Summary information and primary citation
- PDB-id
- 4ea5; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydrolase-DNA
- Method
- X-ray (2.14 Å)
- Summary
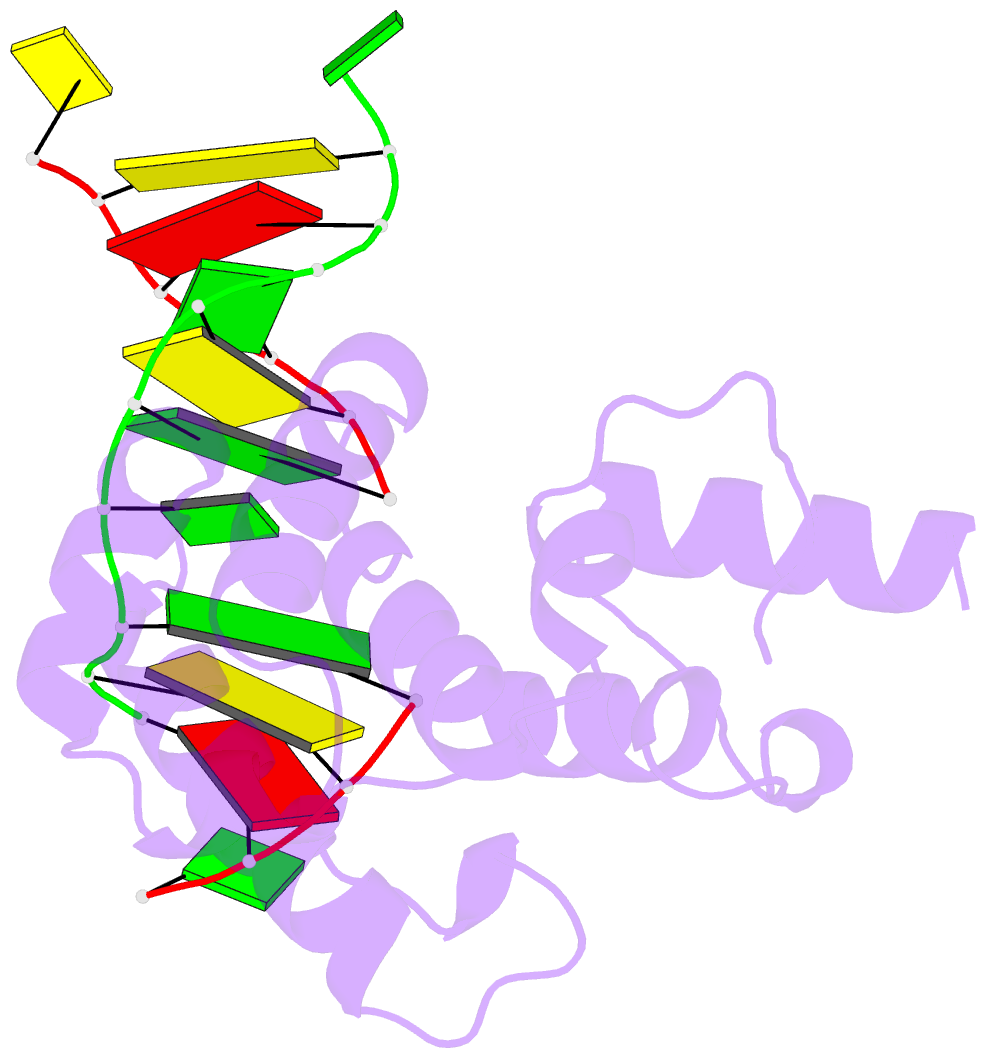
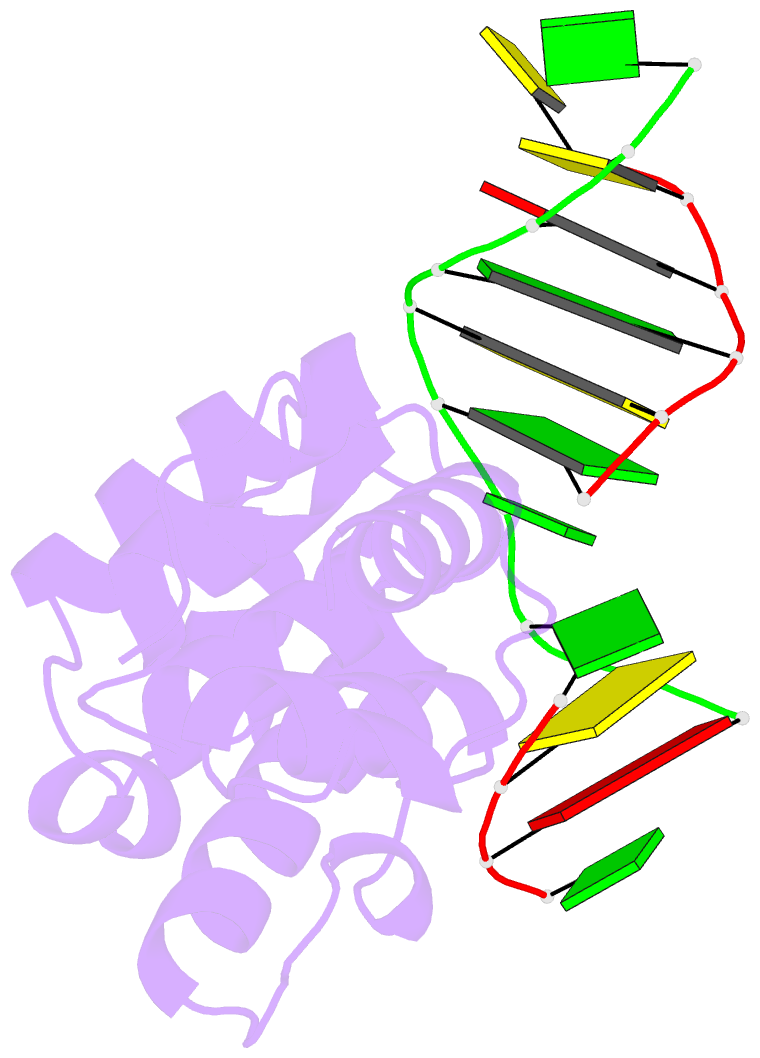
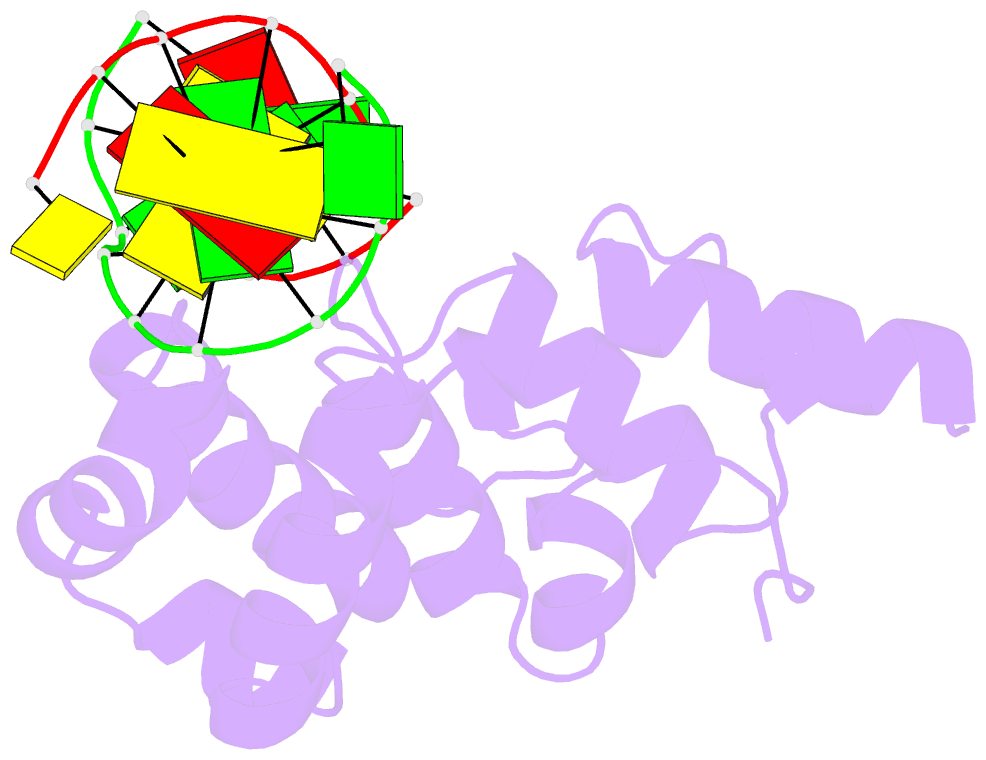
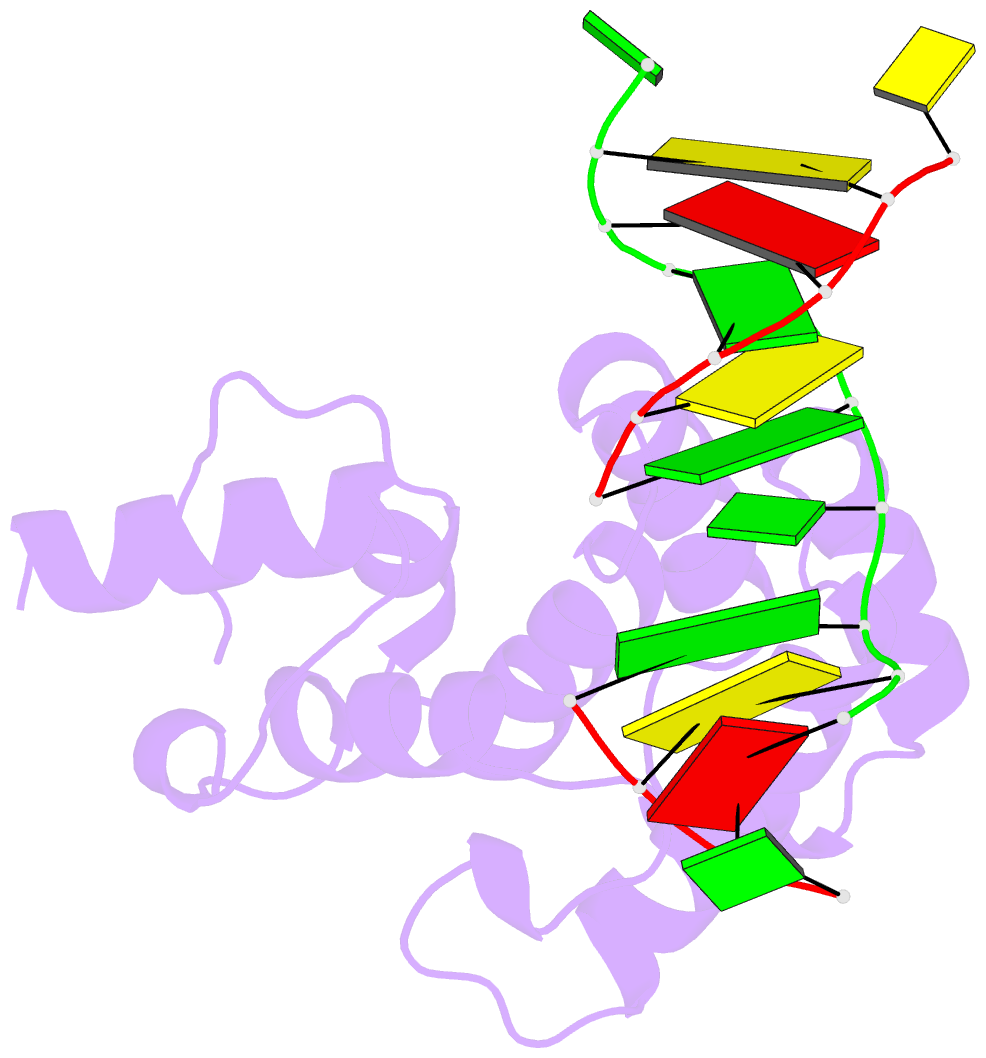
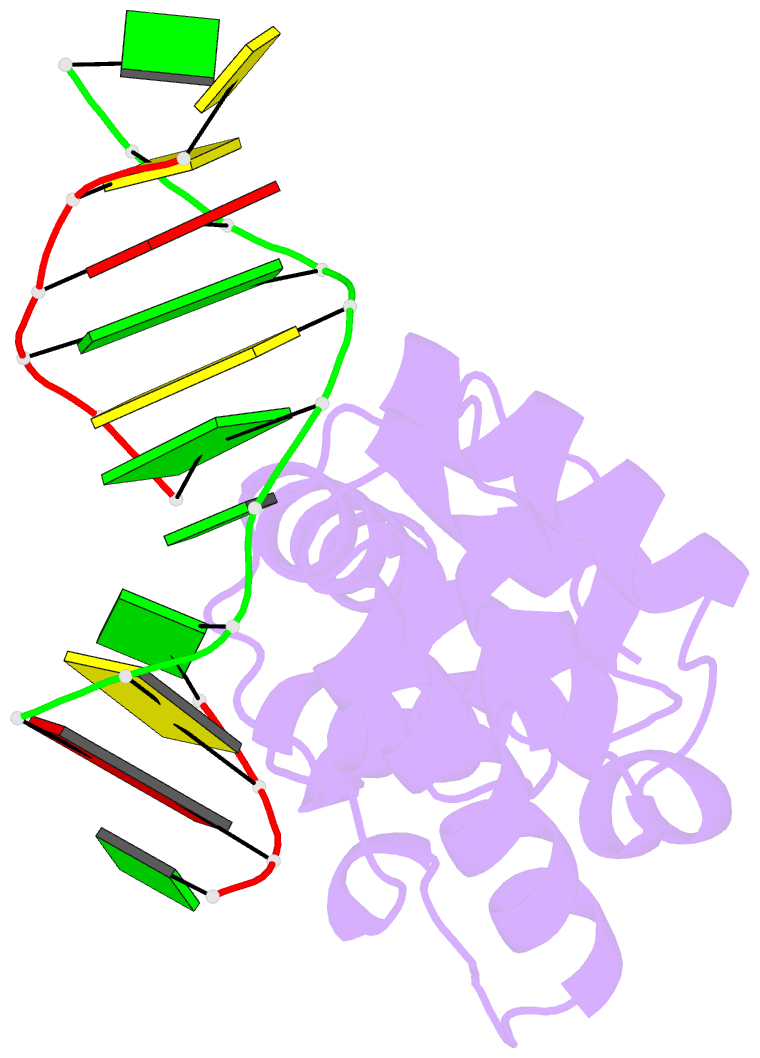
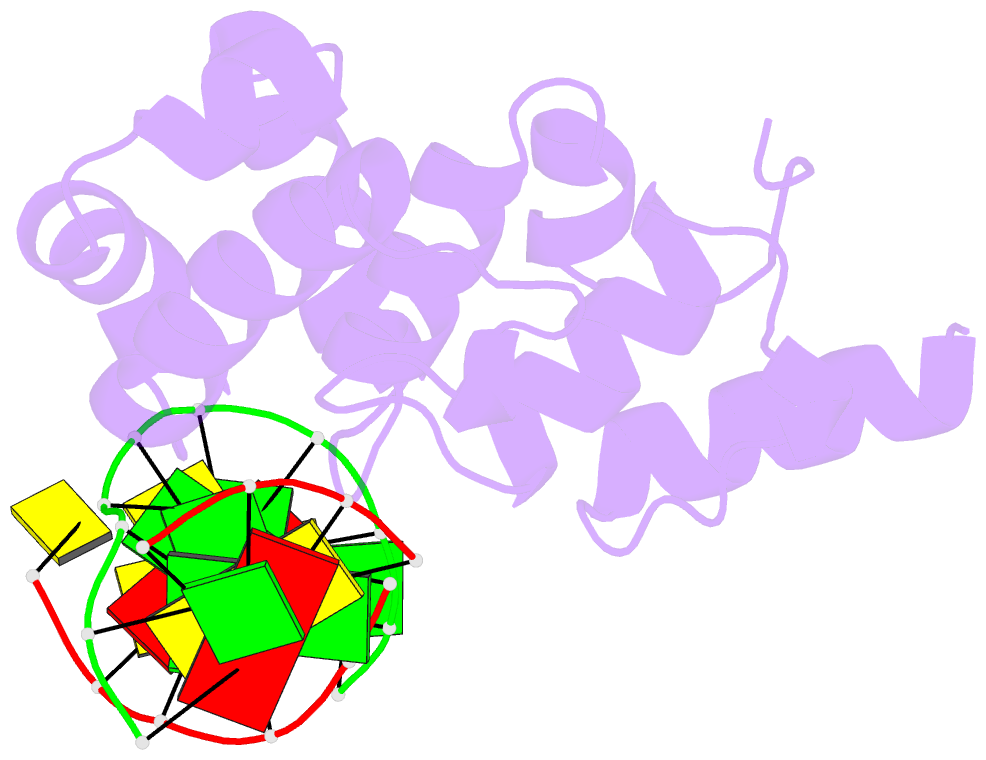
- Structure of the glycoslyase domain of mbd4 bound to a 5hmu containing DNA
- Reference
- Morera S, Grin I, Vigouroux A, Couve S, Henriot V, Saparbaev M, Ishchenko AA (2012): "Biochemical and structural characterization of the glycosylase domain of MBD4 bound to thymine and 5-hydroxymethyuracil-containing DNA." Nucleic Acids Res., 40, 9917-9926. doi: 10.1093/nar/gks714.
- Abstract
- Active DNA demethylation in mammals occurs via hydroxylation of 5-methylcytosine to 5-hydroxymethylcytosine (5hmC) by the ten-eleven translocation family of proteins (TETs). 5hmC residues in DNA can be further oxidized by TETs to 5-carboxylcytosines and/or deaminated by the Activation Induced Deaminase/Apolipoprotein B mRNA-editing enzyme complex family proteins to 5-hydromethyluracil (5hmU). Excision and replacement of these intermediates is initiated by DNA glycosylases such as thymine-DNA glycosylase (TDG), methyl-binding domain protein 4 (MBD4) and single-strand specific monofunctional uracil-DNA glycosylase 1 in the base excision repair pathway. Here, we report detailed biochemical and structural characterization of human MBD4 which contains mismatch-specific TDG activity. Full-length as well as catalytic domain (residues 426-580) of human MBD4 (MBD4(cat)) can remove 5hmU when opposite to G with good efficiency. Here, we also report six crystal structures of human MBD4(cat): an unliganded form and five binary complexes with duplex DNA containing a T•G, 5hmU•G or AP•G (apurinic/apyrimidinic) mismatch at the target base pair. These structures reveal that MBD4(cat) uses a base flipping mechanism to specifically recognize thymine and 5hmU. The recognition mechanism of flipped-out 5hmU bases in MBD4(cat) active site supports the potential role of MBD4, together with TDG, in maintenance of genome stability and active DNA demethylation in mammals.