Summary information and primary citation
- PDB-id
- 5l1z; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription-RNA
- Method
- X-ray (5.9 Å)
- Summary
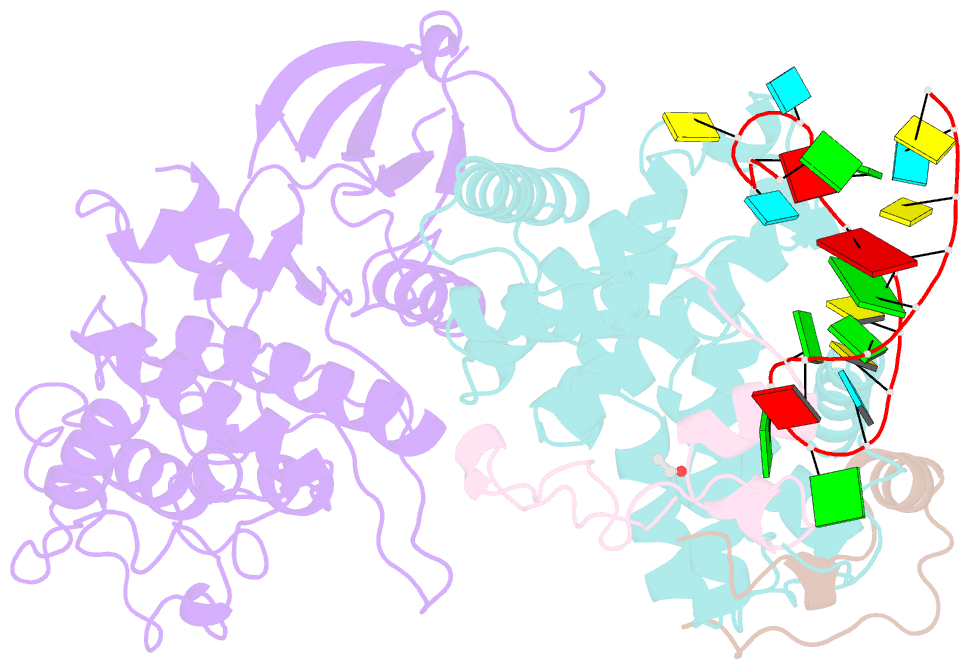
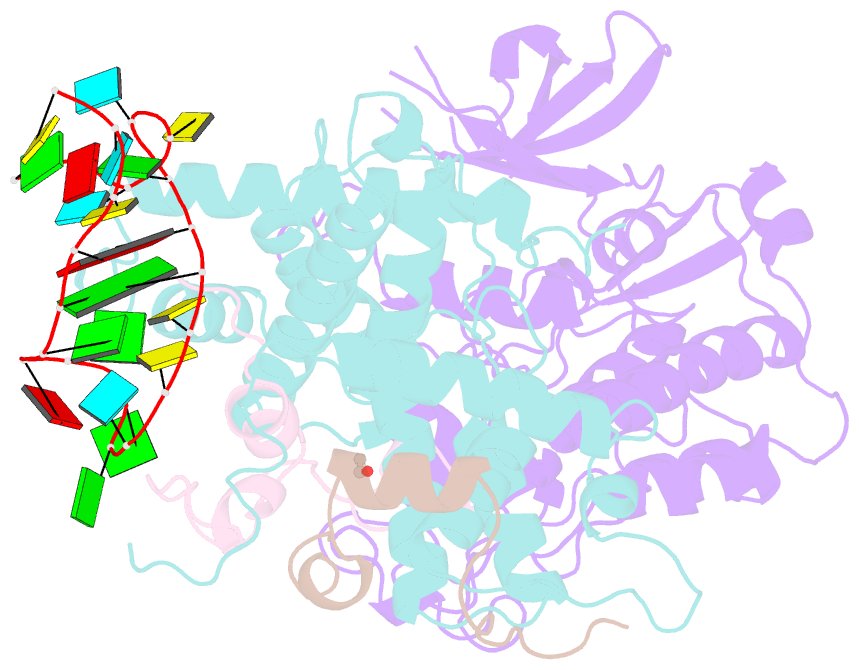
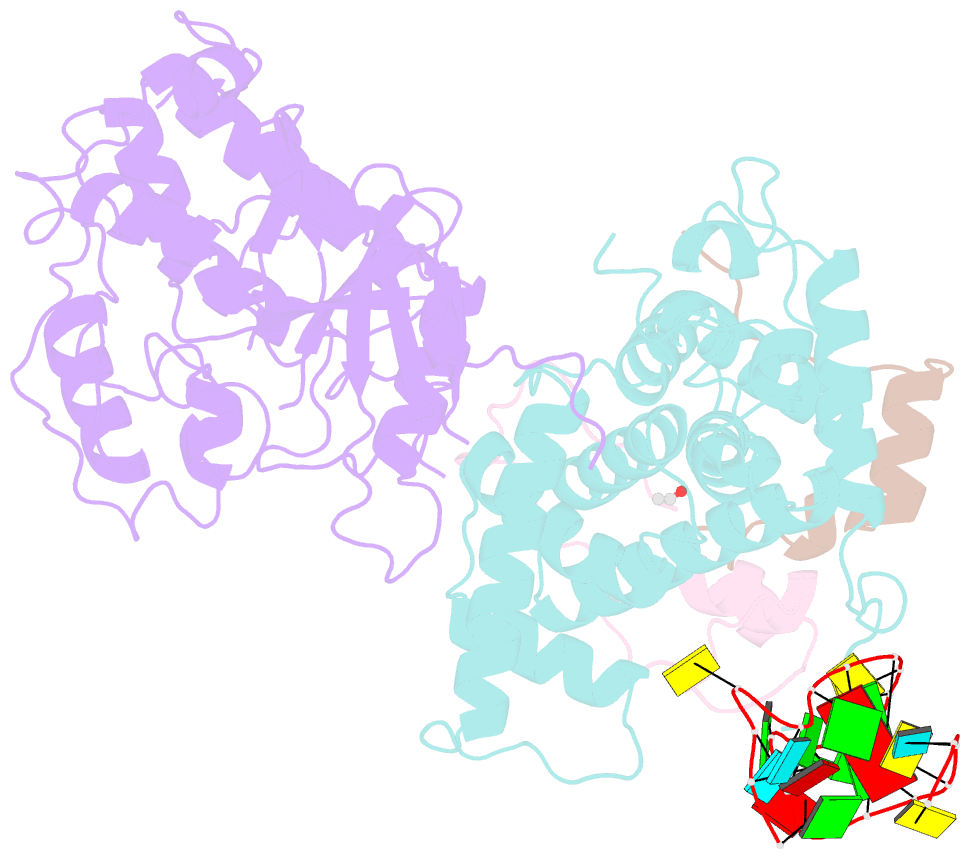
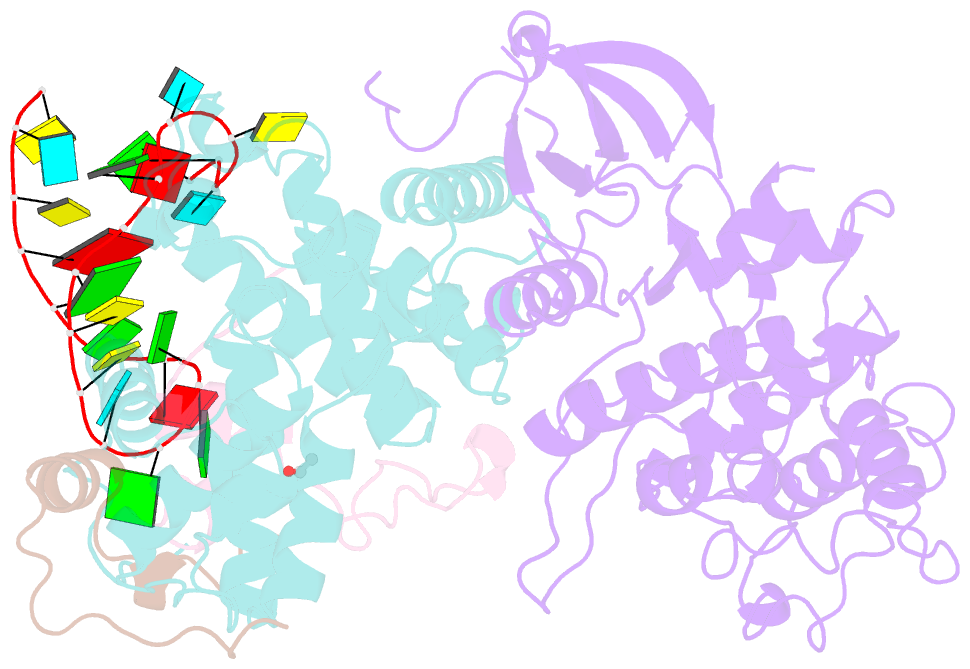
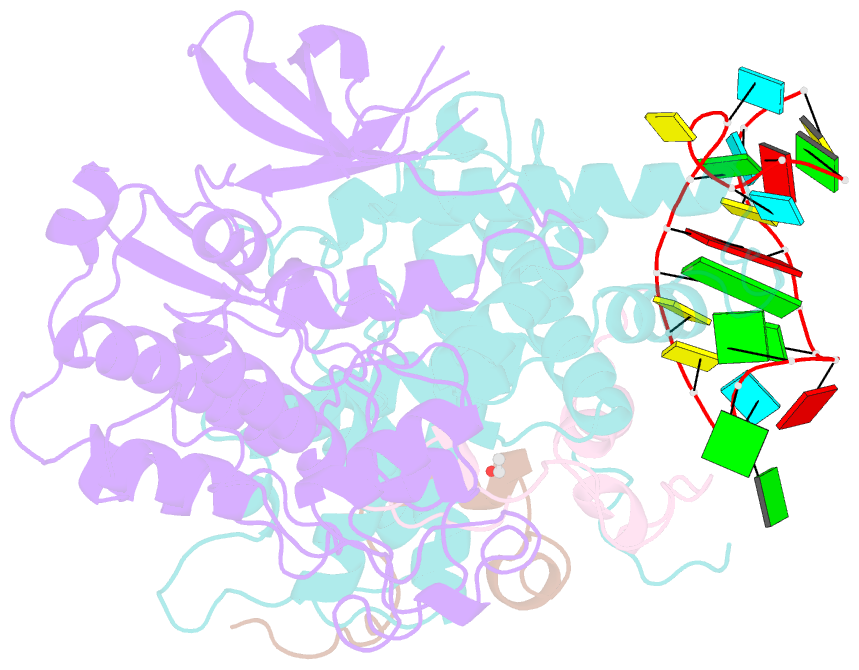
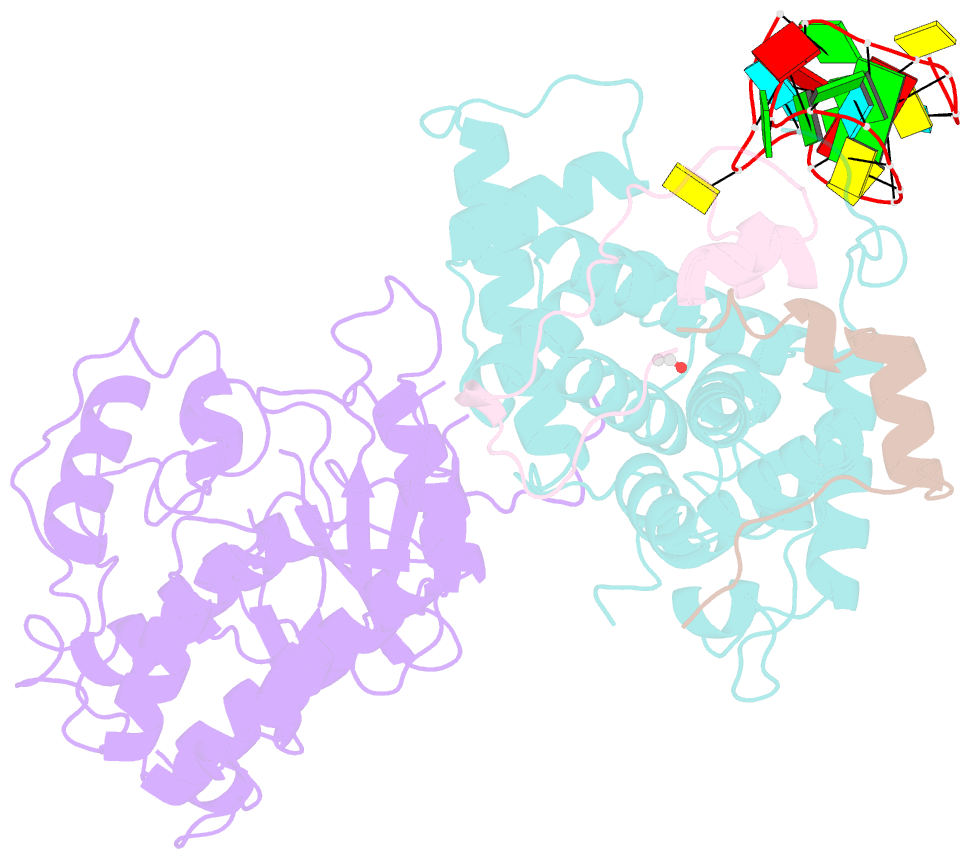
- Tar complex with hiv-1 tat-aff4-p-tefb
- Reference
- Schulze-Gahmen U, Echeverria I, Stjepanovic G, Bai Y, Lu H, Schneidman-Duhovny D, Doudna JA, Zhou Q, Sali A, Hurley JH (2016): "Insights into HIV-1 proviral transcription from integrative structure and dynamics of the Tat:AFF4:P-TEFb:TAR complex." Elife, 5. doi: 10.7554/eLife.15910.
- Abstract
- HIV-1 Tat hijacks the human superelongation complex (SEC) to promote proviral transcription. Here we report the 5.9 Å structure of HIV-1 TAR in complex with HIV-1 Tat and human AFF4, CDK9, and CycT1. The TAR central loop contacts the CycT1 Tat-TAR recognition motif (TRM) and the second Tat Zn2+-binding loop. Hydrogen-deuterium exchange (HDX) shows that AFF4 helix 2 is stabilized in the TAR complex despite not touching the RNA, explaining how it enhances TAR binding to the SEC 50-fold. RNA SHAPE and SAXS data were used to help model the extended (Tat Arginine-Rich Motif) ARM, which enters the TAR major groove between the bulge and the central loop. The structure and functional assays collectively support an integrative structure and a bipartite binding model, wherein the TAR central loop engages the CycT1 TRM and compact core of Tat, while the TAR major groove interacts with the extended Tat ARM.