Summary information and primary citation
- PDB-id
- 3lwr; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- isomerase-RNA binding protein-RNA
- Method
- X-ray (2.203 Å)
- Summary
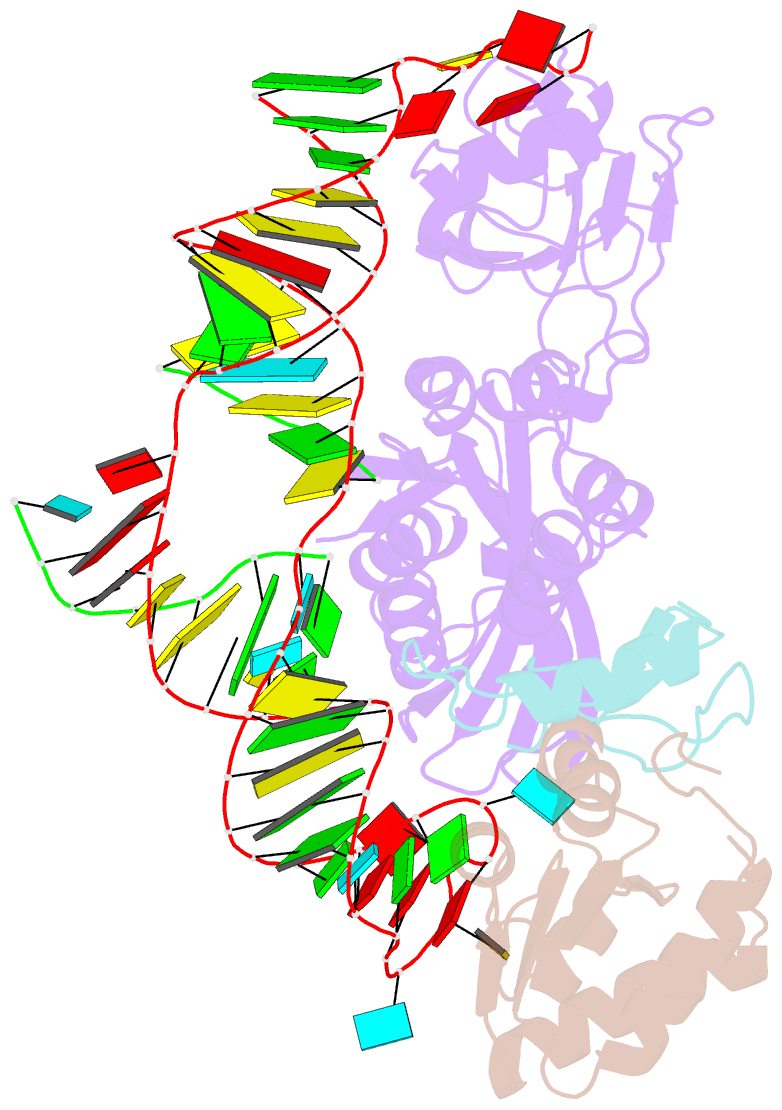
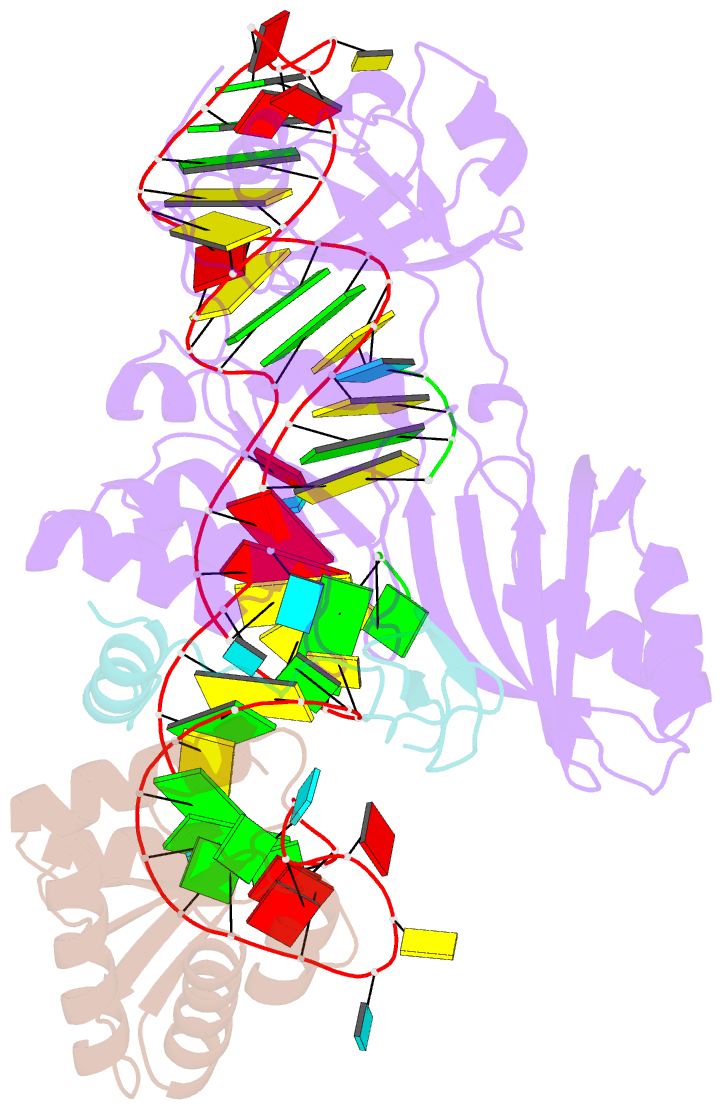
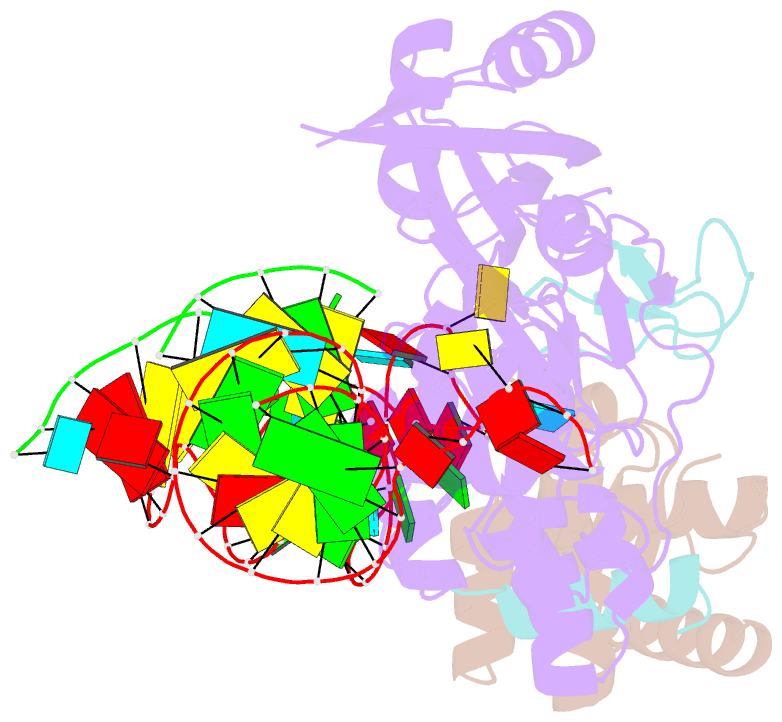
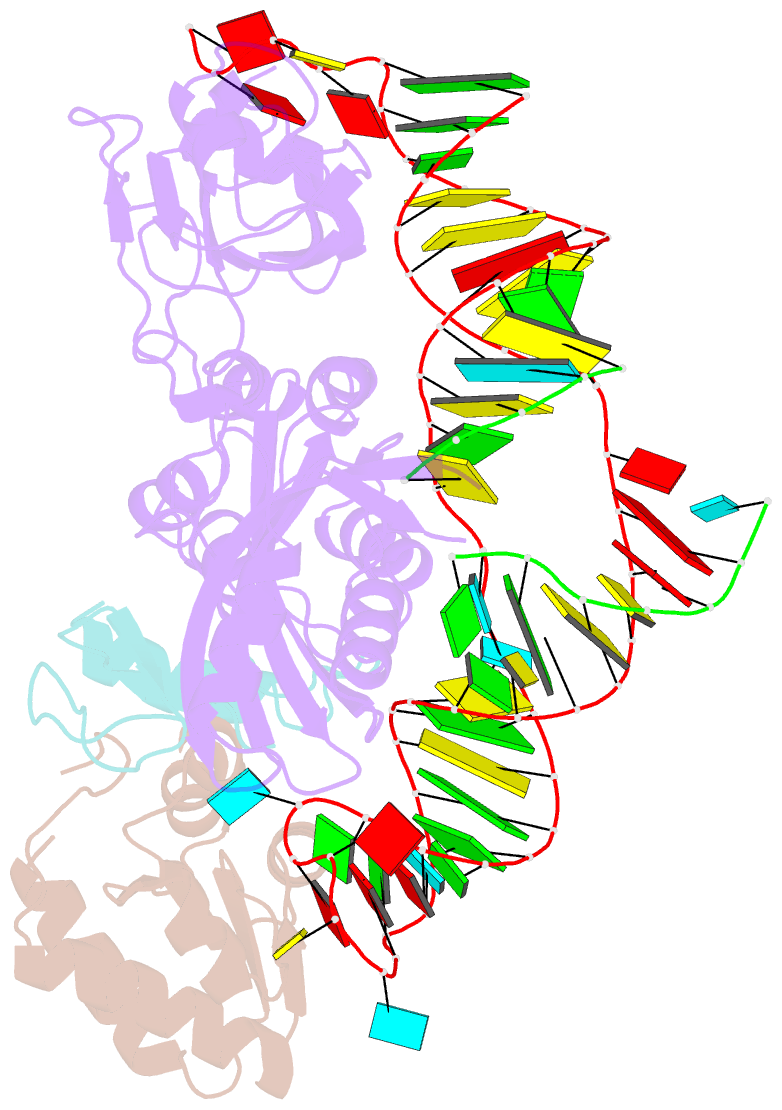
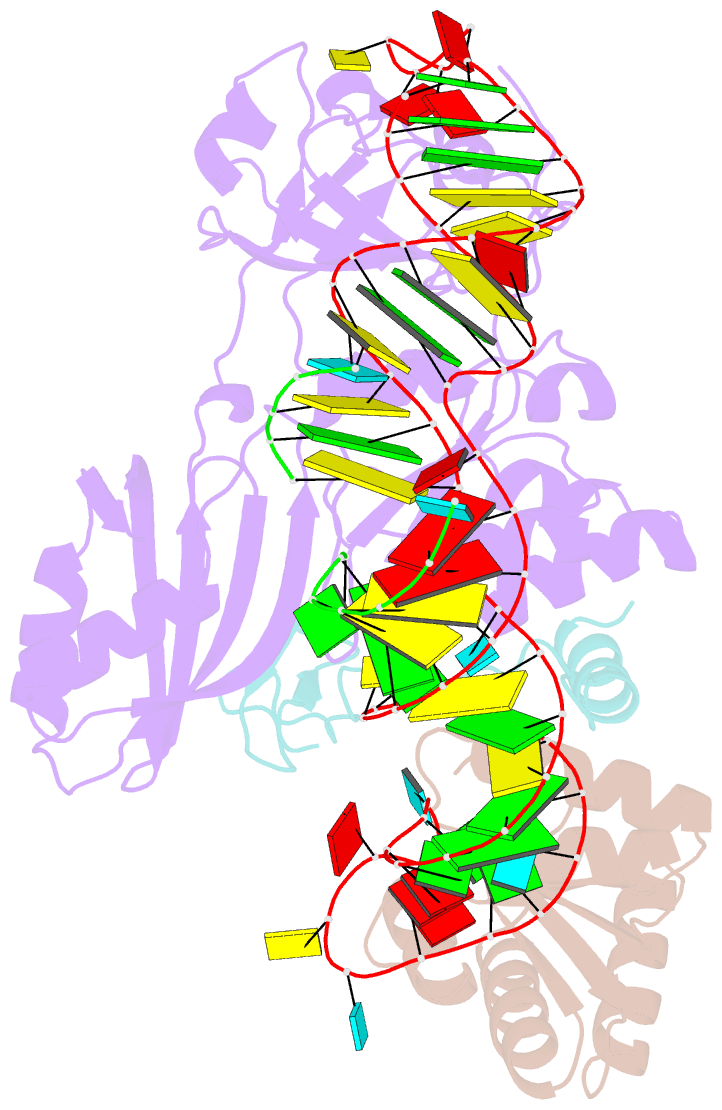
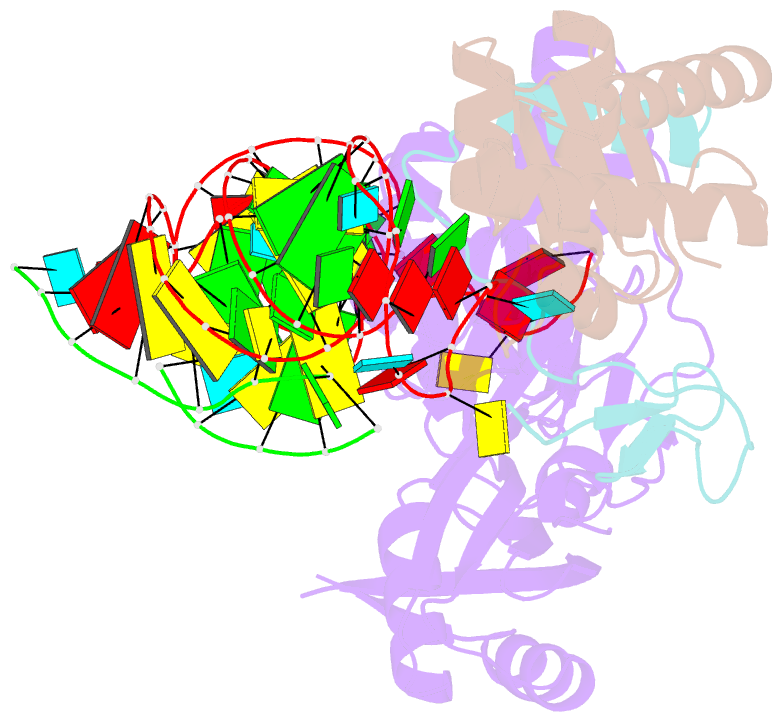
- Structure of h-aca rnp bound to a substrate RNA containing 4su
- Reference
- Zhou J, Liang B, Li H (2010): "Functional and Structural Impact of Target Uridine Substitutions on the H/ACA Ribonucleoprotein Particle Pseudouridine Synthase ." Biochemistry, 49, 6276-6281. doi: 10.1021/bi1006699.
- Abstract
- Box H/ACA ribonucleoprotein protein particles catalyze the majority of pseudouridylation in functional RNA. Different from stand alone pseudouridine synthases, the RNP pseudouridine synthase comprises multiple protein subunits and an RNA subunit. Previous studies showed that each subunit, regardless its location, is sensitive to the step of subunit placement at the catalytic center and potentially to the reaction status of the substrate. Here we describe the impact of chemical substitutions of target uridine on enzyme activity and structure. We found that 3-methyluridine in place of uridine inhibited its isomerization while 2'-deoxyuridine or 4-thiouridine did not. Significantly, crystal structures of an archaeal box H/ACA RNP bound with the nonreactive and the two postreactive substrate analogues showed only subtle structural changes throughout the assembly except for a conserved tyrosine and a substrate anchoring loop of Cbf5. Our results suggest a potential role of these elements and the subunit that contacts them in substrate binding and product release.