Summary information and primary citation
- PDB-id
- 5m0j; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein
- Method
- X-ray (2.8 Å)
- Summary
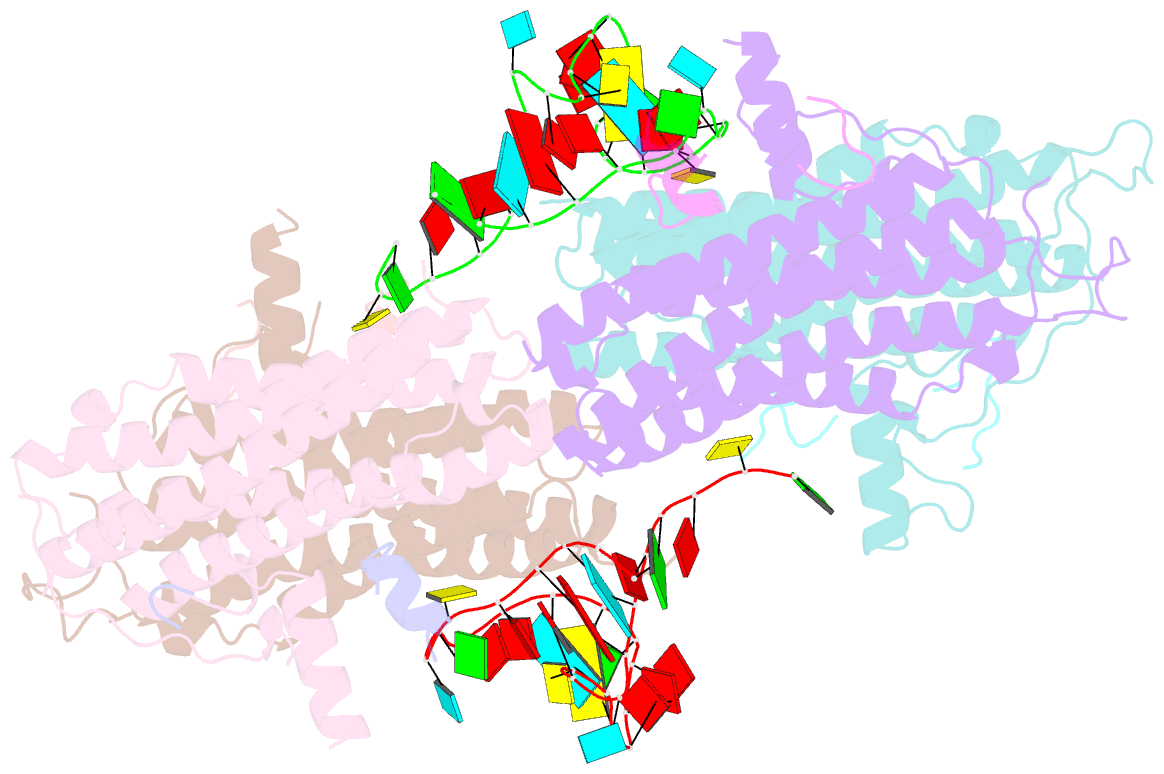
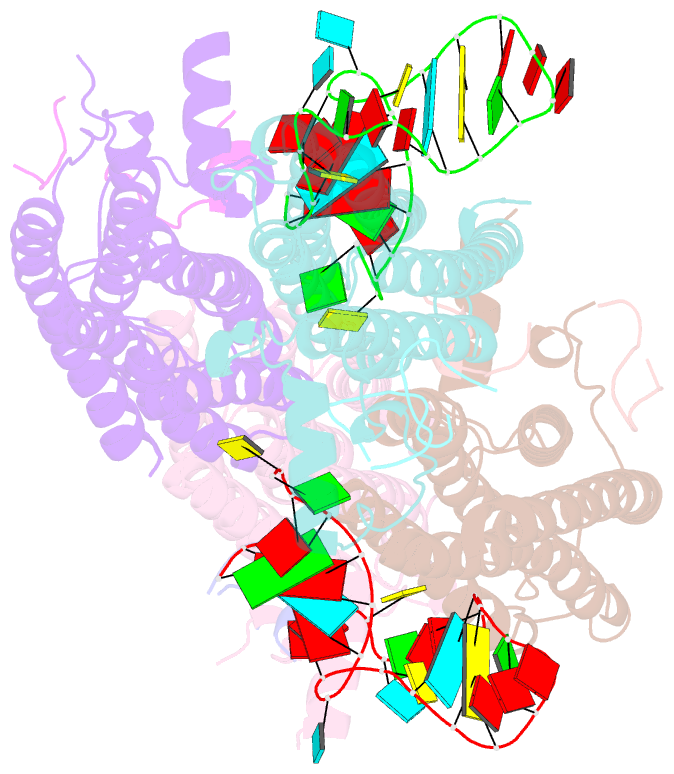
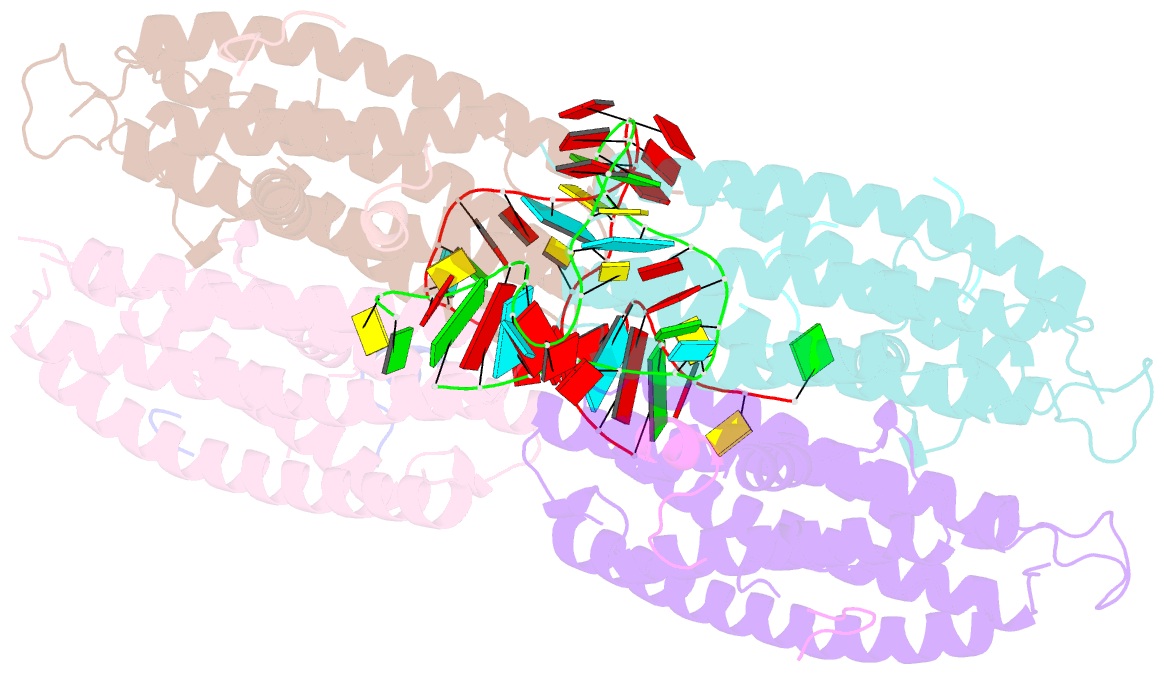
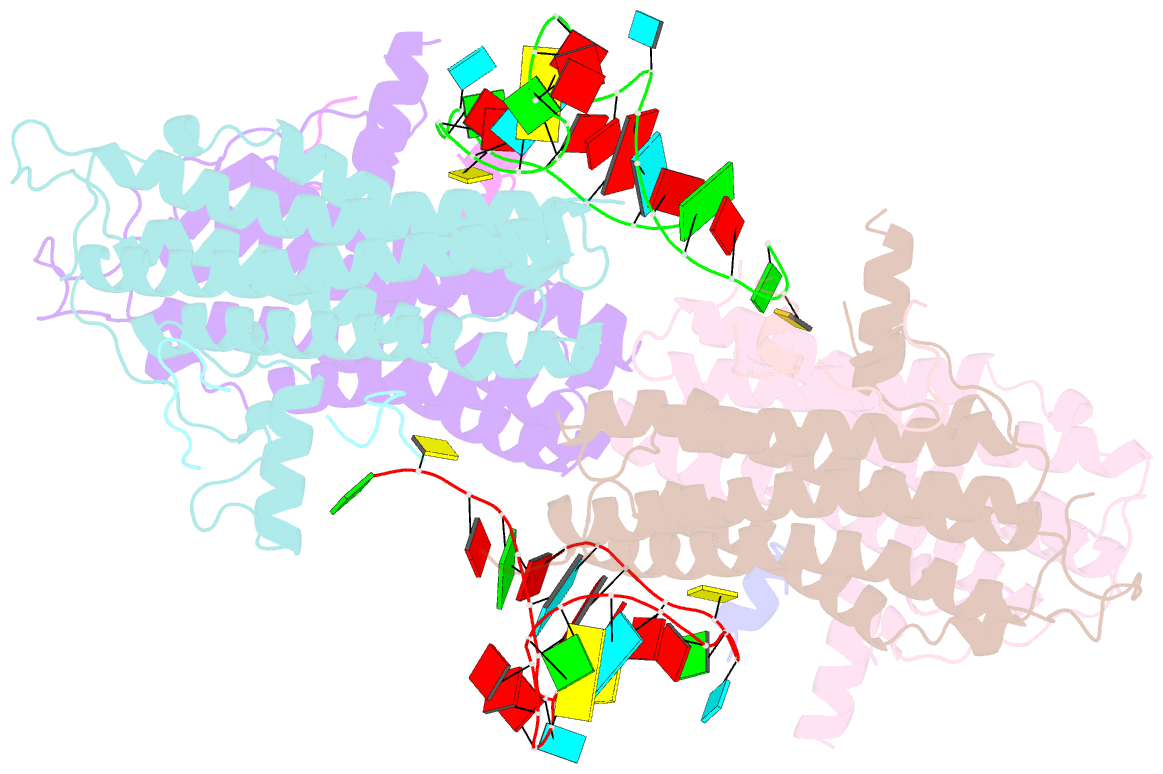
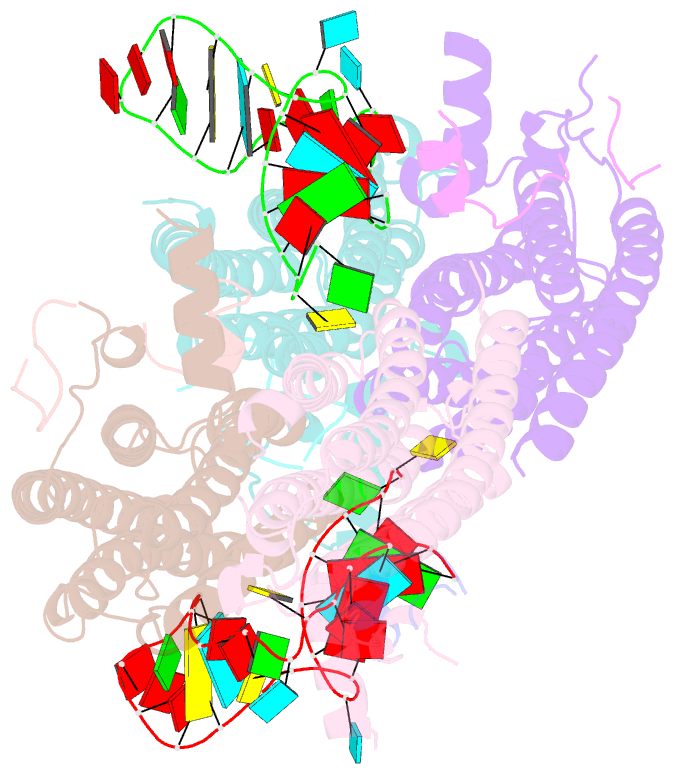
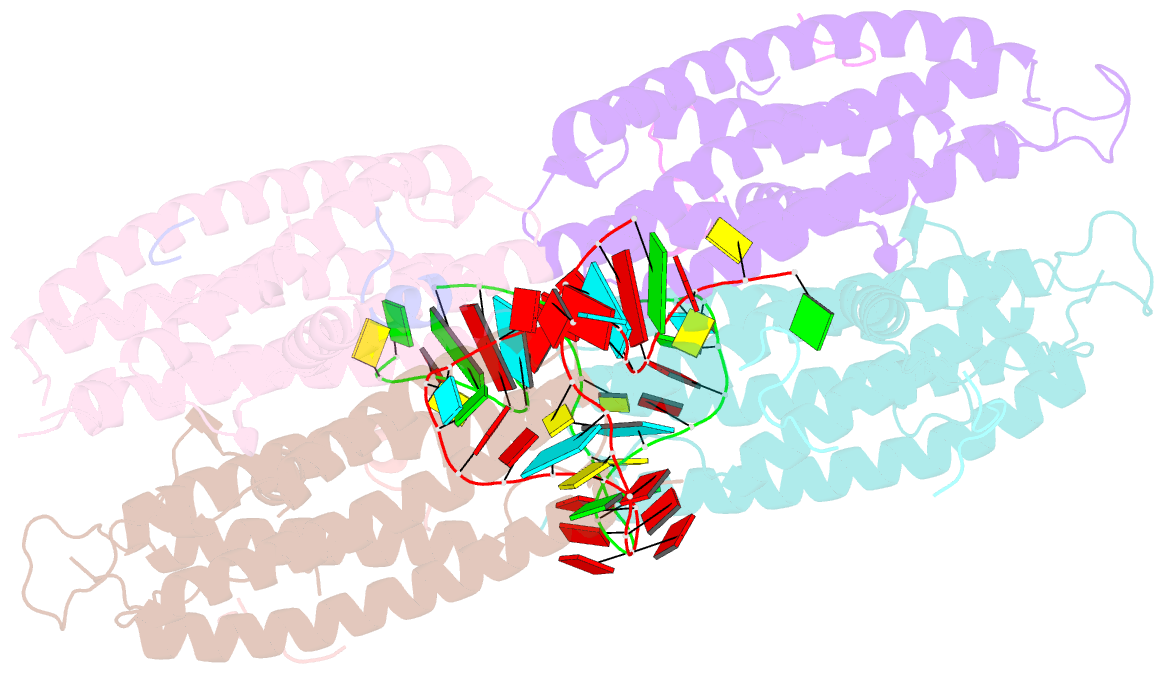
- Crystal structure of the cytoplasmic complex with she2p, she3p, and the ash1 mrna e3-localization element
- Reference
- Edelmann FT, Schlundt A, Heym RG, Jenner A, Niedner-Boblenz A, Syed MI, Paillart JC, Stehle R, Janowski R, Sattler M, Jansen RP, Niessing D (2017): "Molecular architecture and dynamics of ASH1 mRNA recognition by its mRNA-transport complex." Nat. Struct. Mol. Biol., 24, 152-161. doi: 10.1038/nsmb.3351.
- Abstract
- mRNA localization is an essential mechanism of gene regulation and is required for processes such as stem-cell division, embryogenesis and neuronal plasticity. It is not known which features in the cis-acting mRNA localization elements (LEs) are specifically recognized by motor-containing transport complexes. To the best of our knowledge, no high-resolution structure is available for any LE in complex with its cognate protein complex. Using X-ray crystallography and complementary techniques, we carried out a detailed assessment of an LE of the ASH1 mRNA from yeast, its complex with its shuttling RNA-binding protein She2p, and its highly specific, cytoplasmic complex with She3p. Although the RNA alone formed a flexible stem loop, She2p binding induced marked conformational changes. However, only joining by the unstructured She3p resulted in specific RNA recognition. The notable RNA rearrangements and joint action of a globular and an unfolded RNA-binding protein offer unprecedented insights into the step-wise maturation of an mRNA-transport complex.