Summary information and primary citation
- PDB-id
- 2zm5; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-RNA
- Method
- X-ray (2.55 Å)
- Summary
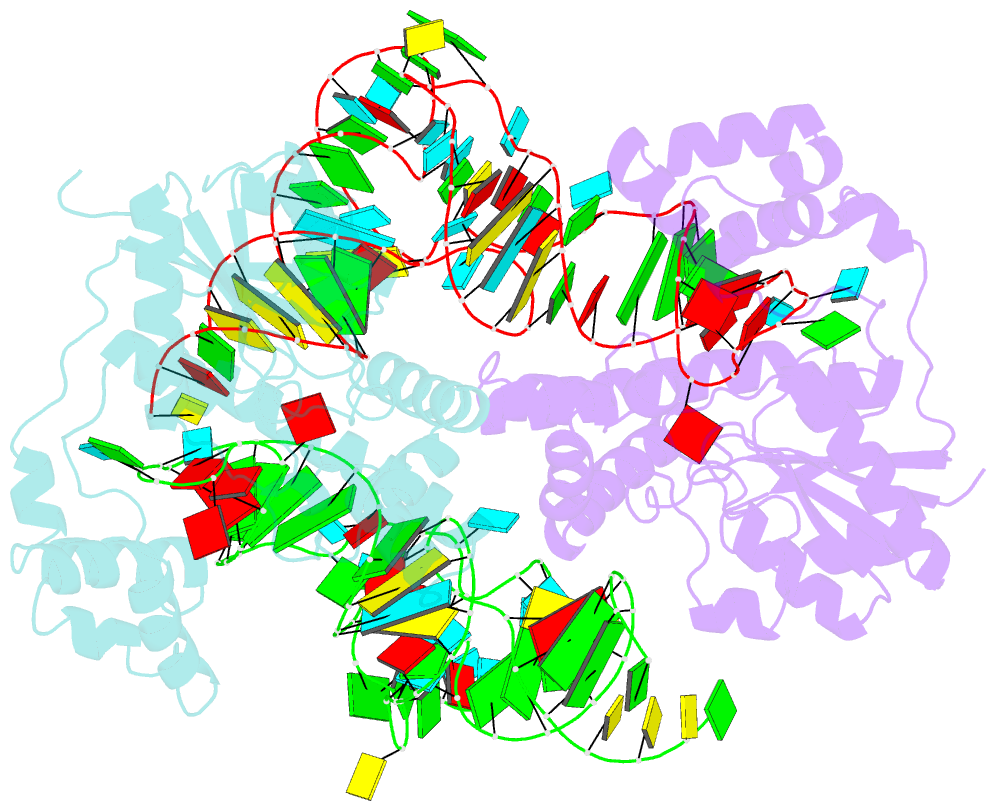
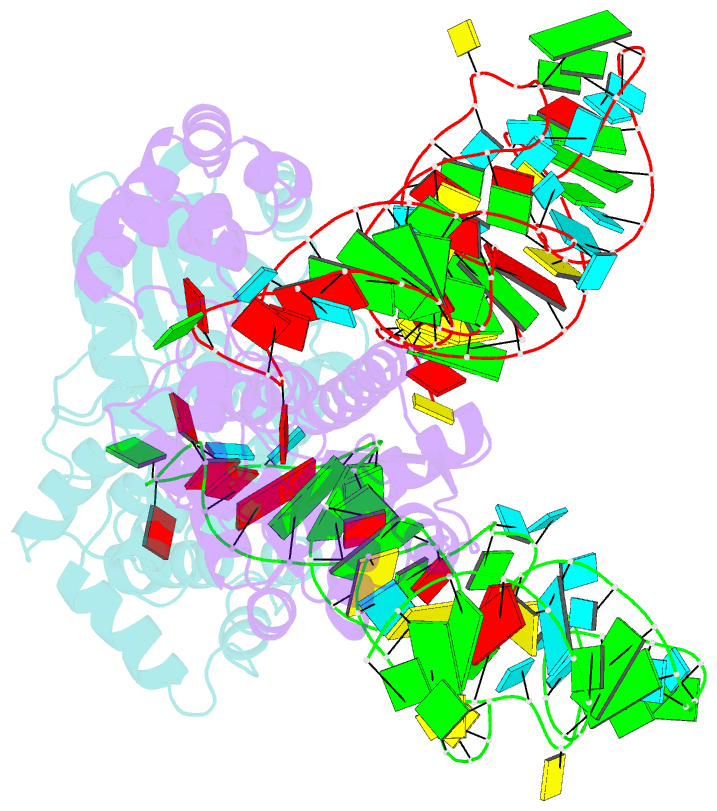
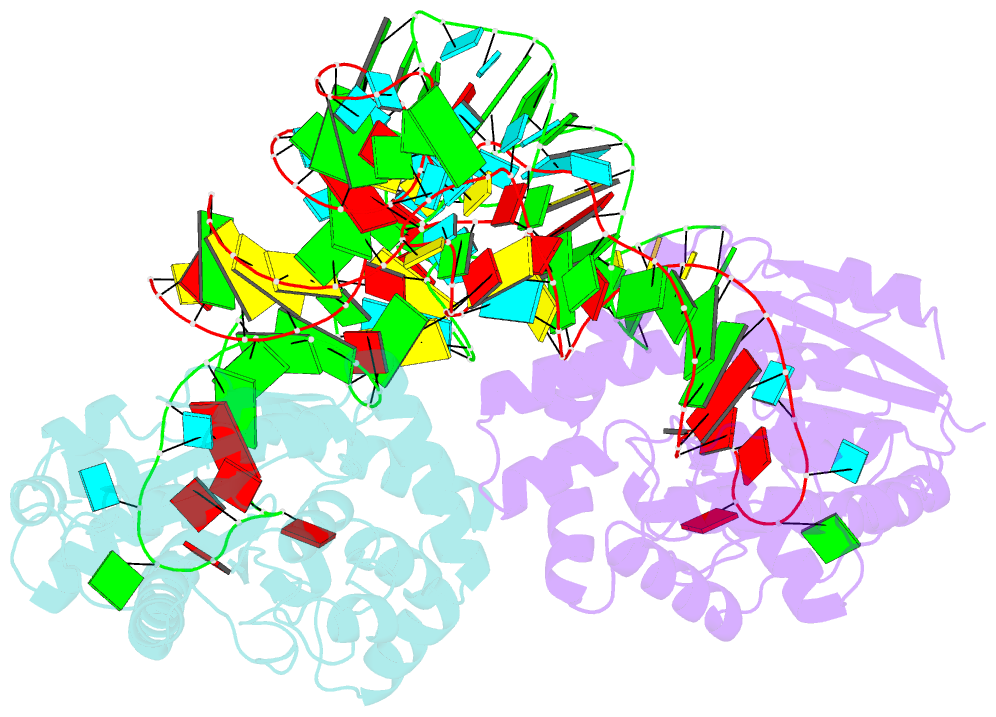
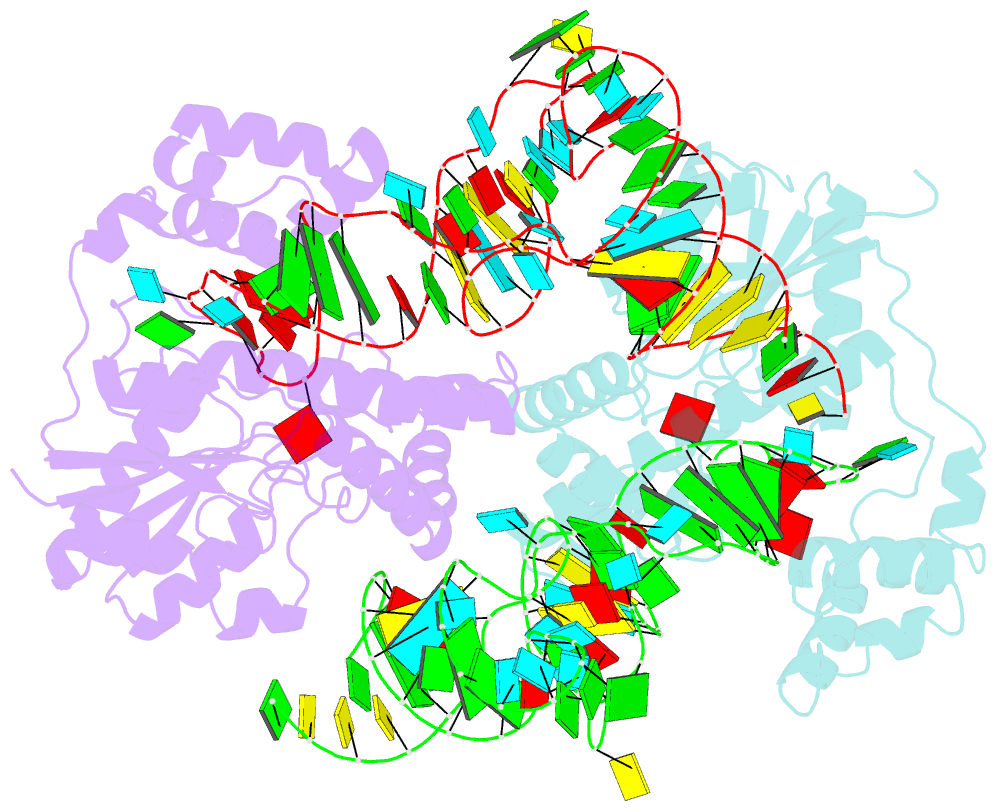
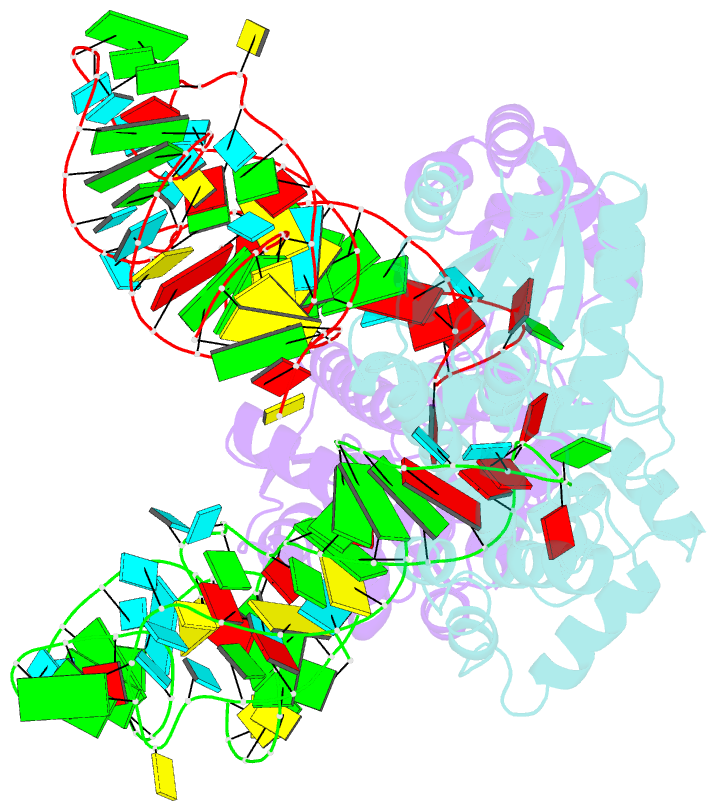
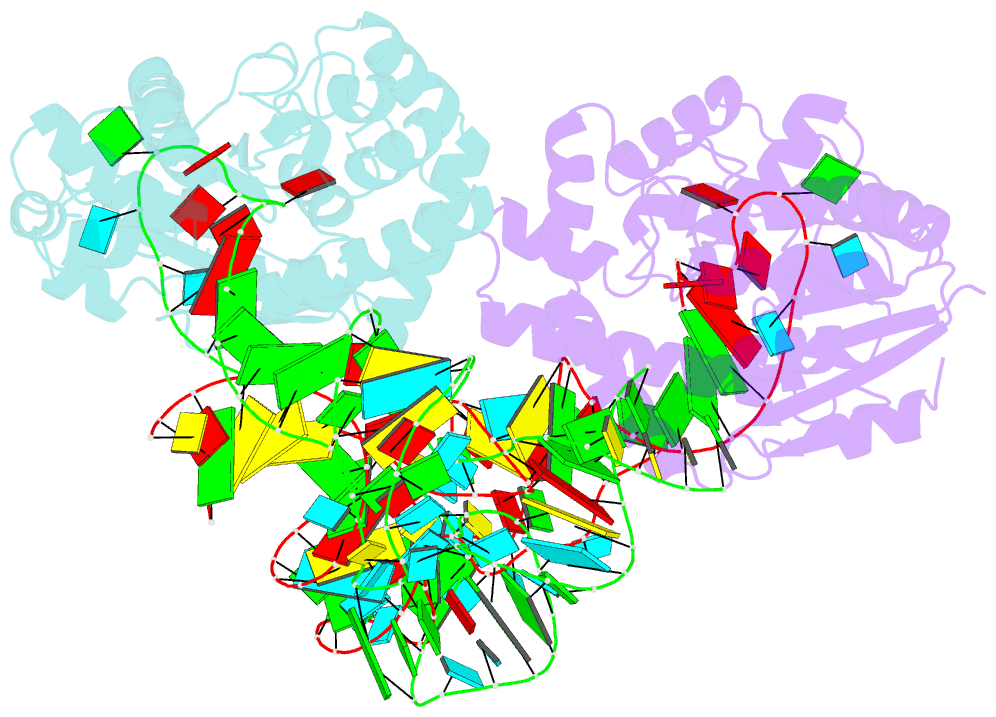
- Crystal structure of trna modification enzyme miaa in the complex with trna(phe)
- Reference
- Chimnaronk S, Forouhar F, Sakai J, Yao M, Tron CM, Atta M, Fontecave M, Hunt JF, Tanaka I (2009): "Snapshots of dynamics in synthesizing N(6)-isopentenyladenosine at the tRNA anticodon." Biochemistry, 48, 5057-5065. doi: 10.1021/bi900337d.
- Abstract
- Bacterial and eukaryotic tRNAs that decode codons starting with uridine have a hydrophobically hypermodified adenosine at position 37 (A(37)) adjacent to the 3'-end of the anticodon, which is essential for efficient and highly accurate protein translation by the ribosome. However, it remains unclear as to how the corresponding tRNAs are selected to be modified by alkylation at the correct position of the adenosine base. We have determined a series of crystal structures of bacterial tRNA isopentenyltransferase (MiaA) in apo- and tRNA-bound forms, which completely render snapshots of substrate selections during the modification of RNA. A compact evolutionary inserted domain (herein swinging domain) in MiaA that exhibits as a highly mobile entity moves around the catalytic domain as likely to reach and trap the tRNA substrate. Thereby, MiaA clamps the anticodon stem loop of the tRNA substrate between the catalytic and swinging domains, where the two conserved elongated residues from the swinging domain pinch the two flanking A(36) and A(38) together to squeeze out A(37) into the reaction tunnel. The site-specific isopentenylation of RNA is thus ensured by a characteristic pinch-and-flip mechanism and by a reaction tunnel to confine the substrate selection. Furthermore, combining information from soaking experiments with structural comparisons, we propose a mechanism for the ordered substrate binding of MiaA.