Summary information and primary citation
- PDB-id
- 3g73; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription-DNA
- Method
- X-ray (2.21 Å)
- Summary
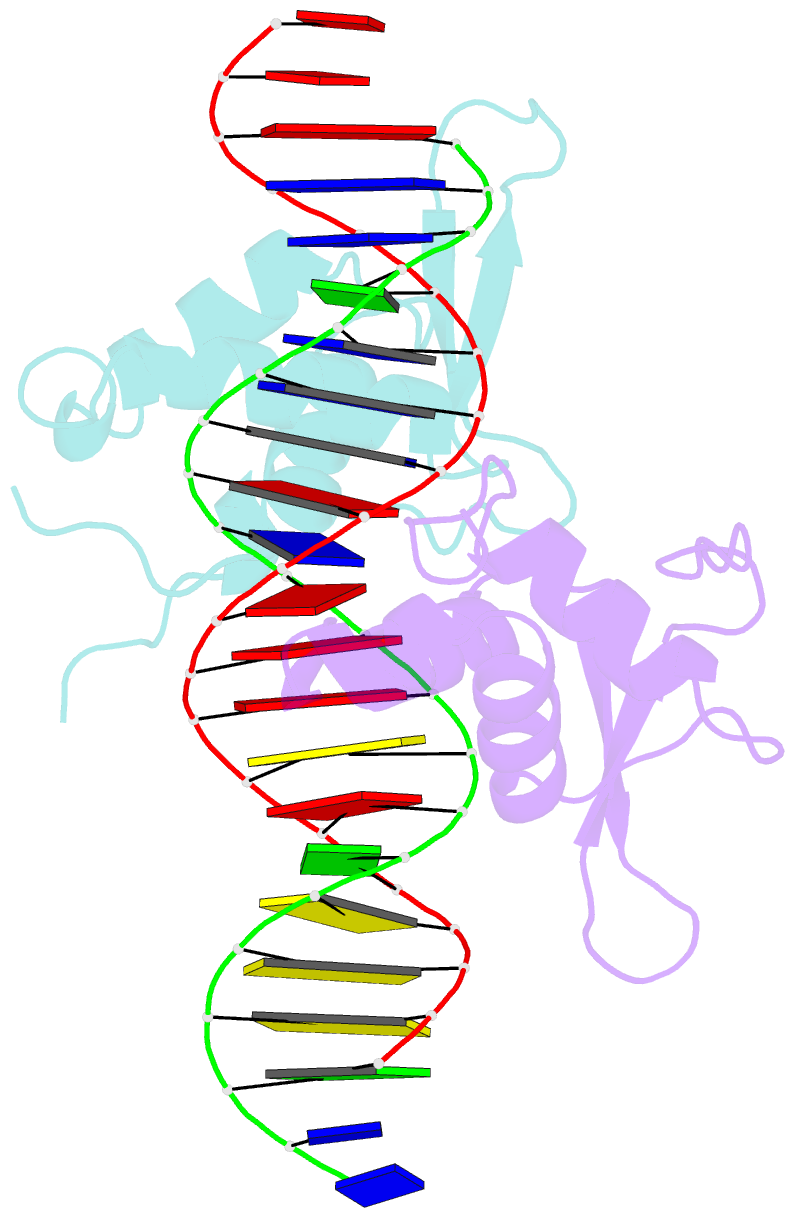
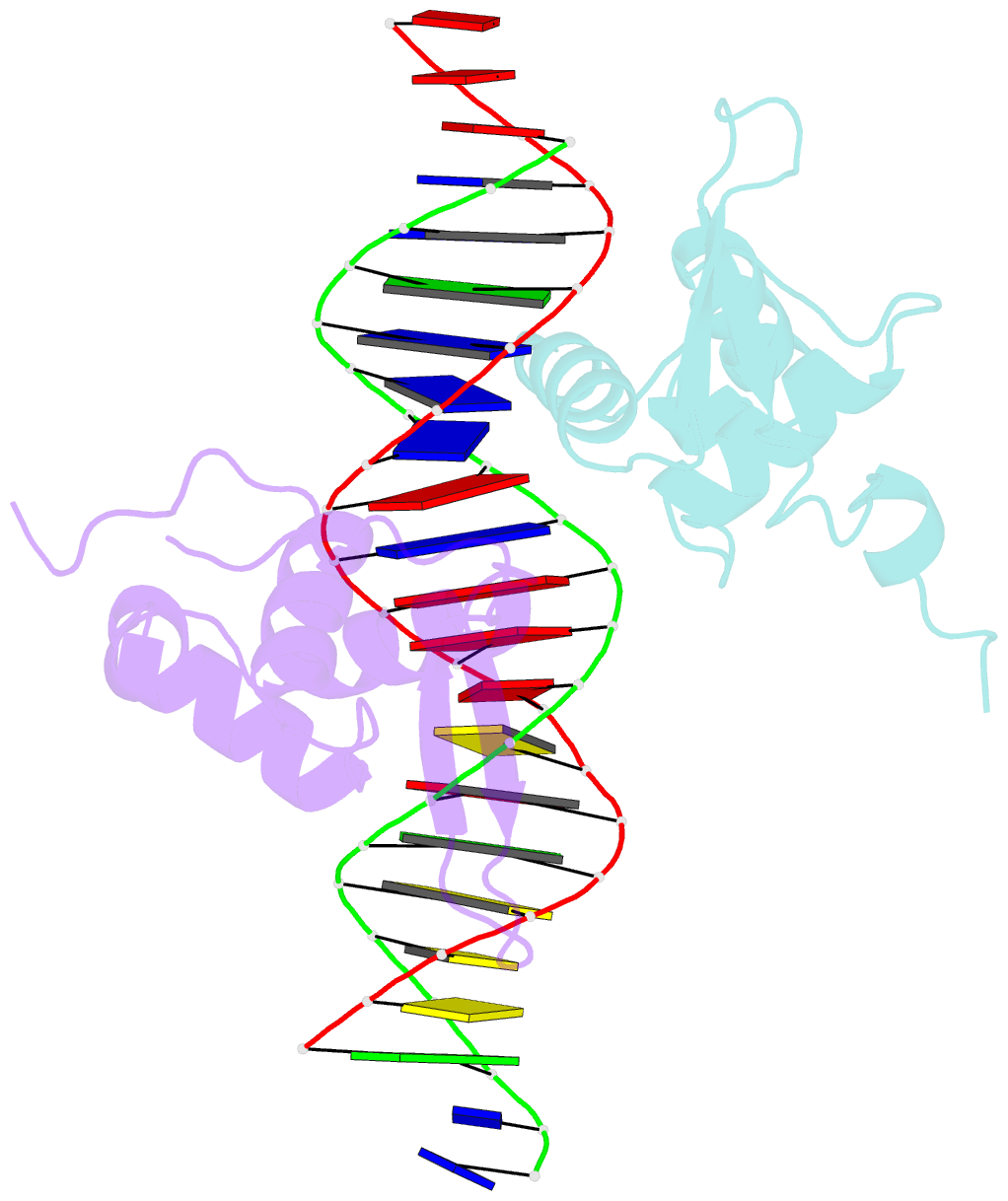
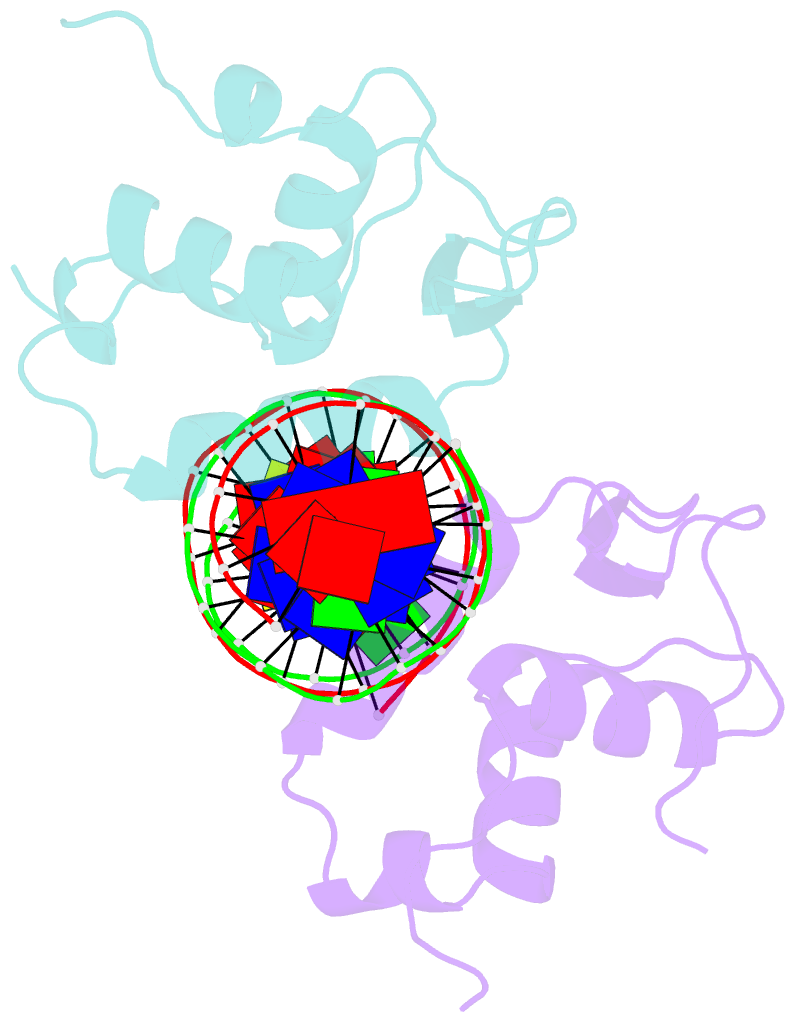
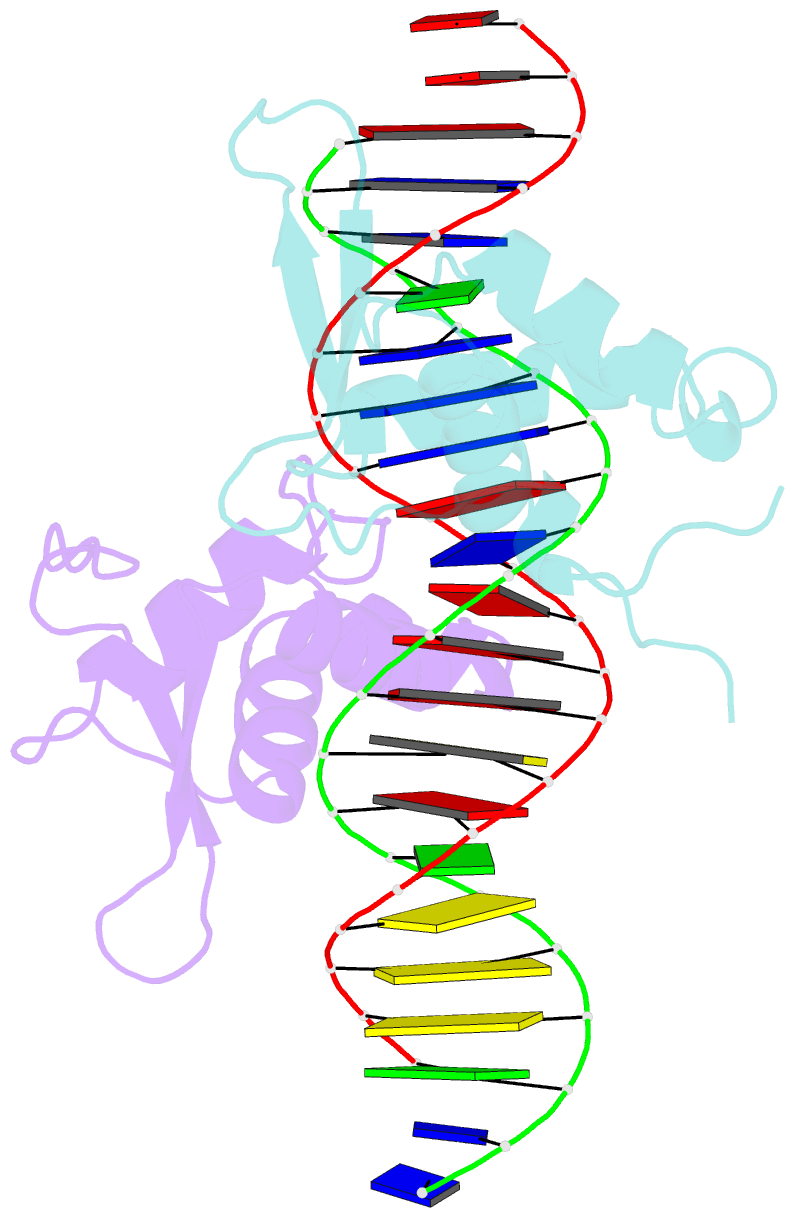
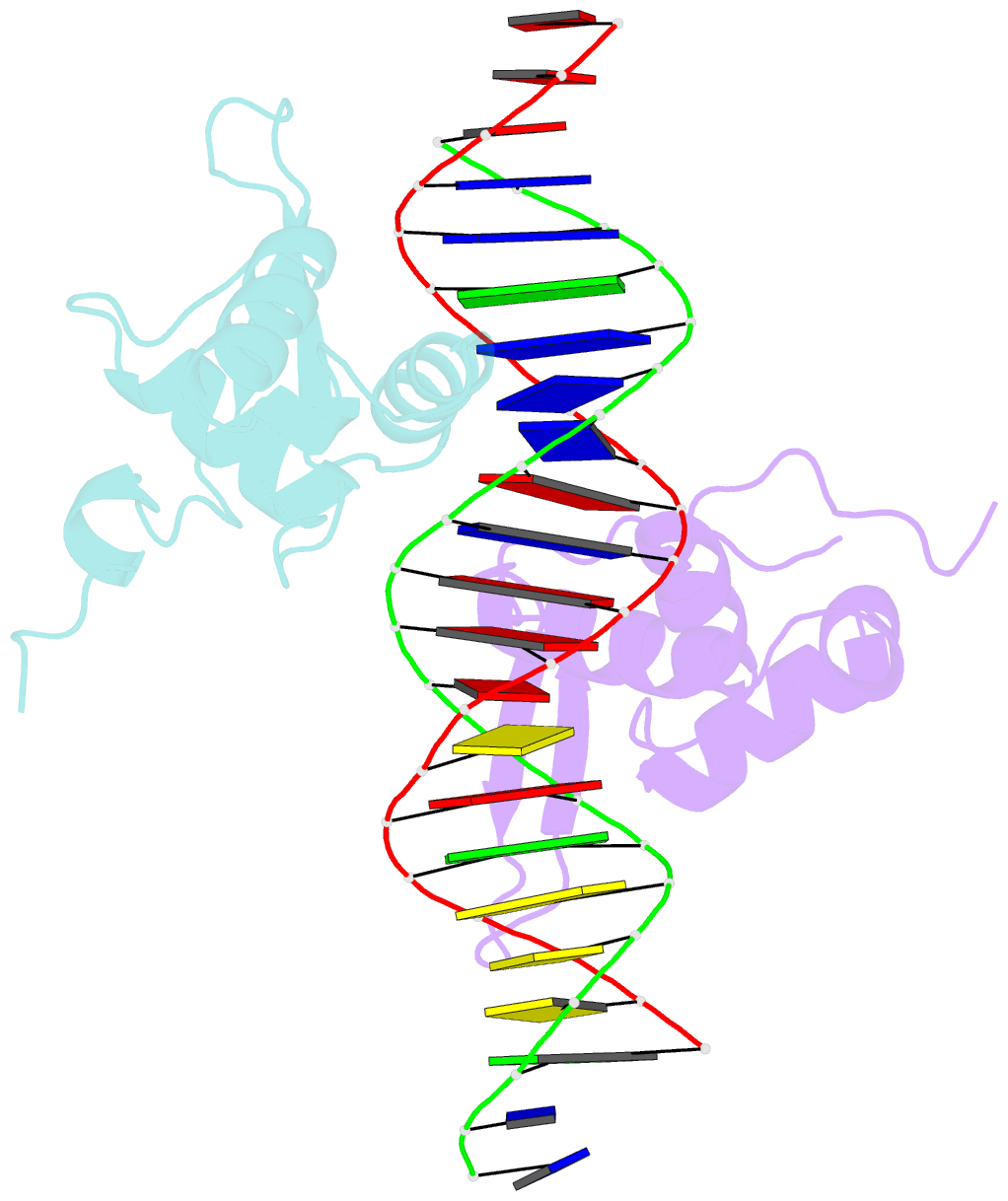
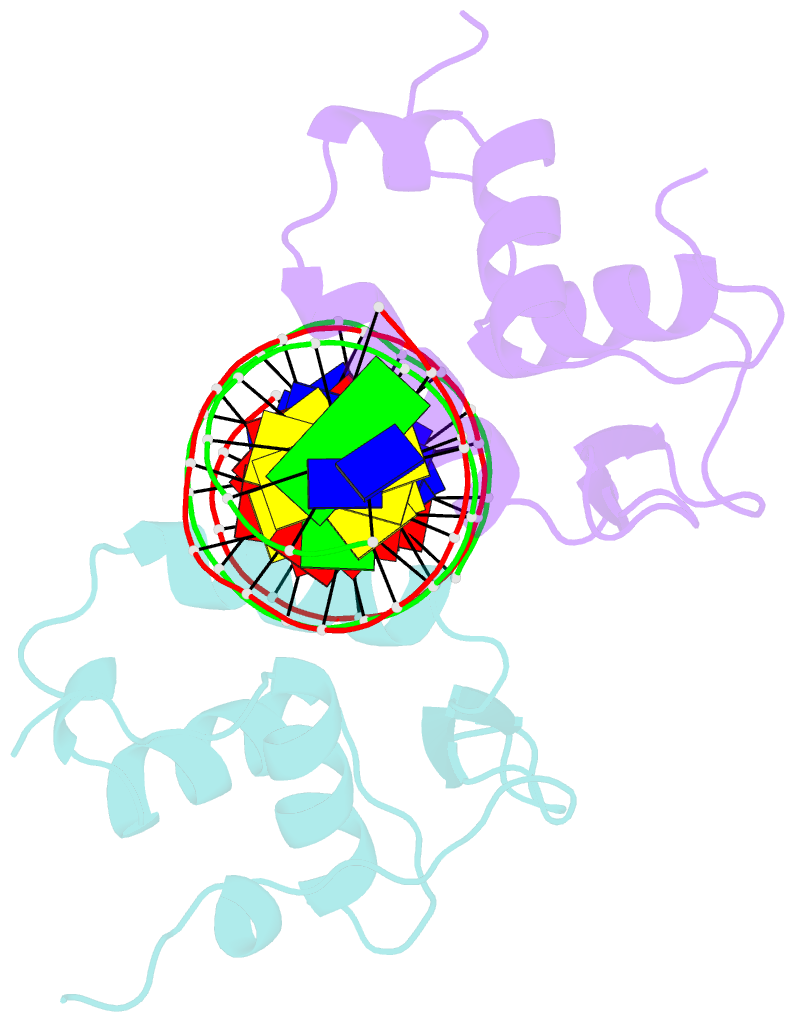
- Structure of the foxm1 DNA binding
- Reference
- Littler DR, Alvarez-Fernandez M, Stein A, Hibbert RG, Heidebrecht T, Aloy P, Medema RH, Perrakis A (2010): "Structure of the FoxM1 DNA-recognition domain bound to a promoter sequence." Nucleic Acids Res. doi: 10.1093/nar/gkq194.
- Abstract
- FoxM1 is a member of the Forkhead family of transcription factors and is implicated in inducing cell proliferation and some forms of tumorigenesis. It binds promoter regions with a preference for tandem repeats of a consensus 'TAAACA' recognition sequence. The affinity of the isolated FoxM1 DNA-binding domain for this site is in the micromolar range, lower than observed for other Forkhead proteins. To explain these FoxM1 features, we determined the crystal structure of its DNA-binding domain in complex with a tandem recognition sequence. FoxM1 adopts the winged-helix fold, typical of the Forkhead family. Neither 'wing' of the fold however, makes significant contacts with the DNA, while the second, C-terminal, wing adopts an unusual ordered conformation across the back of the molecule. The lack of standard DNA-'wing' interactions may be a reason for FoxM1's relatively low affinity. The role of the 'wings' is possibly undertaken by other FoxM1 regions outside the DBD, that could interact with the target DNA directly or mediate interactions with other binding partners. Finally, we were unable to show a clear preference for tandem consensus site recognition in DNA-binding, transcription activation or bioinformatics analysis; FoxM1's moniker, 'Trident', is not supported by our data.