Summary information and primary citation
- PDB-id
- 4pdb; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- ribosomal protein-RNA
- Method
- X-ray (2.6 Å)
- Summary
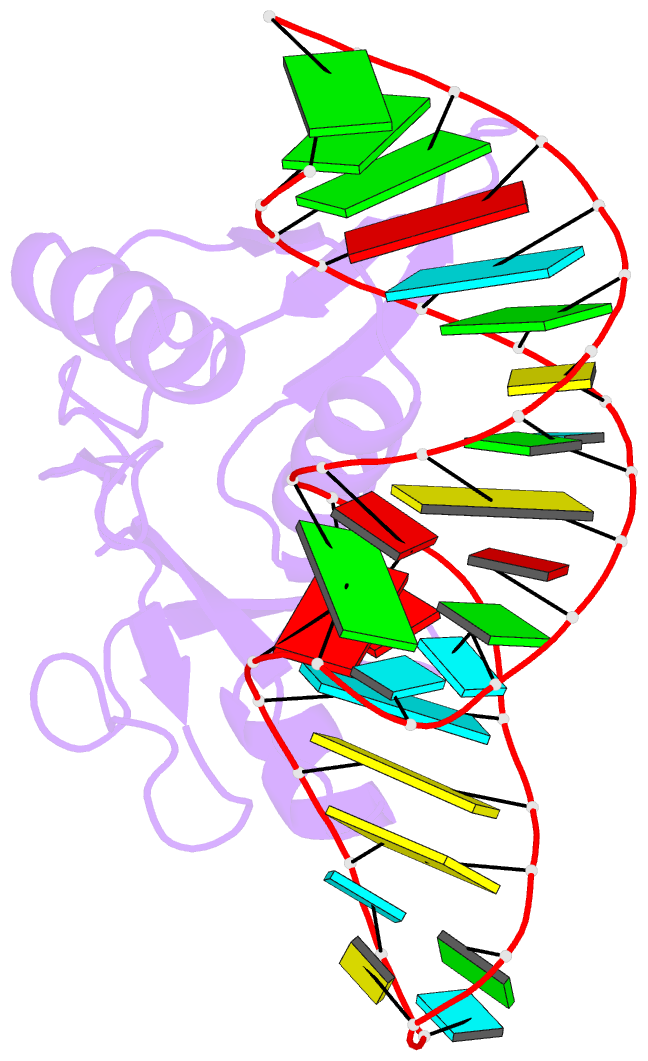
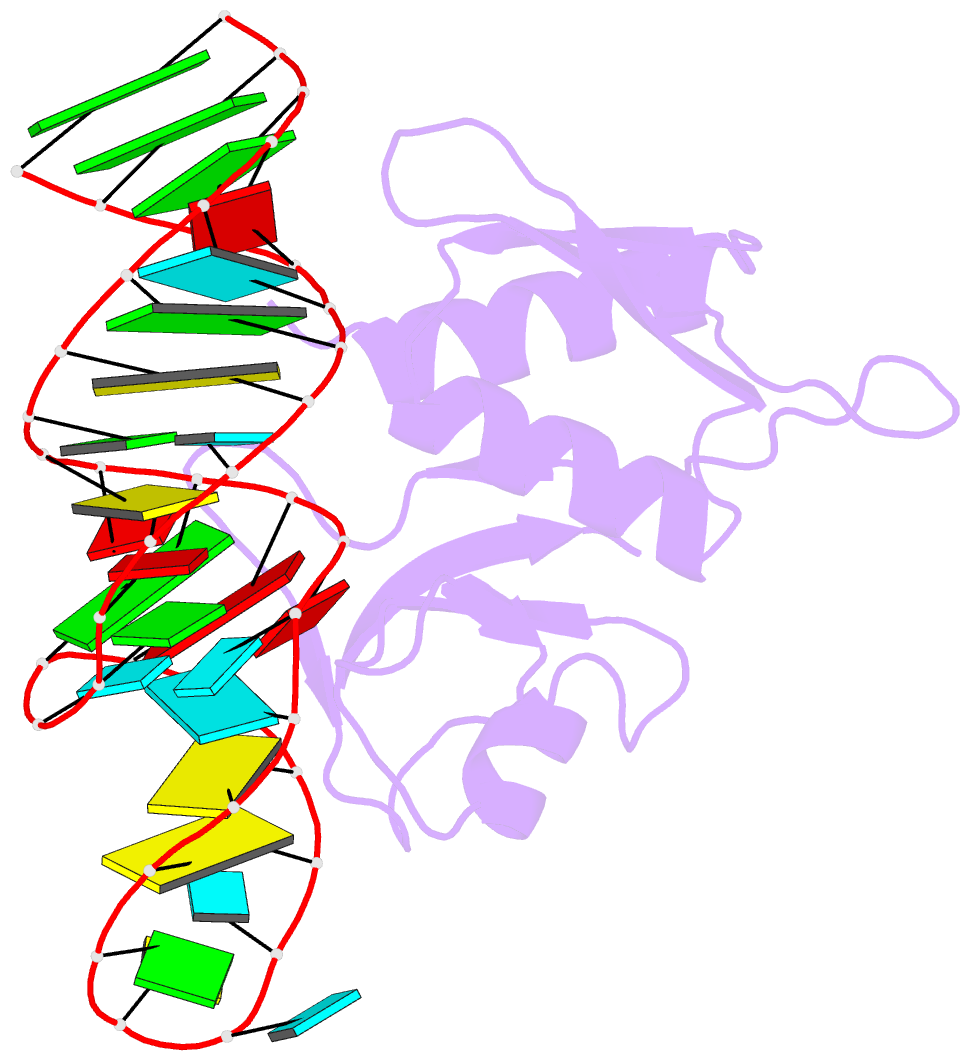
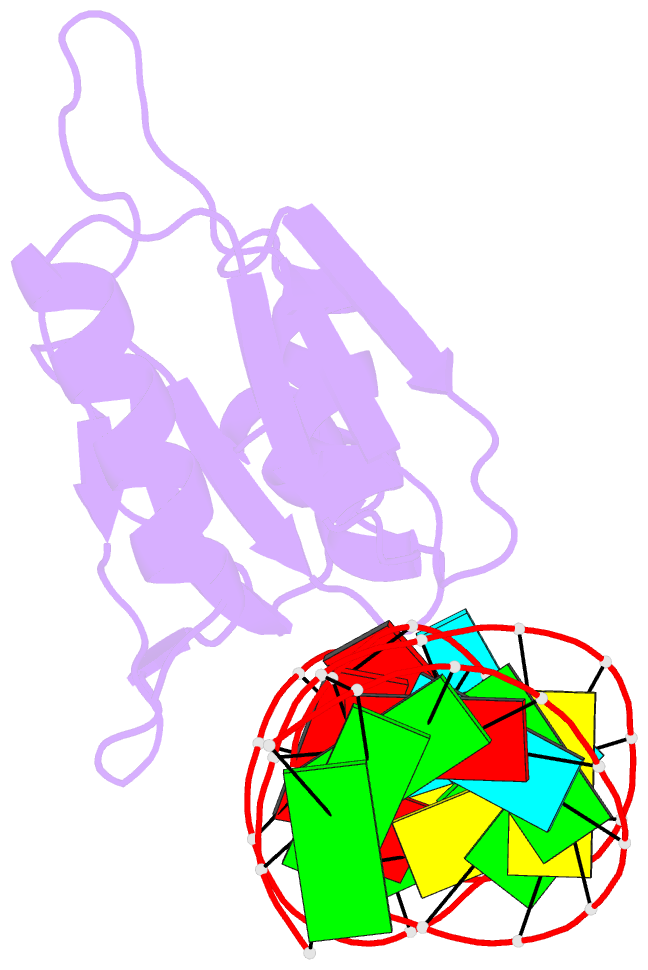
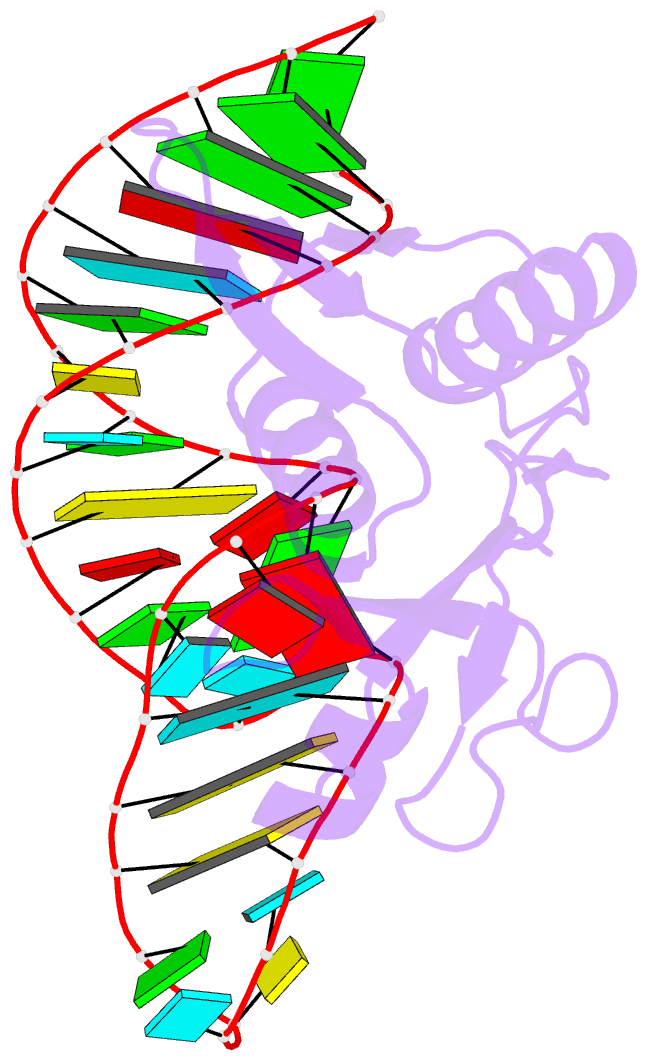
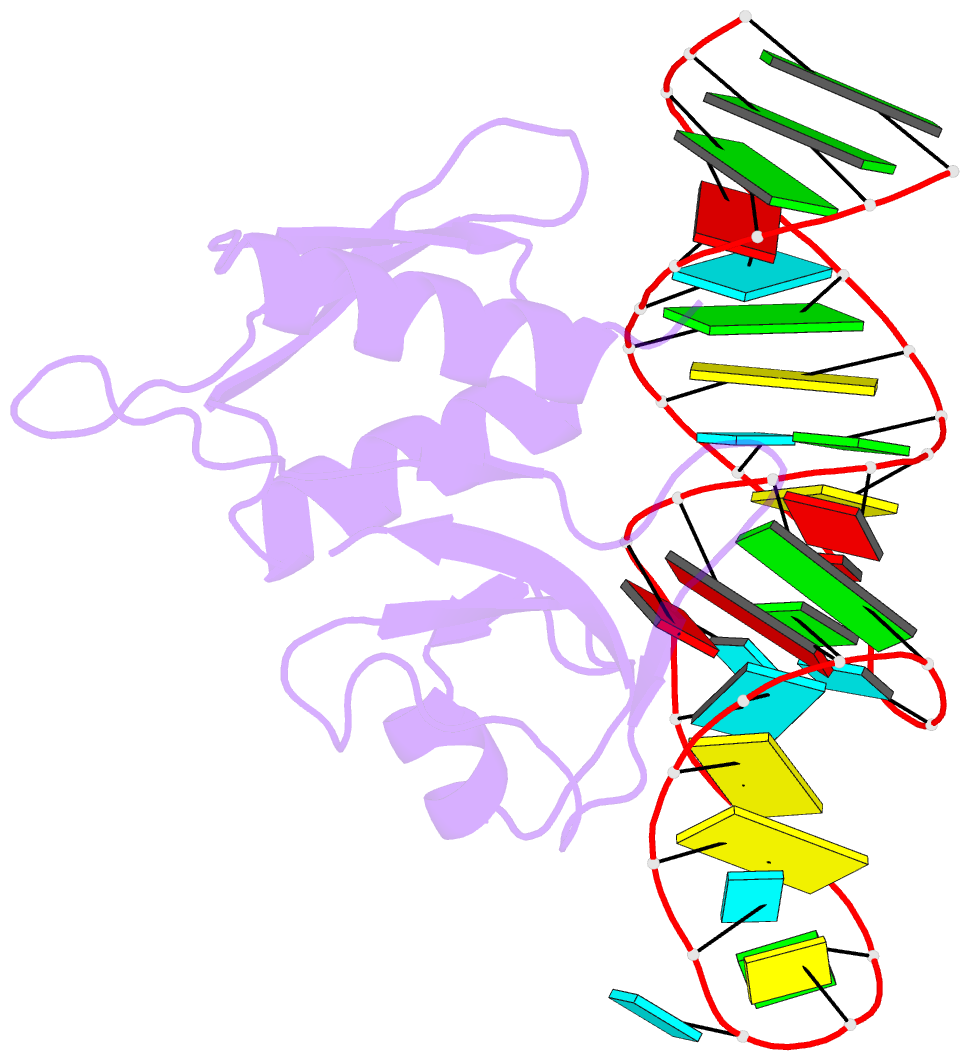
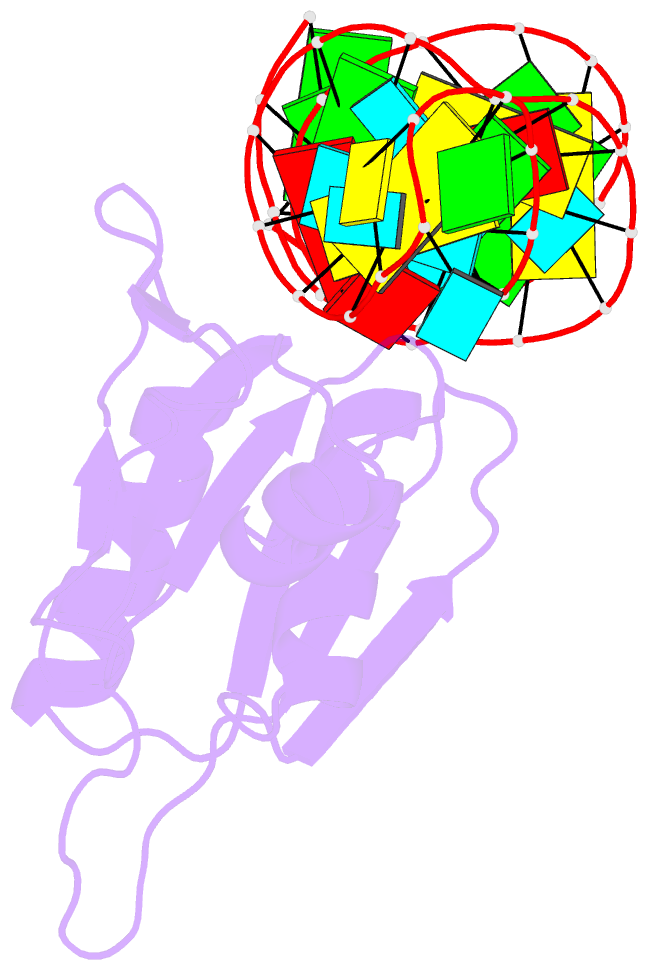
- Crystal structure of bacillus anthracis ribosomal protein s8 in complex with an RNA aptamer
- Reference
- Davlieva M, Donarski J, Wang J, Shamoo Y, Nikonowicz EP (2014): "Structure analysis of free and bound states of an RNA aptamer against ribosomal protein S8 from Bacillus anthracis." Nucleic Acids Res., 42, 10795-10808. doi: 10.1093/nar/gku743.
- Abstract
- Several protein-targeted RNA aptamers have been identified for a variety of applications and although the affinities of numerous protein-aptamer complexes have been determined, the structural details of these complexes have not been widely explored. We examined the structural accommodation of an RNA aptamer that binds bacterial r-protein S8. The core of the primary binding site for S8 on helix 21 of 16S rRNA contains a pair of conserved base triples that mold the sugar-phosphate backbone to S8. The aptamer, which does not contain the conserved sequence motif, is specific for the rRNA binding site of S8. The protein-free RNA aptamer adopts a helical structure with multiple non-canonical base pairs. Surprisingly, binding of S8 leads to a dramatic change in the RNA conformation that restores the signature S8 recognition fold through a novel combination of nucleobase interactions. Nucleotides within the non-canonical core rearrange to create a G-(G-C) triple and a U-(A-U)-U quartet. Although native-like S8-RNA interactions are present in the aptamer-S8 complex, the topology of the aptamer RNA differs from that of the helix 21-S8 complex. This is the first example of an RNA aptamer that adopts substantially different secondary structures in the free and protein-bound states and highlights the remarkable plasticity of RNA secondary structure.