Summary information and primary citation
- PDB-id
- 5d4c; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription-DNA
- Method
- X-ray (3.28 Å)
- Summary
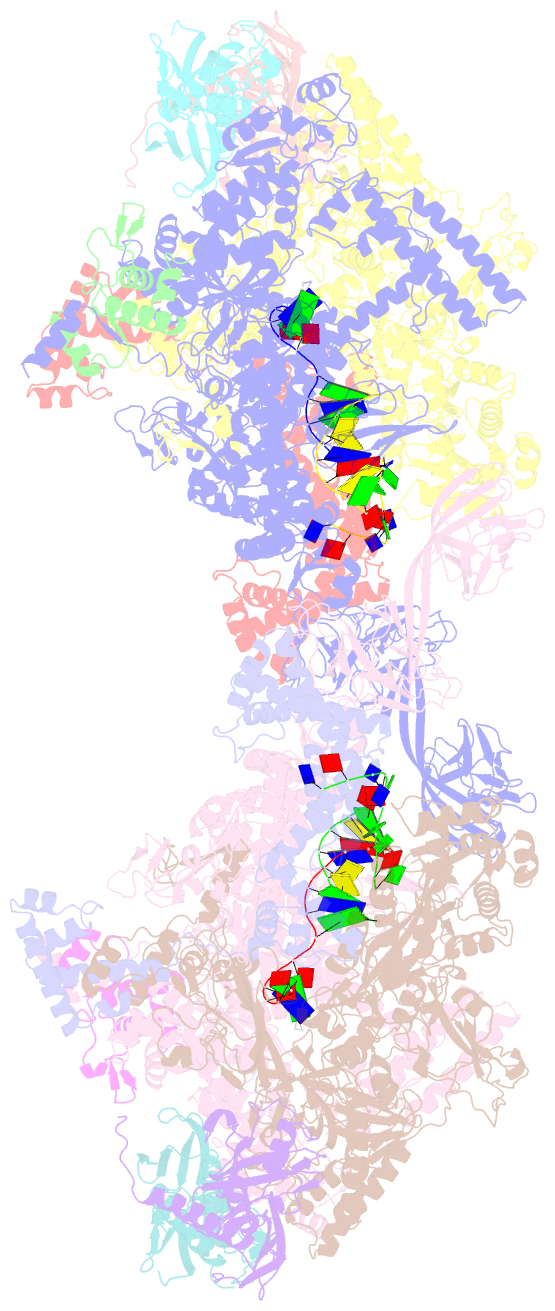
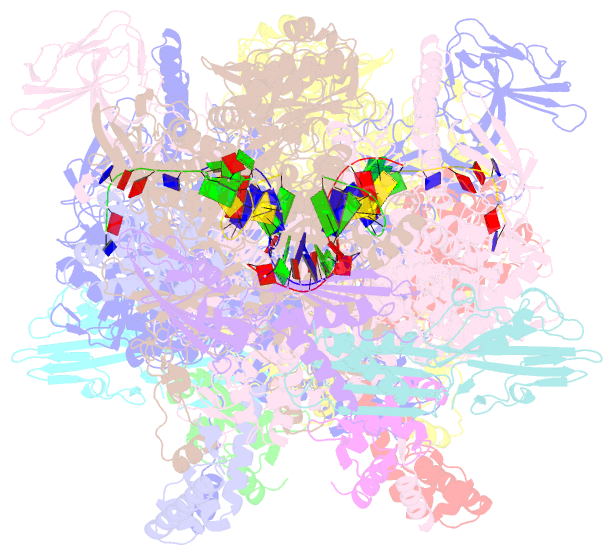
- Crystal structure of thermus thermophilus product complex for transcription initiation with atp and ctp
- Reference
- Bird JG, Zhang Y, Tian Y, Panova N, Barvik I, Greene L, Liu M, Buckley B, Krasny L, Lee JK, Kaplan CD, Ebright RH, Nickels BE (2016): "The mechanism of RNA 5' capping with NAD(+), NADH and desphospho-CoA." Nature, 535, 444-447. doi: 10.1038/nature18622.
- Abstract
- The chemical nature of the 5′ end of RNA is a key determinant of RNA stability, processing, localization and translation efficiency, and has been proposed to provide a layer of ‘epitranscriptomic’ gene regulation. Recently it has been shown that some bacterial RNA species carry a 5′-end structure reminiscent of the 5′ 7-methylguanylate ‘cap’ in eukaryotic RNA. In particular, RNA species containing a 5′-end nicotinamide adenine dinucleotide (NAD+) or 3′-desphospho-coenzyme A (dpCoA) have been identified in both Gram-negative and Gram-positive bacteria. It has been proposed that NAD+, reduced NAD+ (NADH) and dpCoA caps are added to RNA after transcription initiation, in a manner analogous to the addition of 7-methylguanylate caps. Here we show instead that NAD+, NADH and dpCoA are incorporated into RNA during transcription initiation, by serving as non-canonical initiating nucleotides (NCINs) for de novo transcription initiation by cellular RNA polymerase (RNAP). We further show that both bacterial RNAP and eukaryotic RNAP II incorporate NCIN caps, that promoter DNA sequences at and upstream of the transcription start site determine the efficiency of NCIN capping, that NCIN capping occurs in vivo, and that NCIN capping has functional consequences. We report crystal structures of transcription initiation complexes containing NCIN-capped RNA products. Our results define the mechanism and structural basis of NCIN capping, and suggest that NCIN-mediated ‘ab initio capping’ may occur in all organisms.