Summary information and primary citation
- PDB-id
- 3jsm; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-DNA complex
- Method
- X-ray (3.0 Å)
- Summary
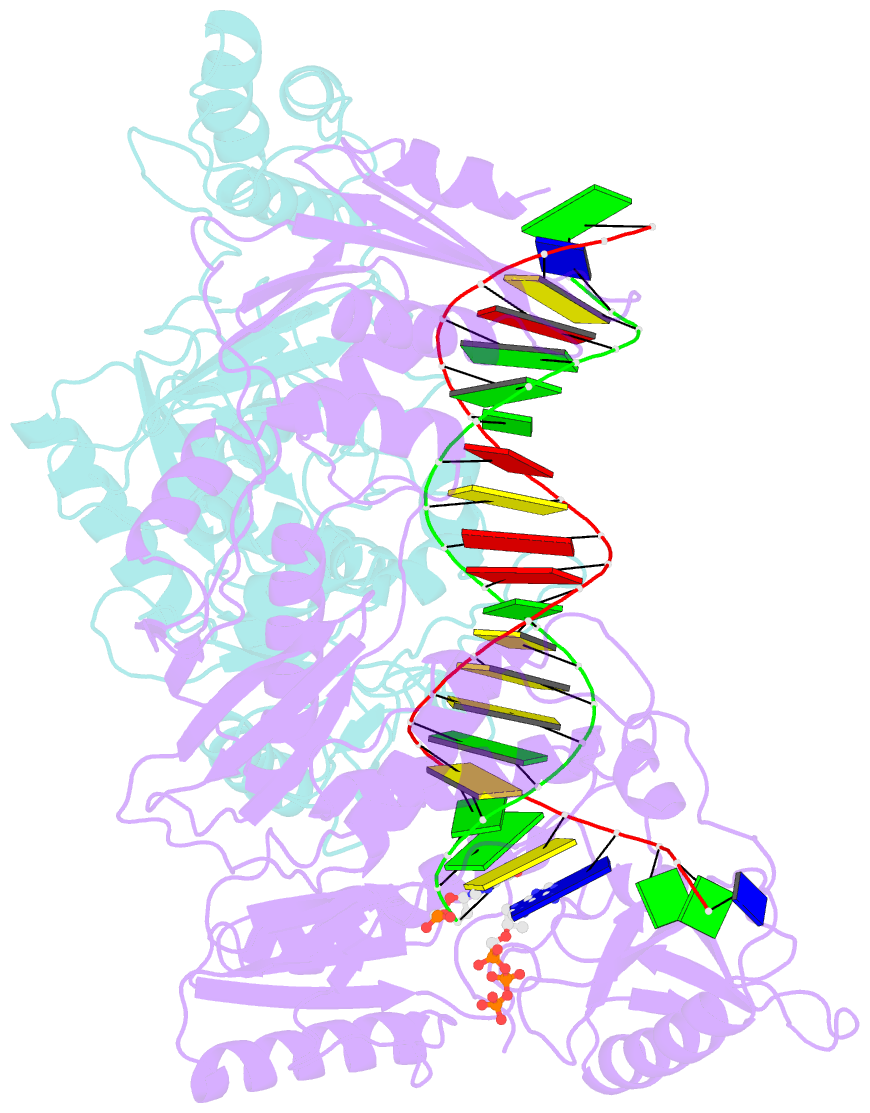
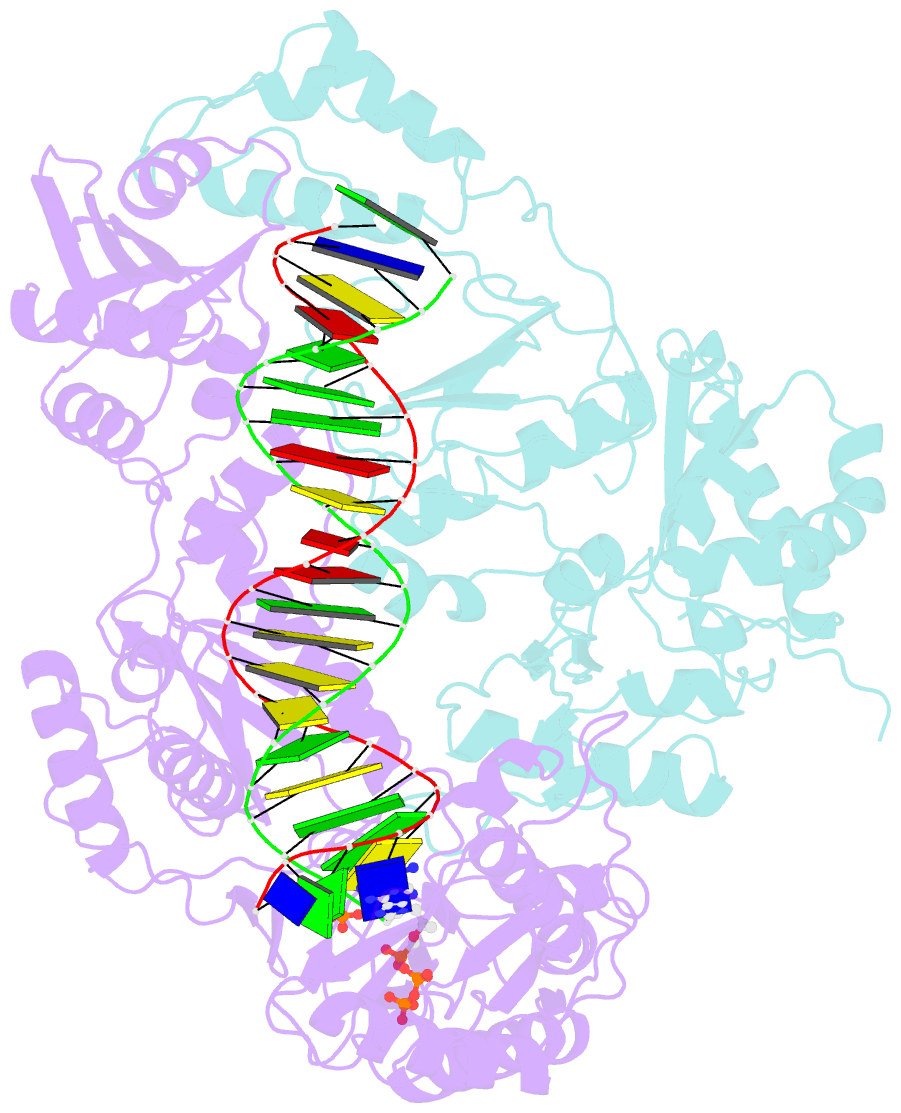
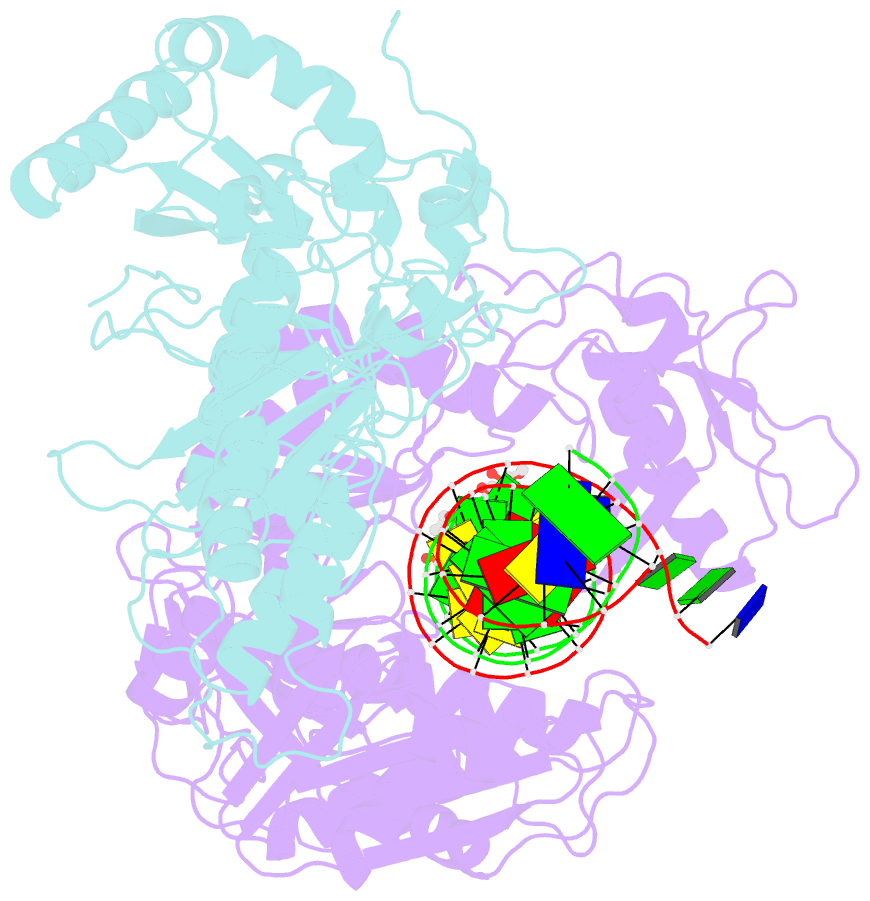
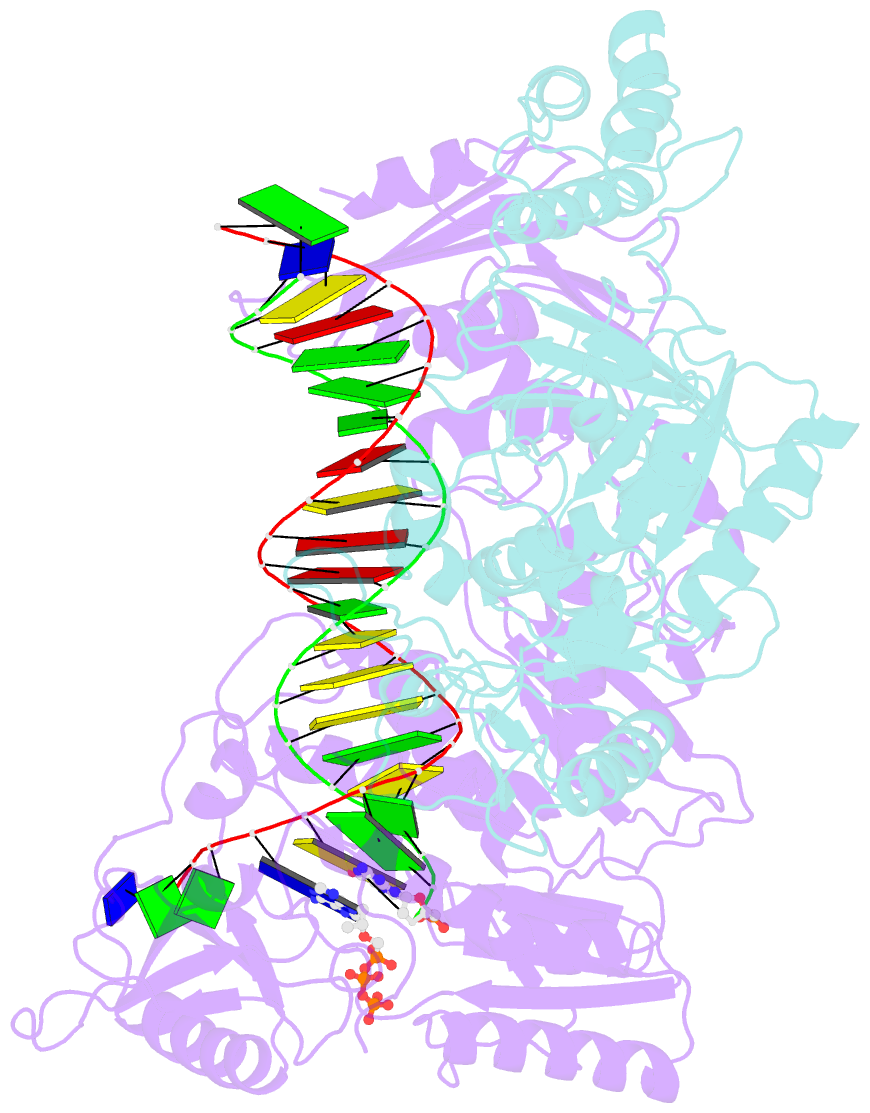
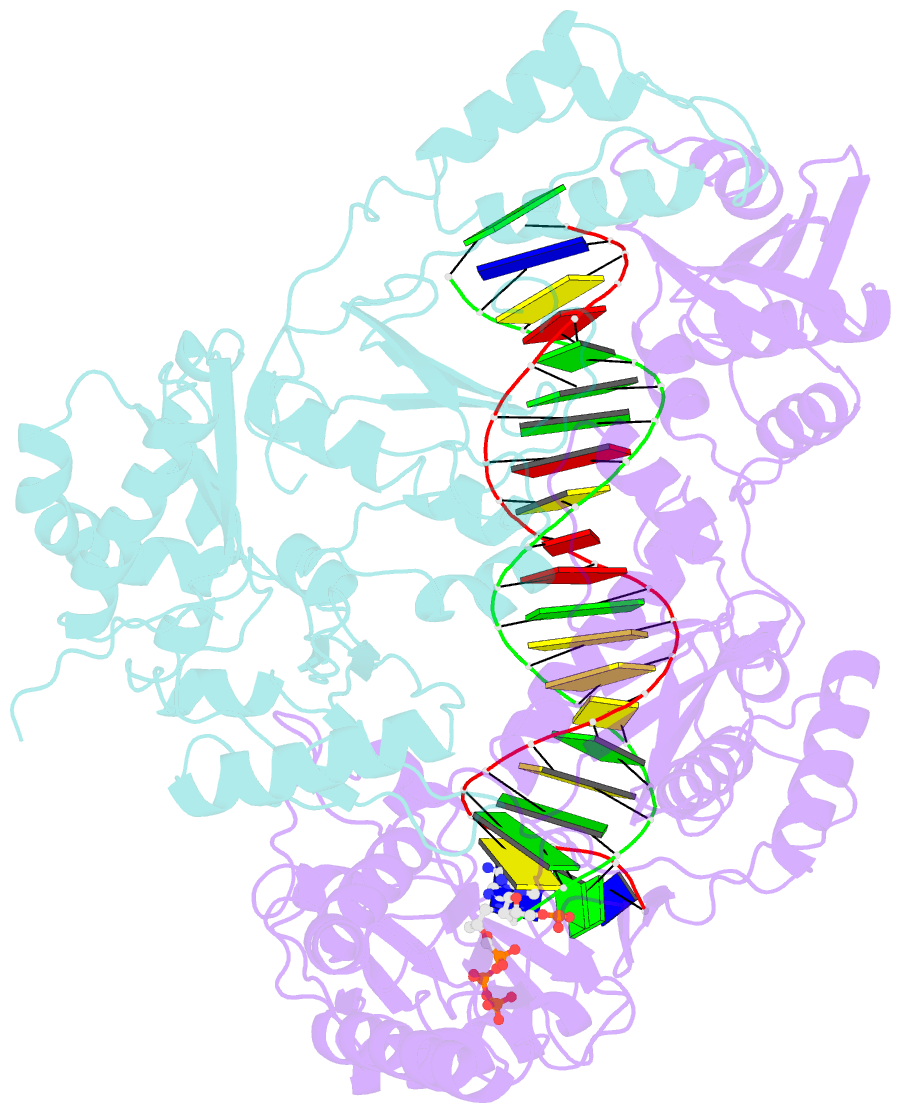
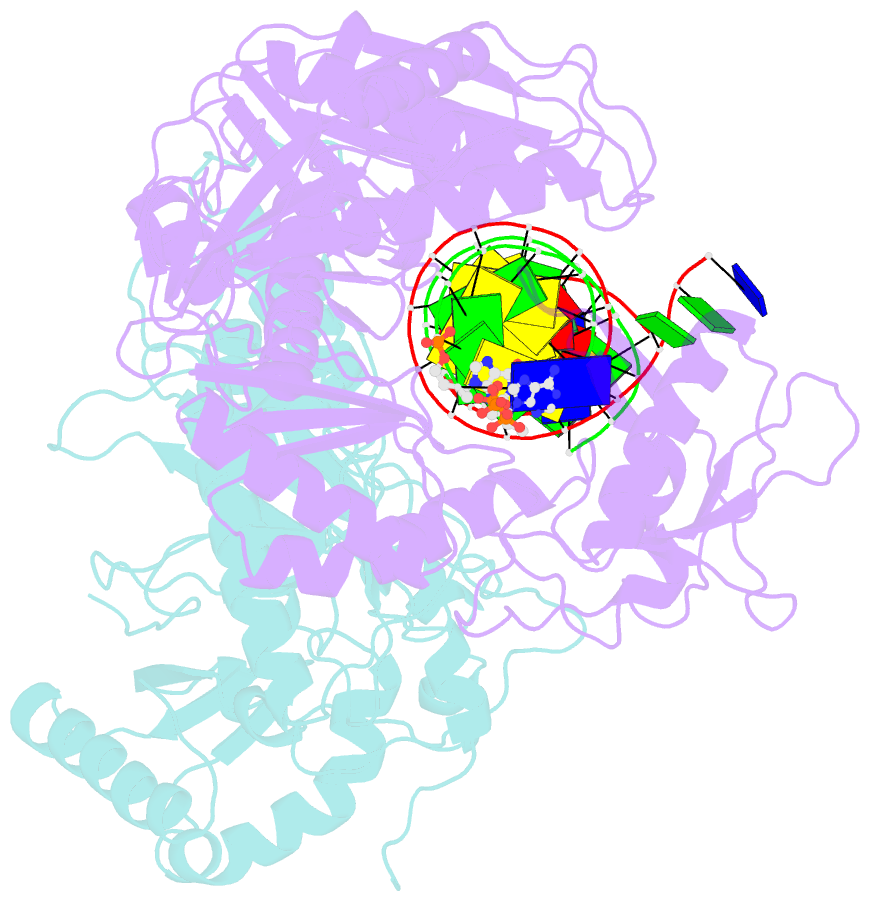
- K65r mutant hiv-1 reverse transcriptase cross-linked to ds-DNA and complexed with tenofovir-diphosphate as the incoming nucleotide substrate
- Reference
- Das K, Bandwar RP, White KL, Feng JY, Sarafianos SG, Tuske S, Tu X, Clark AD, Boyer PL, Hou X, Gaffney BL, Jones RA, Miller MD, Hughes SH, Arnold E (2009): "Structural basis for the role of the K65r mutation in HIV-1 reverse transcriptase polymerization, excision antagonism, and tenofovir resistance." J.Biol.Chem., 284, 35092-35100. doi: 10.1074/jbc.M109.022525.
- Abstract
- K65R is a primary reverse transcriptase (RT) mutation selected in human immunodeficiency virus type 1-infected patients taking antiretroviral regimens containing tenofovir disoproxil fumarate or other nucleoside analog RT drugs. We determined the crystal structures of K65R mutant RT cross-linked to double-stranded DNA and in complexes with tenofovir diphosphate (TFV-DP) or dATP. The crystals permit substitution of TFV-DP with dATP at the dNTP-binding site. The guanidinium planes of the arginines K65R and Arg(72) were stacked to form a molecular platform that restricts the conformational adaptability of both of the residues, which explains the negative effects of the K65R mutation on nucleotide incorporation and on excision. Furthermore, the guanidinium planes of K65R and Arg(72) were stacked in two different rotameric conformations in TFV-DP- and dATP-bound structures that may help explain how K65R RT discriminates the drug from substrates. These K65R-mediated effects on RT structure and function help us to visualize the complex interaction with other key nucleotide RT drug resistance mutations, such as M184V, L74V, and thymidine analog resistance mutations.