Summary information and primary citation
- PDB-id
- 4aad; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydrolase-DNA
- Method
- X-ray (3.1 Å)
- Summary
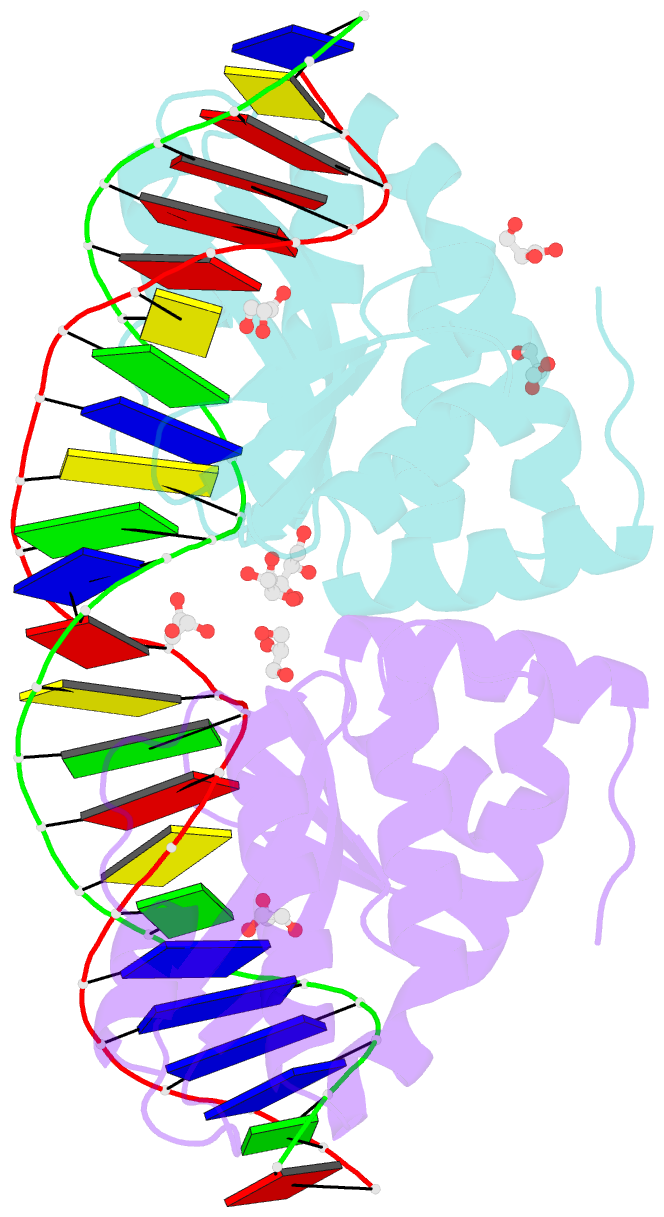
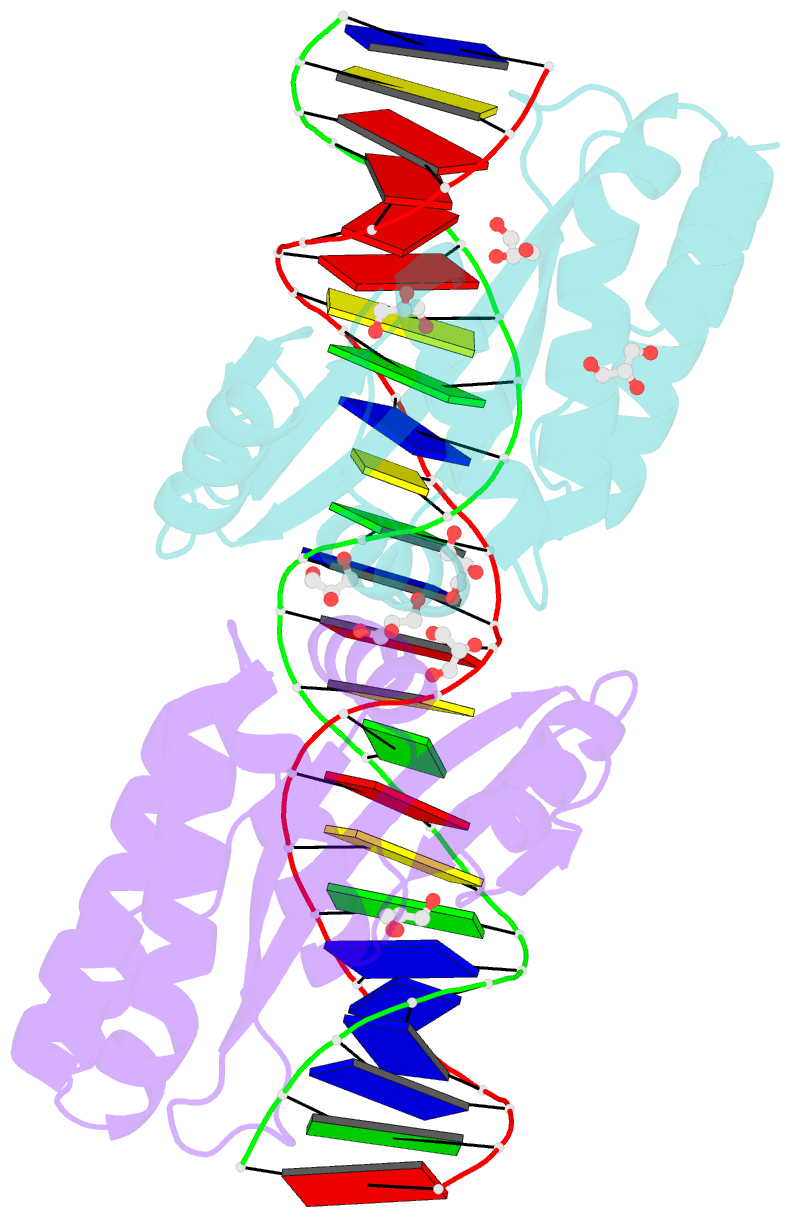
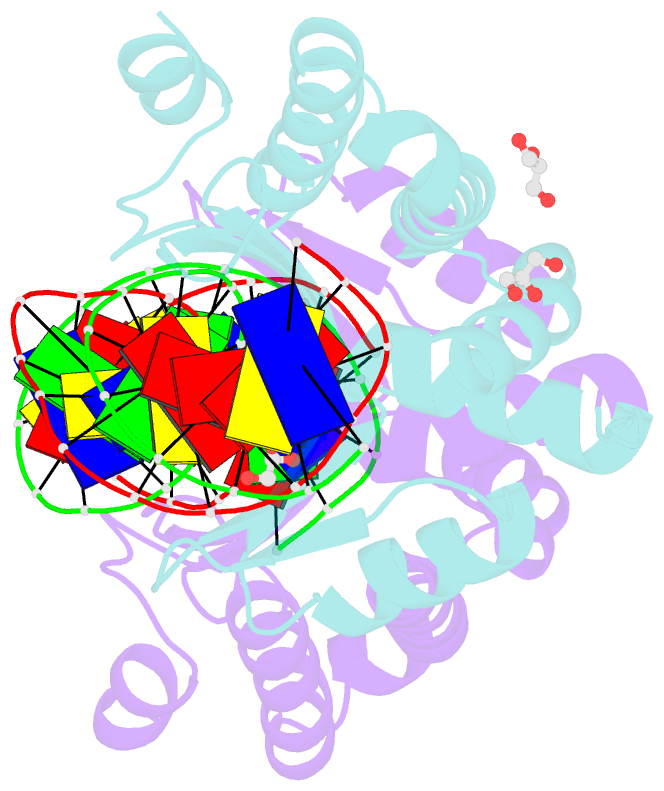
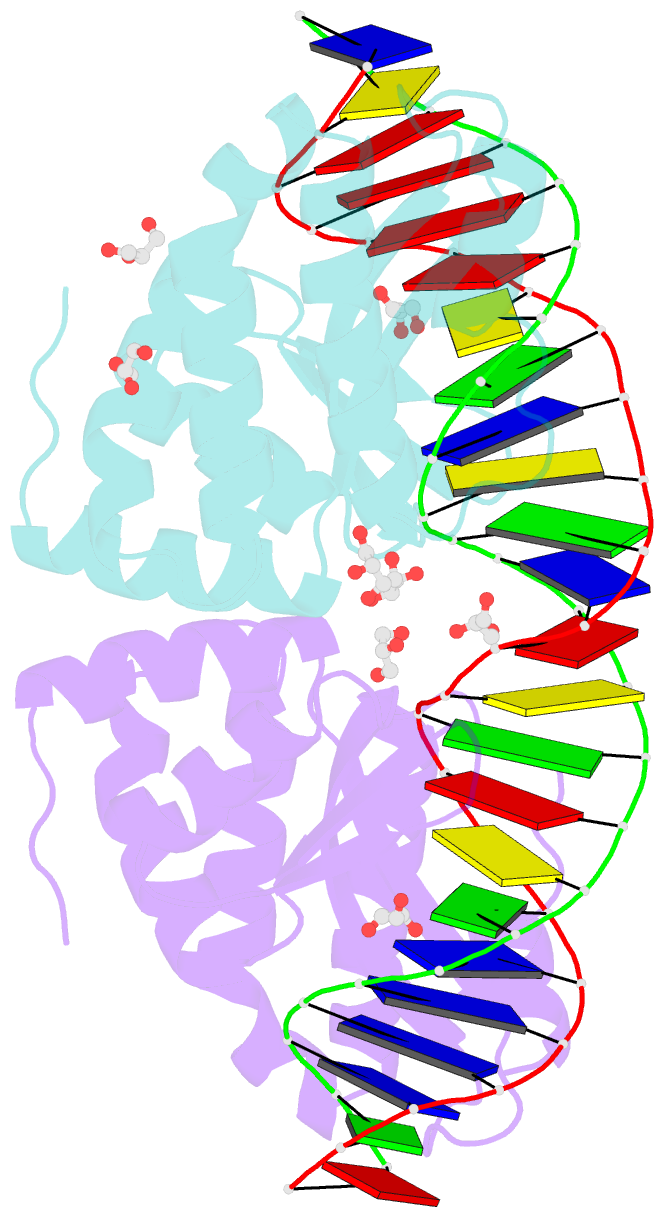
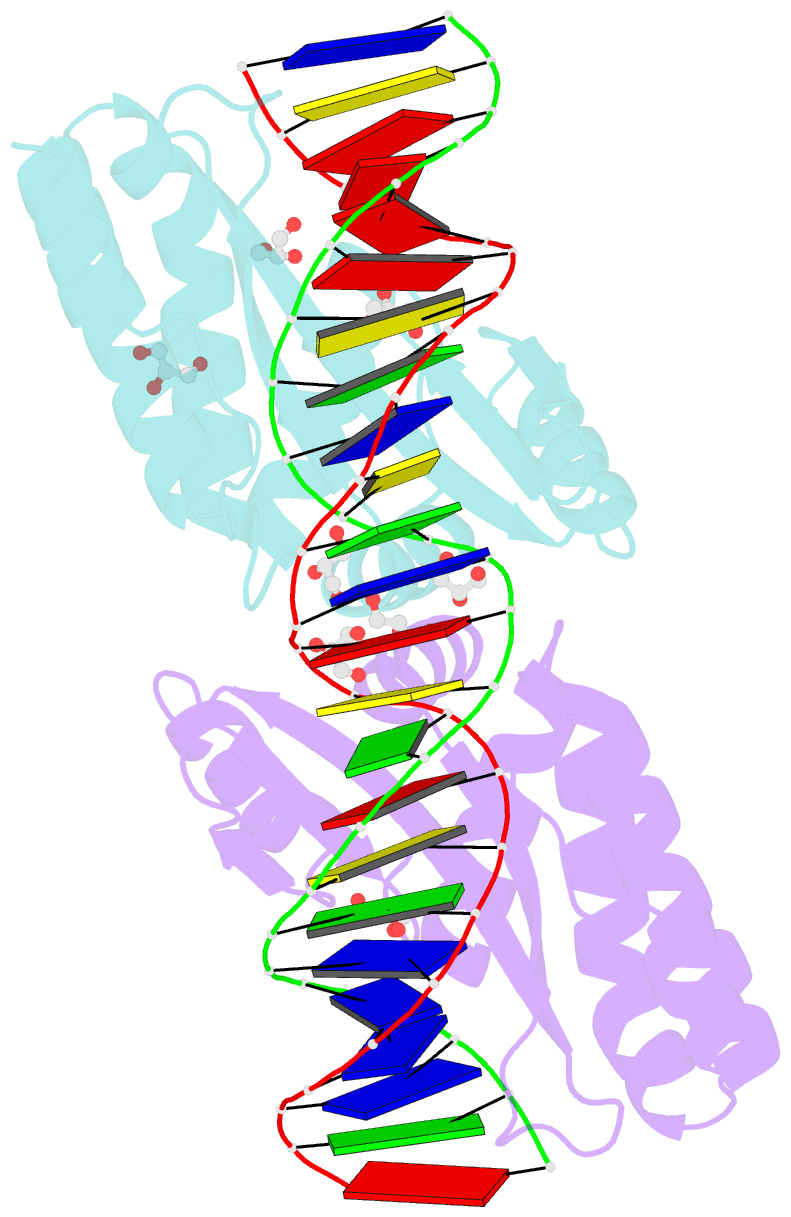
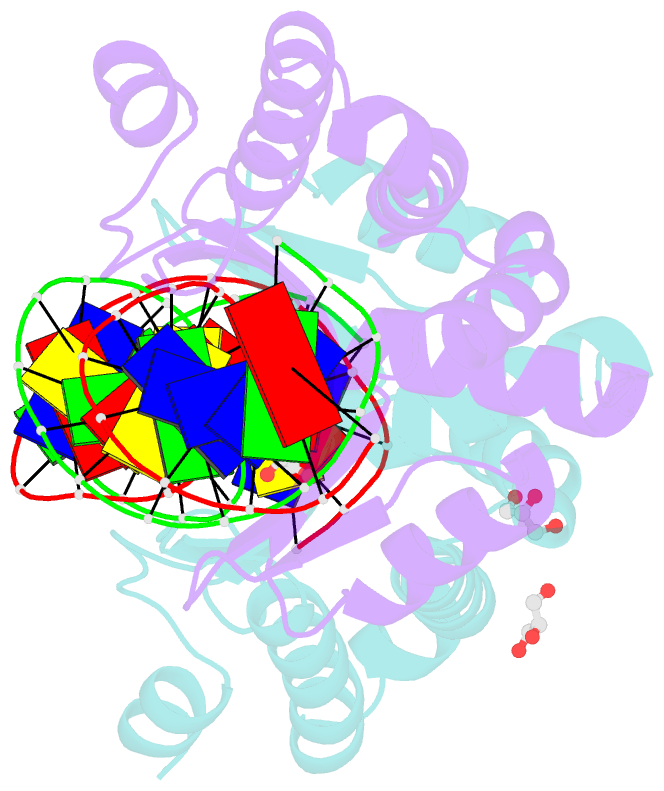
- Crystal structure of the mutant d75n i-crei in complex with its wild- type target in absence of metal ions at the active site (the four central bases, 2nn region, are composed by gtac from 5' to 3')
- Reference
- Molina R, Redondo P, Stella S, Marenchino M, D'Abramo M, Gervasio FL, Charles Epinat J, Valton J, Grizot S, Duchateau P, Prieto J, Montoya G (2012): "Non-Specific Protein-DNA Interactions Control I-Crei Target Binding and Cleavage." Nucleic Acids Res., 40, 6936-6945. doi: 10.1093/NAR/GKS320.
- Abstract
- Homing endonucleases represent protein scaffolds that provide powerful tools for genome manipulation, as these enzymes possess a very low frequency of DNA cleavage in eukaryotic genomes due to their high specificity. The basis of protein-DNA recognition must be understood to generate tailored enzymes that target the DNA at sites of interest. Protein-DNA interaction engineering of homing endonucleases has demonstrated the potential of these approaches to create new specific instruments to target genes for inactivation or repair. Protein-DNA interface studies have been focused mostly on specific contacts between amino acid side chains and bases to redesign the binding interface. However, it has been shown that 4 bp in the central DNA sequence of the 22-bp substrate of a homing endonuclease (I-CreI), which do not show specific protein-DNA interactions, is not devoid of content information. Here, we analyze the mechanism of target discrimination in this substrate region by the I-CreI protein, determining how it can occur independently of the specific protein-DNA interactions. Our data suggest the important role of indirect readout in this substrate region, opening the possibility for a fully rational search of new target sequences, thus improving the development of redesigned enzymes for therapeutic and biotechnological applications.