Summary information and primary citation
- PDB-id
- 3kmq; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-RNA
- Method
- X-ray (2.11 Å)
- Summary
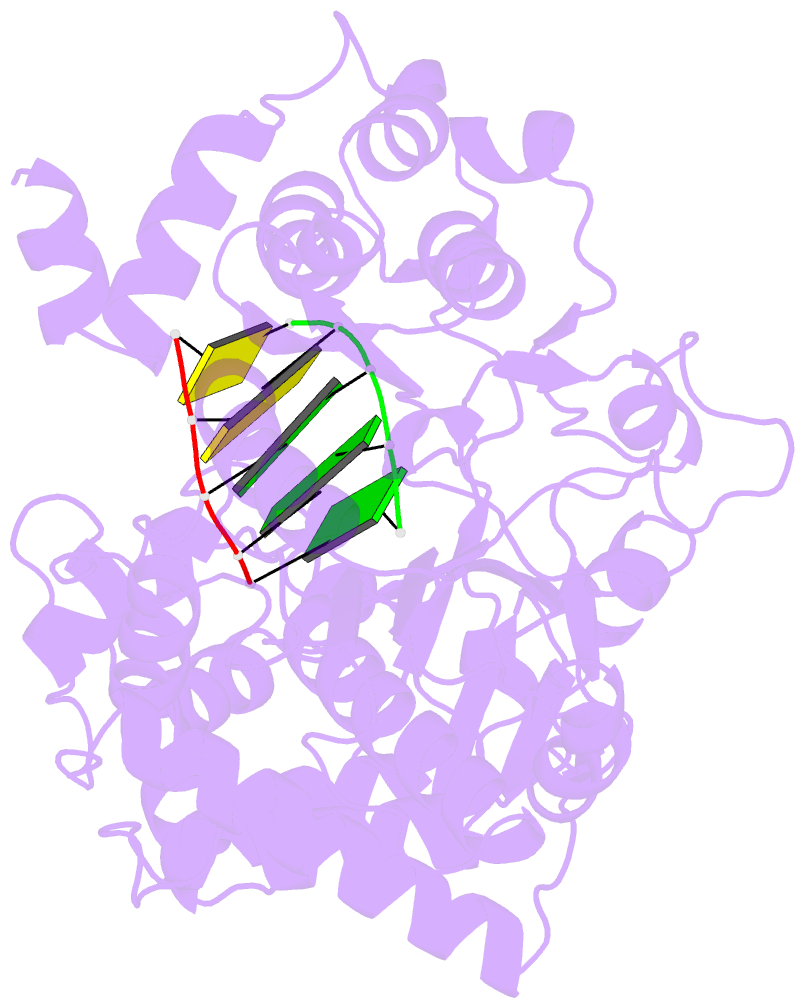
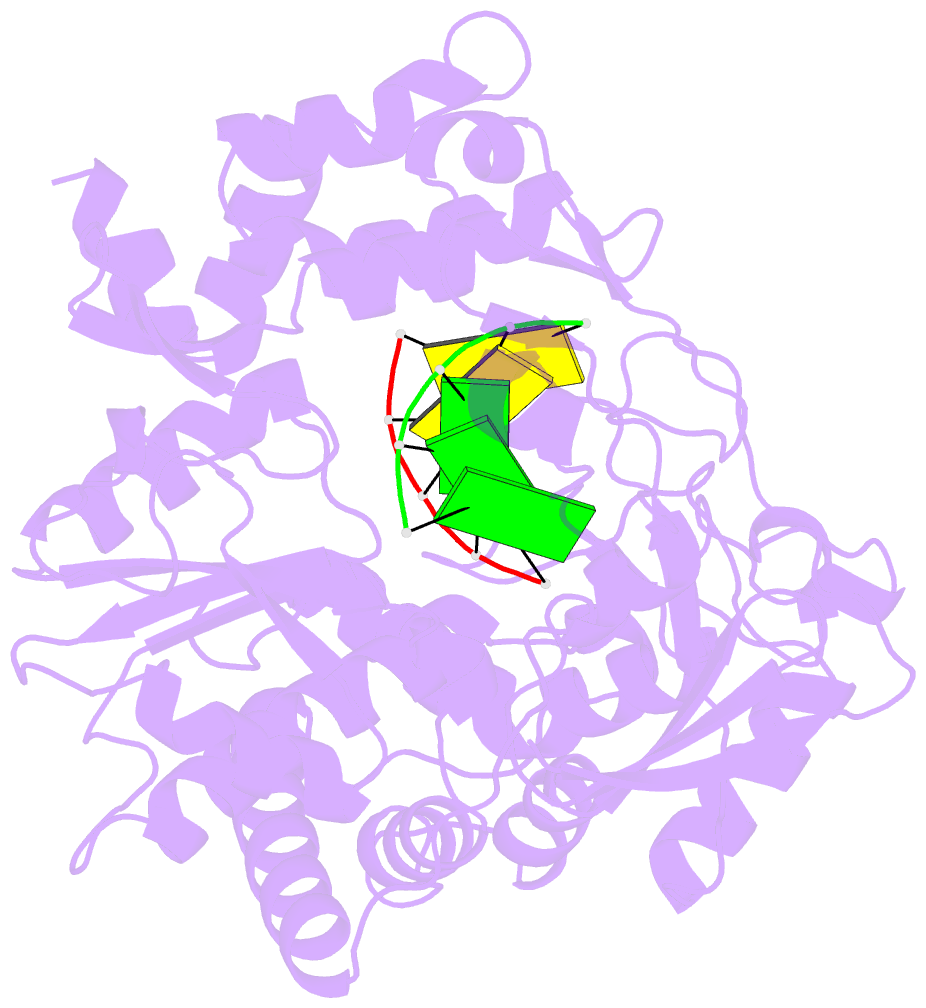
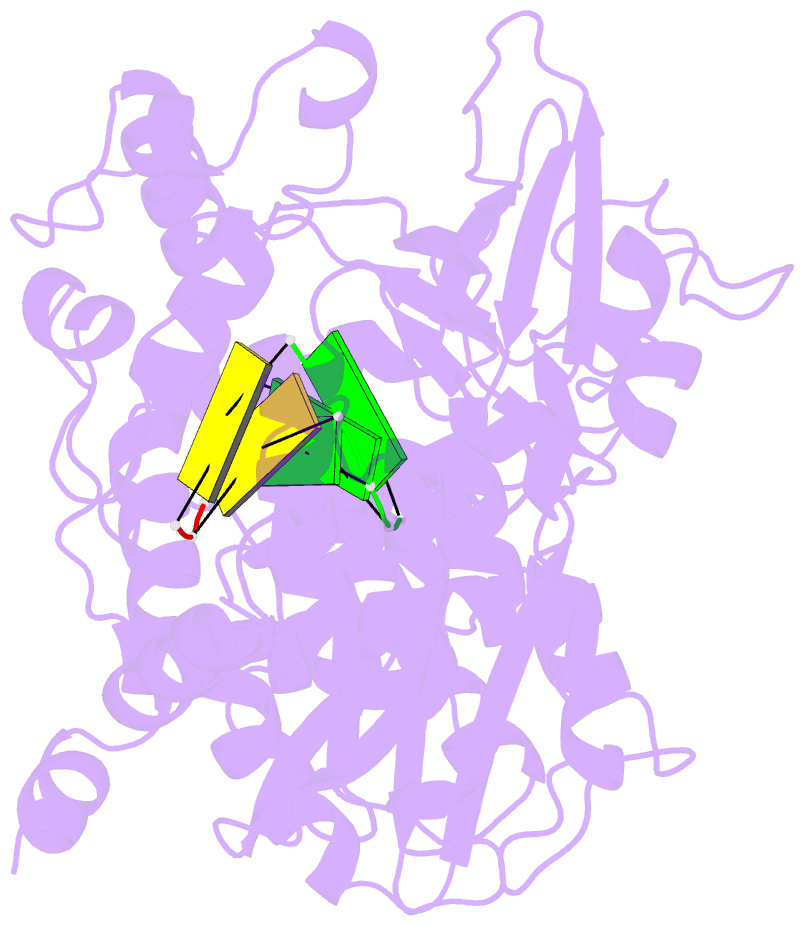
- G62s mutant of foot-and-mouth disease virus RNA-polymerase in complex with a template- primer RNA, tetragonal structure
- Reference
- Ferrer-Orta C, Sierra M, Agudo R, de la Higuera I, Arias A, Perez-Luque R, Escarmis C, Domingo E, Verdaguer N (2010): "Structure of foot-and-mouth disease virus mutant polymerases with reduced sensitivity to ribavirin." J.Virol., 84, 6188-6199. doi: 10.1128/JVI.02420-09.
- Abstract
- Passage of poliovirus (PV) or foot-and-mouth disease virus (FMDV) in the presence of ribavirin selected for viruses with decreased sensitivity to R, which included different mutations in their polymerase (3D): G64S located in the finger subdomain in the case of PV and M296I located within loop beta9-alpha11 at the active site in the case of FMDV. To investigate why disparate substitutions were selected in two closely related 3Ds, we constructed FMDVs with a 3D that included either G62S (the equivalent replacement in FMDV of PV G64S), M296I, or both substitutions. G62S, but not M296I, inflicts upon FMDV a strong selective disadvantage which is partially compensated for by the substitution M296I. The corresponding mutant polymerases, 3D(G62S), 3D(M296I), and 3D(G62S-M296I), were analyzed functionally and structurally. G62S in 3D impairs RNA-binding, polymerization, and R monophosphate incorporation activities. The X-ray structures of the 3D(G62S)-RNA, 3D(M296I)-RNA, and 3D(G62S-M296I)-RNA complexes show that although the two positions are separated by 13.1 A, the loops where the replacements reside are tightly connected through an extensive network of interactions that reach the polymerase active site. In particular, G62S seems to restrict the flexibility of loop beta9-alpha11 and, as a consequence, the flexibility of the active site and its ability to bind the RNA template. Thus, a localized change in the finger subdomain of 3D may affect the catalytic domain. The results provide a structural interpretation of why different amino acid substitutions were selected to confer R resistance in closely related viruses and reveal a complex network of intra-3D interactions that can affect the recognition of both the RNA template and incoming nucleotide.