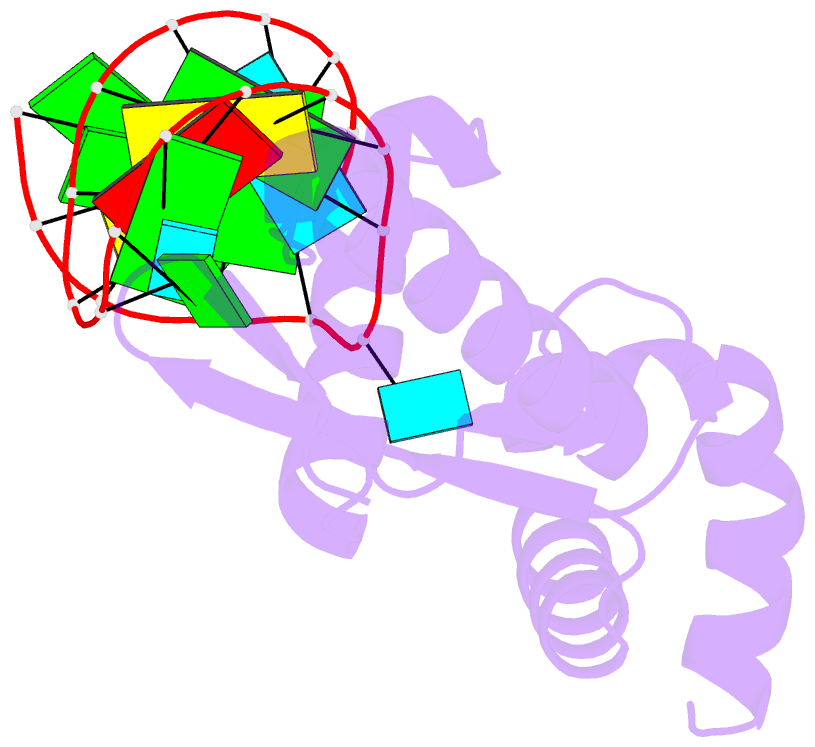
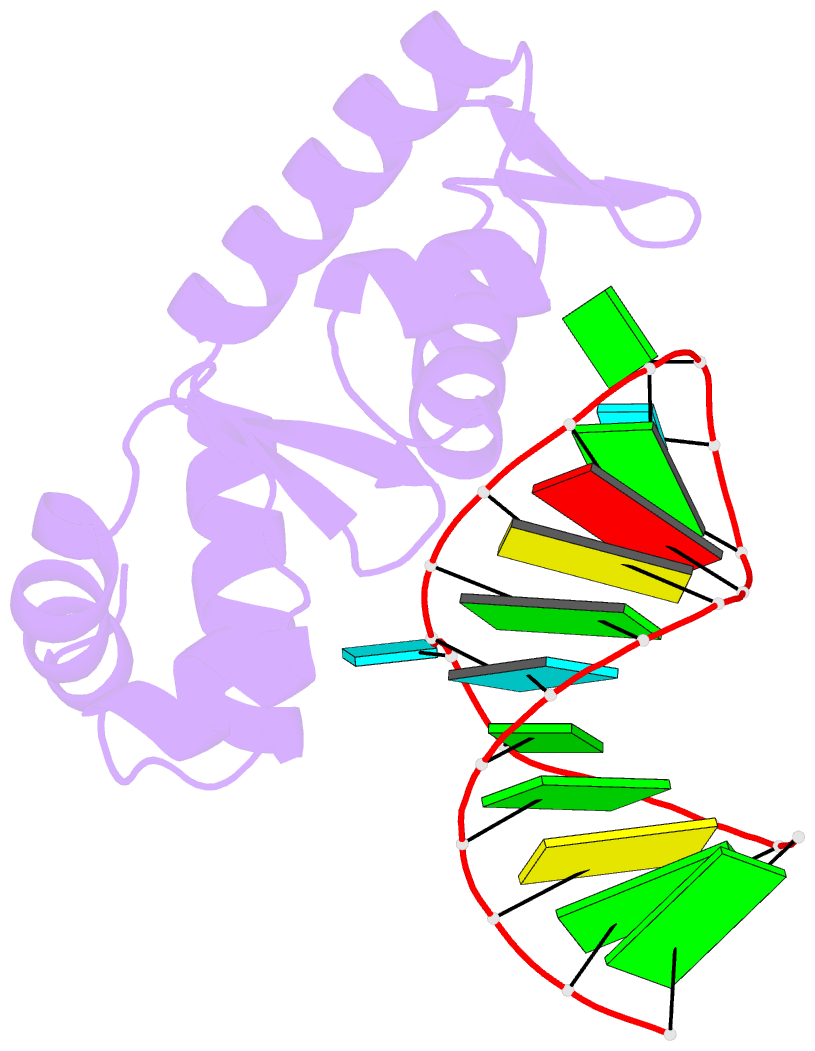
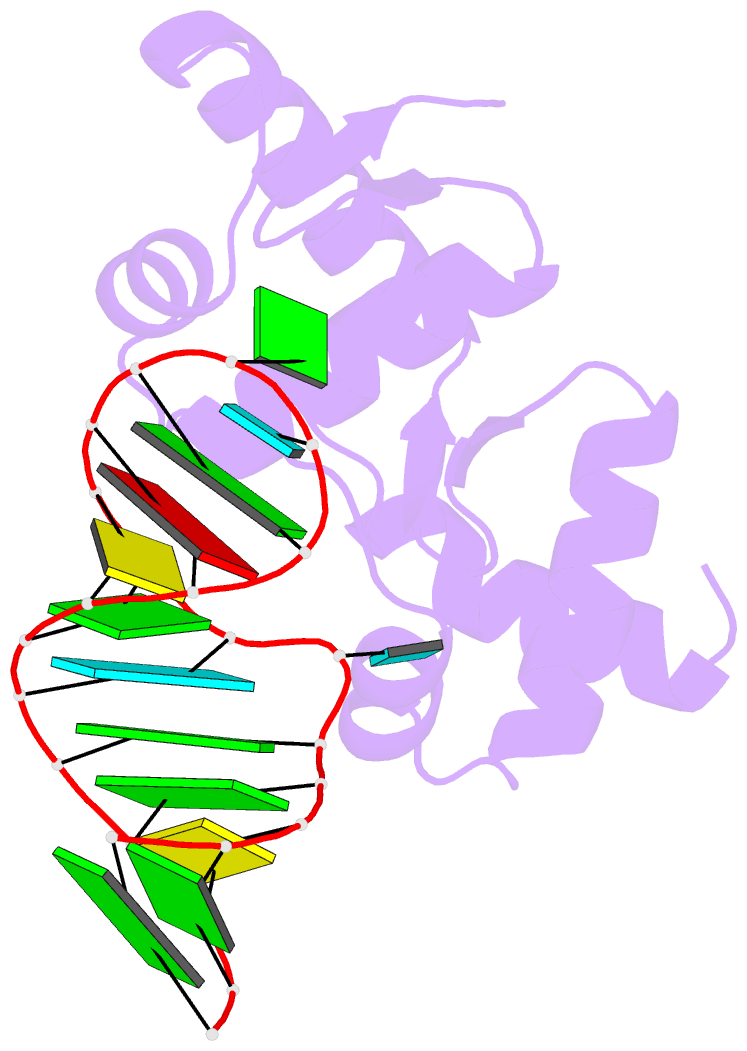
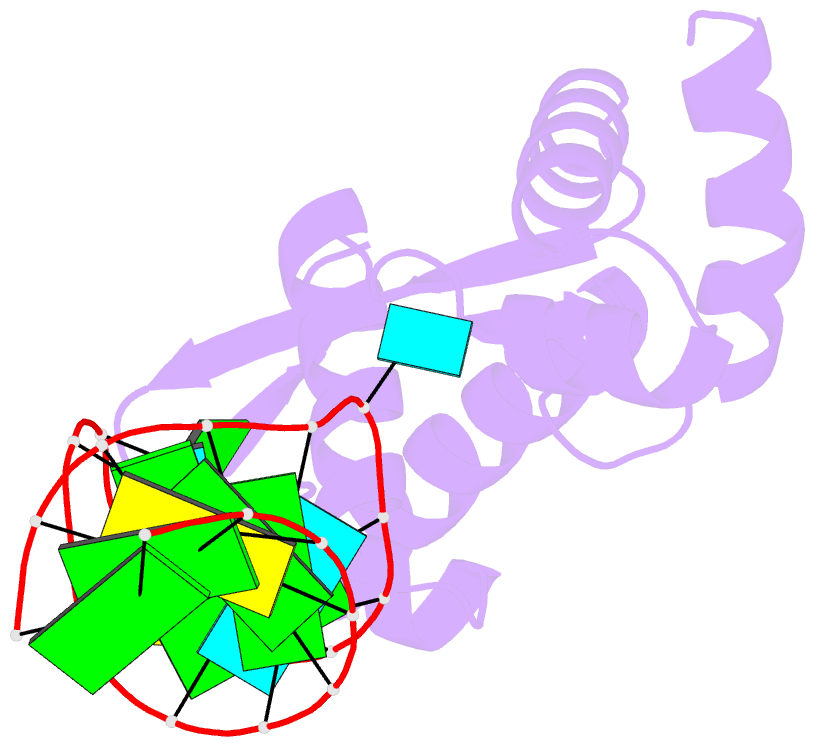
- Structure of the mrna-binding domain of elongation factor selb from e.coli in complex with secis RNA
- Reference
- Soler N, Fourmy D, Yoshizawa S (2007): "Structural insight into a molecular switch in tandem winged-helix motifs from elongation factor SelB." J.Mol.Biol., 370, 728-741. doi: 10.1016/j.jmb.2007.05.001.
- Abstract
- Elongation factor SelB is responsible for co-translational incorporation of selenocysteine (Sec) into proteins. The UGA stop codon is recoded as a Sec codon in the presence of a downstream mRNA hairpin. In prokaryotes, in addition to the EF-Tu-like N-terminal domains, a C-terminal extension containing four tandem winged-helix motifs (WH1-4) recognizes the mRNA hairpin. The 2.3-A resolution crystal structure of the Escherichia coli WH3/4 domains bound to mRNA with mutagenesis data reveal that the two WH motifs use the same structural elements to bind RNA. The structure together with the 2.6-A resolution structure of the WH1-4 domains from Moorella thermoacetica bound to RNA revealed that a salt bridge connecting WH2 to WH3 modules is disrupted upon mRNA binding. The results provide a structural basis for the molecular switch that may allow communication between tRNA and mRNA binding sites and illustrate how RNA acts as an activator of the switch. The structures show that tandem WH motifs not only provide an excellent scaffold for RNA binding but can also have an active role in the function of protein-RNA complexes.