Summary information and primary citation
- PDB-id
- 4a0k; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- ligase-DNA-binding protein-DNA
- Method
- X-ray (5.93 Å)
- Summary
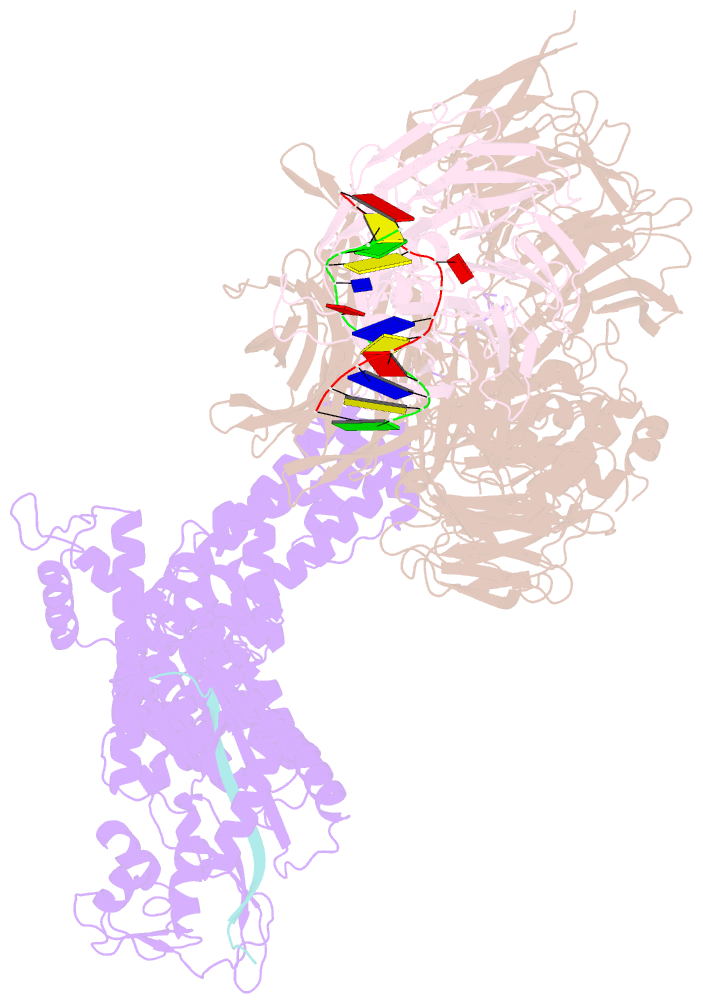
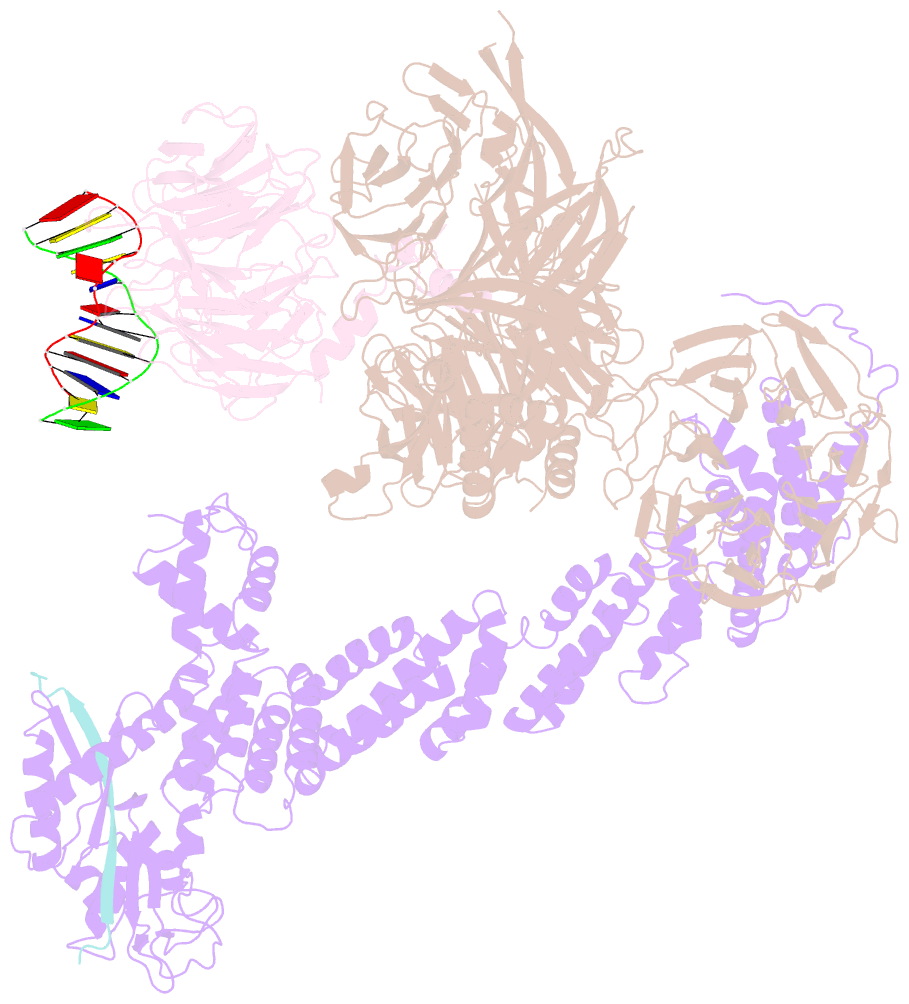
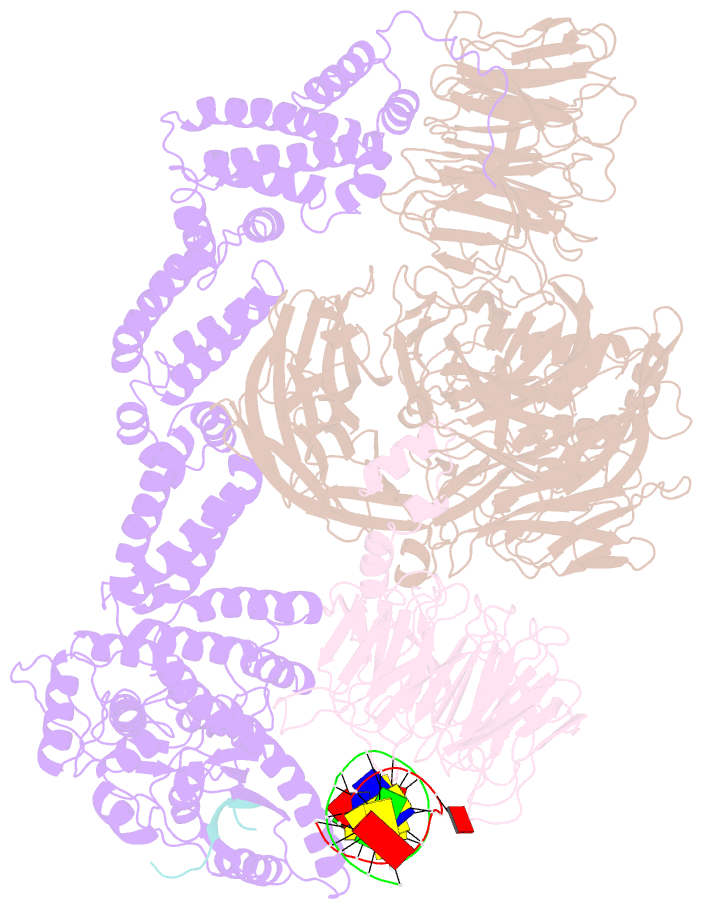
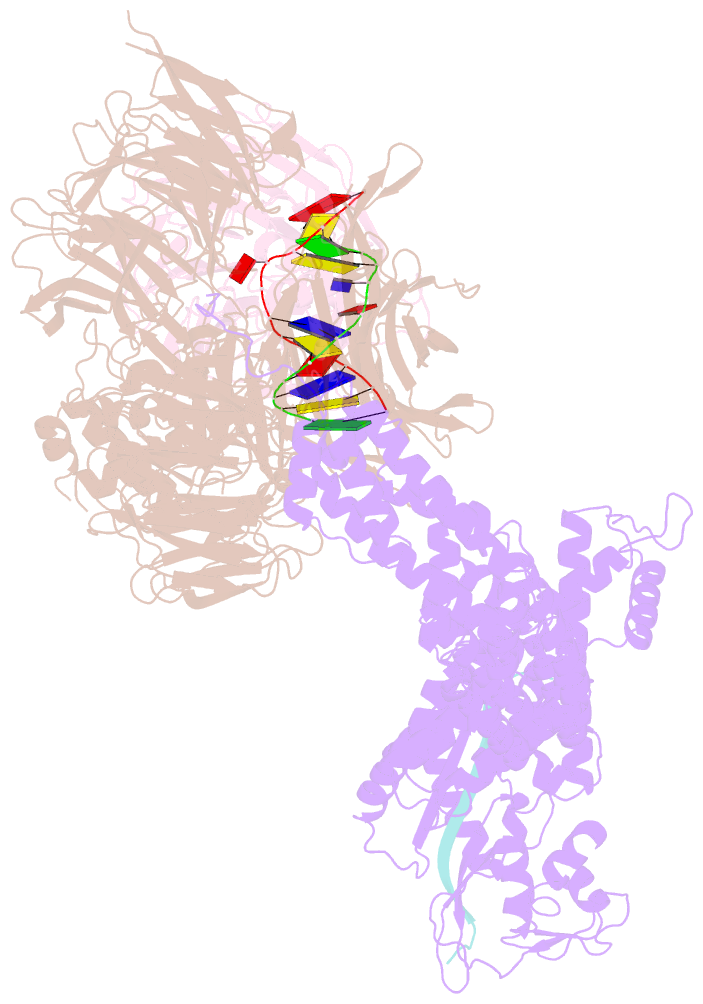
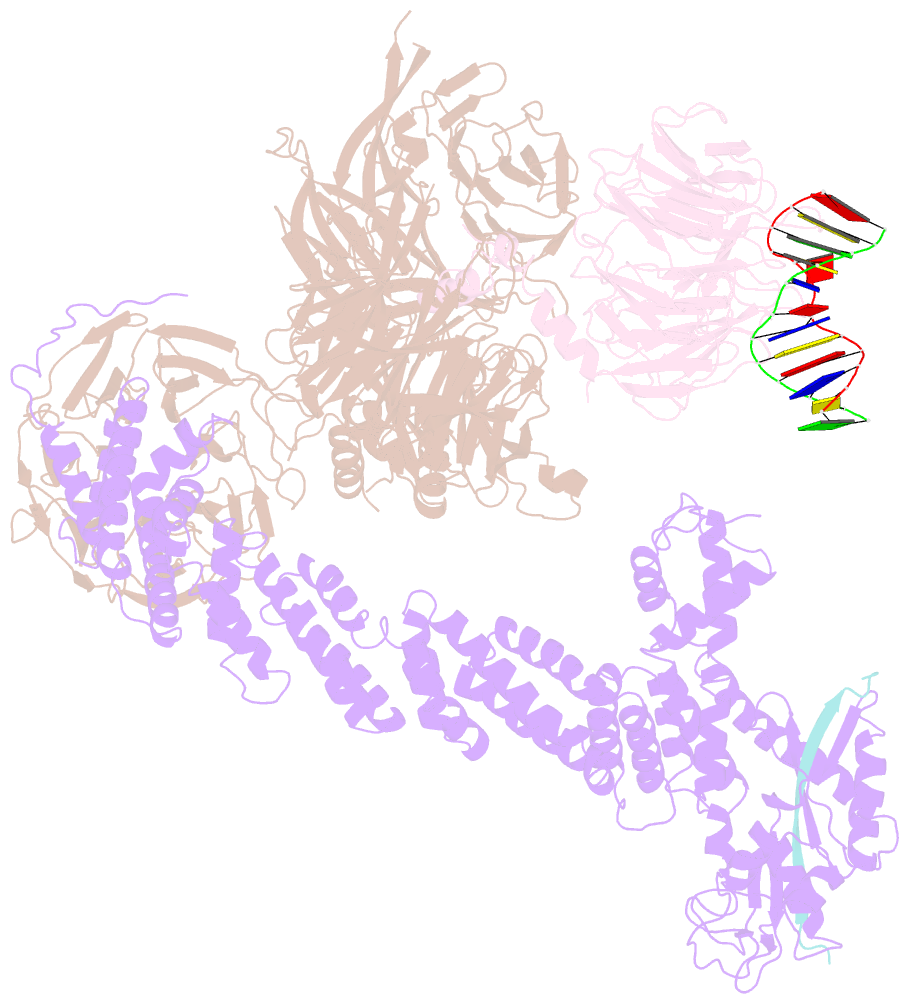
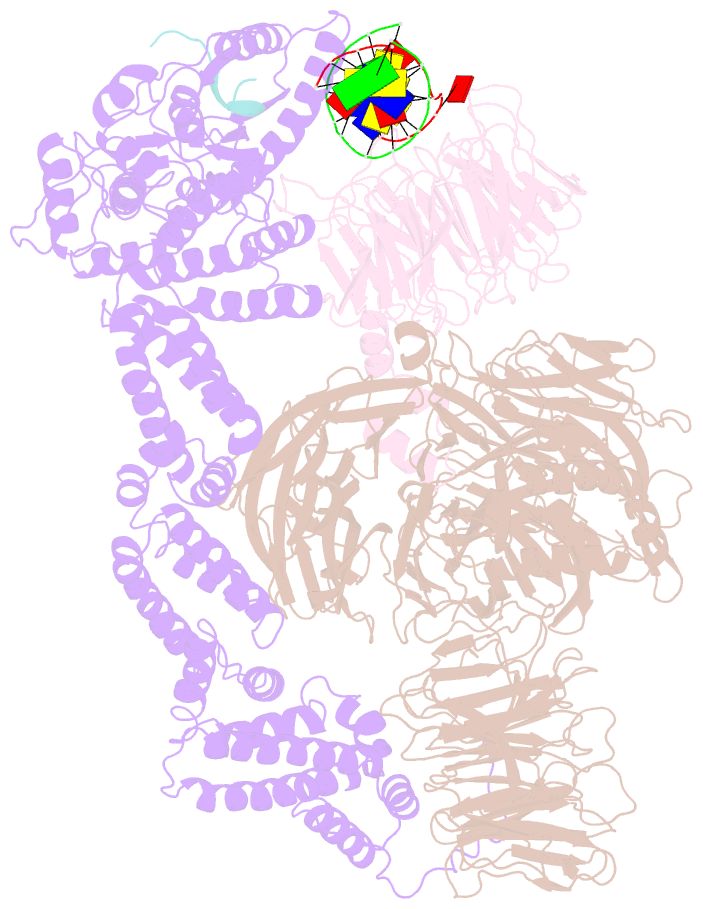
- Structure of ddb1-ddb2-cul4a-rbx1 bound to a 12 bp abasic site containing DNA-duplex
- Reference
- Fischer ES, Scrima A, Bohm K, Matsumoto S, Lingaraju GM, Faty M, Yasuda T, Cavadini S, Wakasugi M, Hanaoka F, Iwai S, Gut H, Sugasawa K, Thoma NH (2011): "The Molecular Basis of Crl4(Ddb2/Csa) Ubiquitin Ligase Architecture, Targeting, and Activation." Cell(Cambridge,Mass.), 147, 1024. doi: 10.1016/J.CELL.2011.10.035.
- Abstract
- The DDB1-CUL4-RBX1 (CRL4) ubiquitin ligase family regulates a diverse set of cellular pathways through dedicated substrate receptors (DCAFs). The DCAF DDB2 detects UV-induced pyrimidine dimers in the genome and facilitates nucleotide excision repair. We provide the molecular basis for DDB2 receptor-mediated cyclobutane pyrimidine dimer recognition in chromatin. The structures of the fully assembled DDB1-DDB2-CUL4A/B-RBX1 (CRL4(DDB2)) ligases reveal that the mobility of the ligase arm creates a defined ubiquitination zone around the damage, which precludes direct ligase activation by DNA lesions. Instead, the COP9 signalosome (CSN) mediates the CRL4(DDB2) inhibition in a CSN5 independent, nonenzymatic, fashion. In turn, CSN inhibition is relieved upon DNA damage binding to the DDB2 module within CSN-CRL4(DDB2). The Cockayne syndrome A DCAF complex crystal structure shows that CRL4(DCAF(WD40)) ligases share common architectural features. Our data support a general mechanism of ligase activation, which is induced by CSN displacement from CRL4(DCAF) on substrate binding to the DCAF.