Summary information and primary citation
- PDB-id
- 3wfq; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-RNA
- Method
- X-ray (3.619 Å)
- Summary
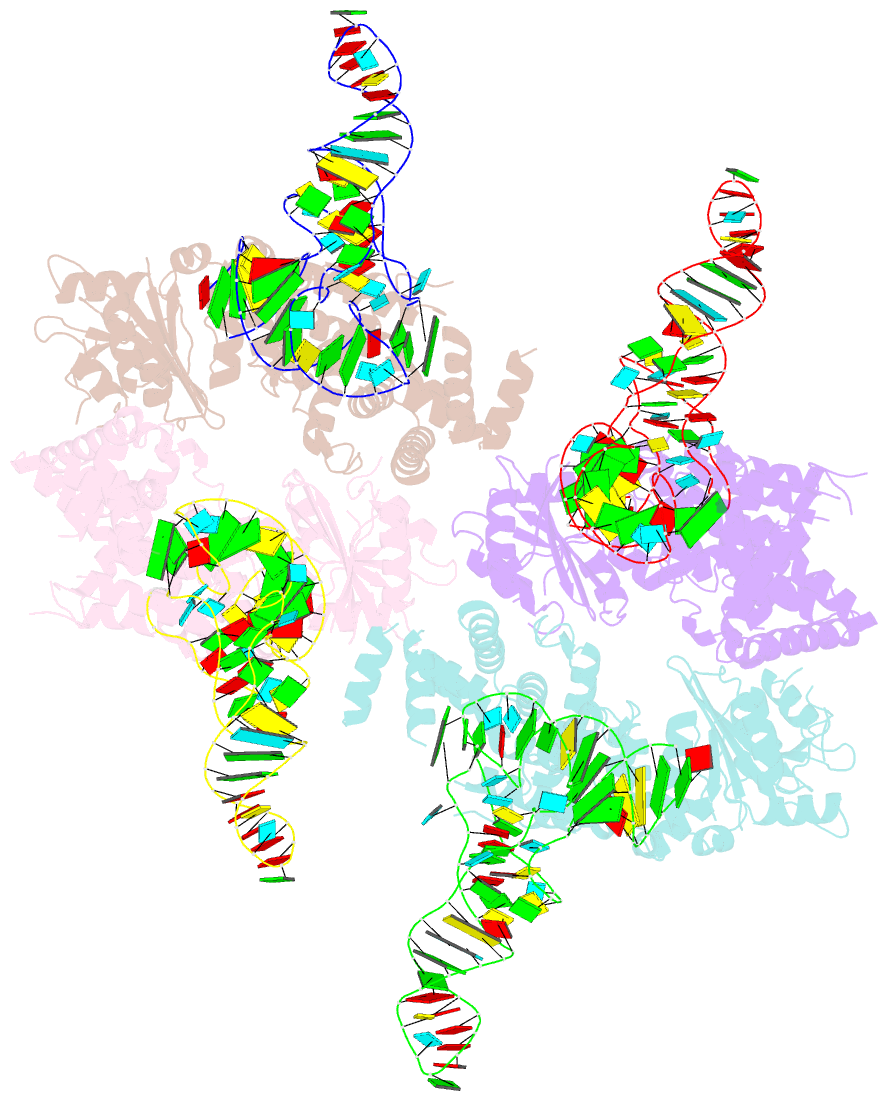
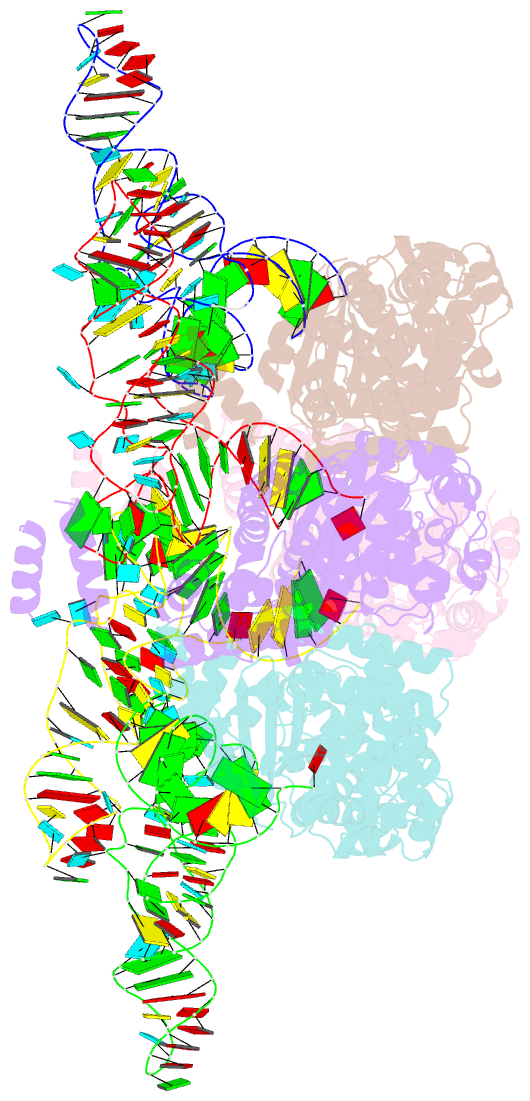
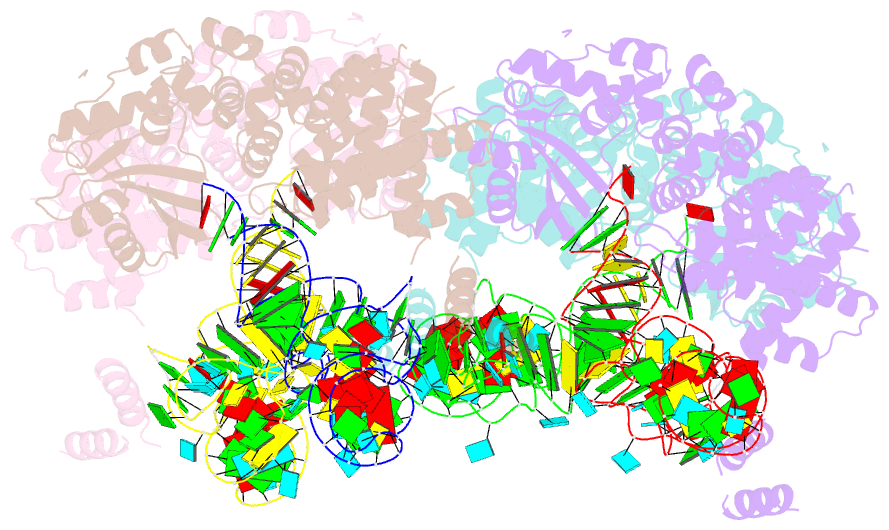
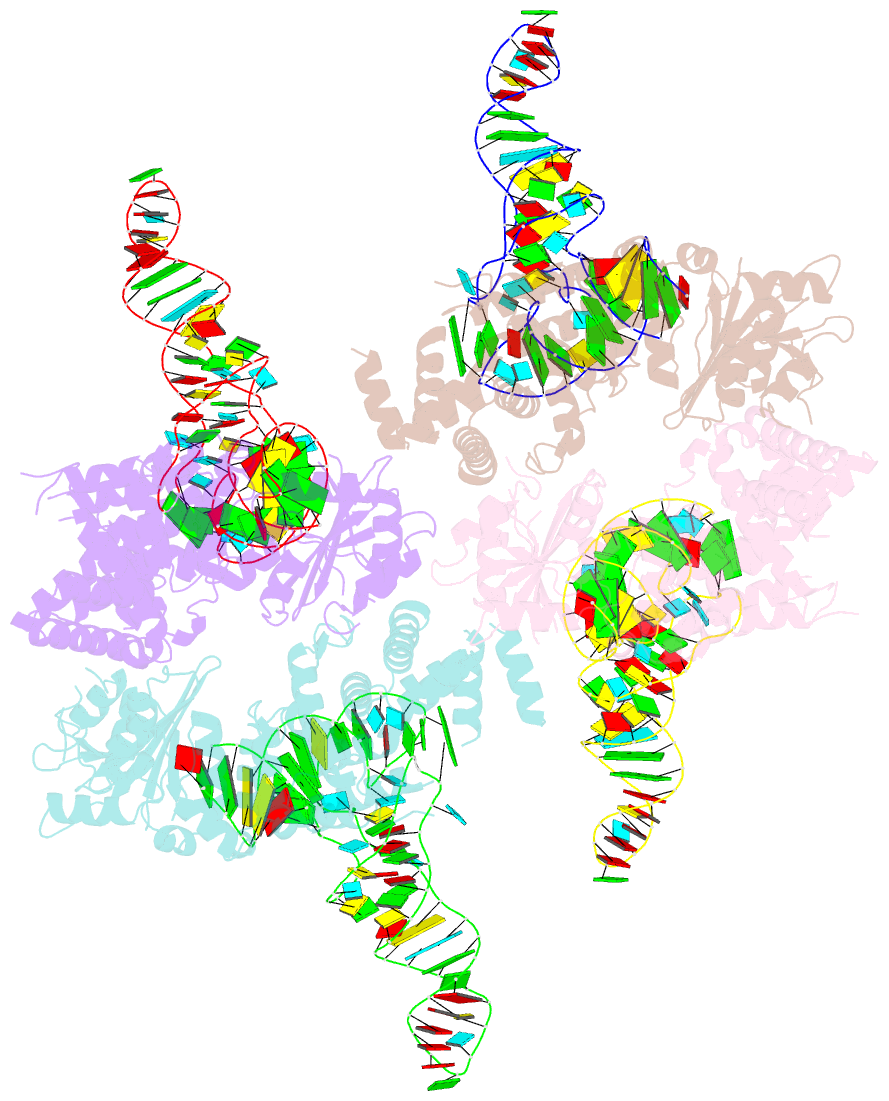
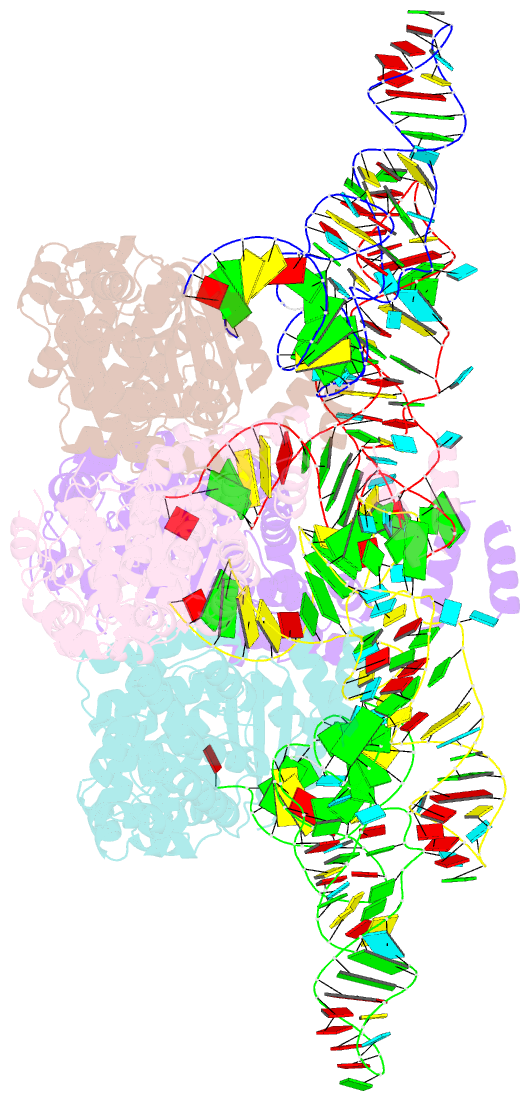
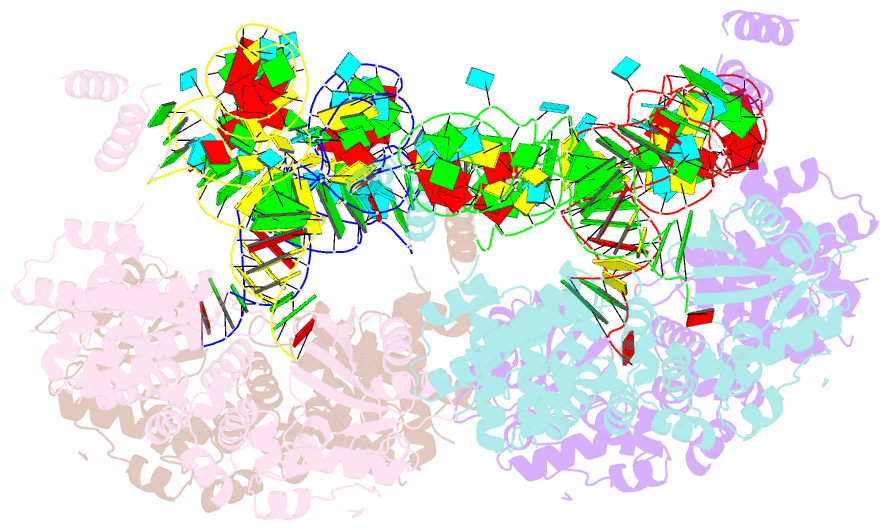
- Trna processing enzyme complex 1
- Reference
- Yamashita S, Takeshita D, Tomita K (2014): "Translocation and rotation of tRNA during template-independent RNA polymerization by tRNA nucleotidyltransferase." Structure, 22, 315-325. doi: 10.1016/j.str.2013.12.002.
- Abstract
- The 3'-terminal CCA (CCA-3' at positions 74-76) of tRNA is synthesized by CCA-adding enzyme using CTP and ATP as substrates, without a nucleic acid template. In Aquifex aeolicus, CC-adding and A-adding enzymes collaboratively synthesize the CCA-3'. The mechanism of CCA-3' synthesis by these two enzymes remained obscure. We now present crystal structures representing CC addition onto tRNA by A. aeolicus CC-adding enzyme. After C₇₄ addition in an enclosed active pocket and pyrophosphate release, the tRNA translocates and rotates relative to the enzyme, and C₇₅ addition occurs in the same active pocket as C₇₄ addition. At both the C₇₄-adding and C₇₅-adding stages, CTP is selected by Watson-Crick-like hydrogen bonds between the cytosine of CTP and conserved Asp and Arg residues in the pocket. After C₇₄C₇₅ addition and pyrophosphate release, the tRNA translocates further and drops off the enzyme, and the CC-adding enzyme terminates RNA polymerization.