Summary information and primary citation
- PDB-id
- 5mq0; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- splicing
- Method
- cryo-EM (4.17 Å)
- Summary
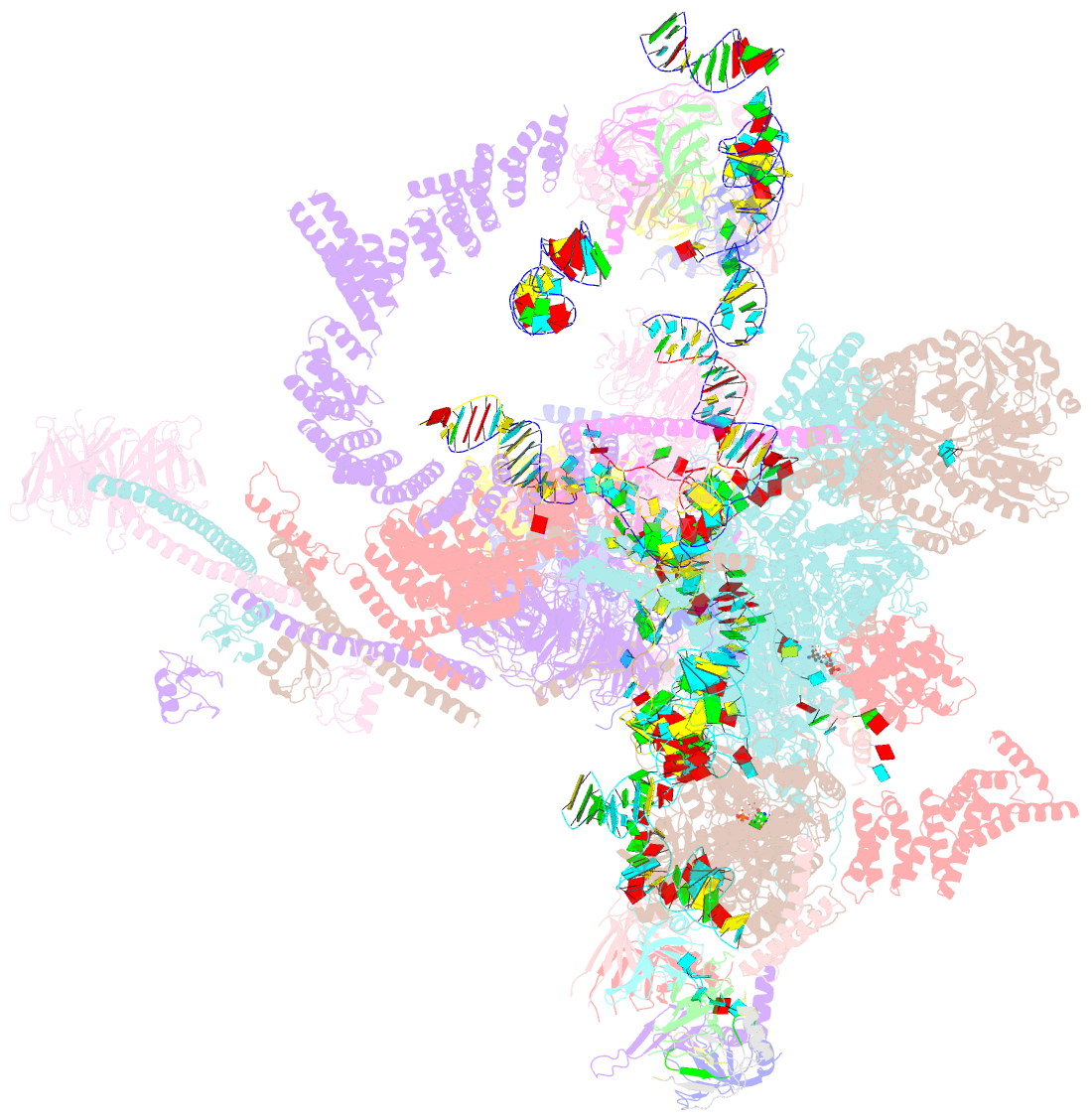
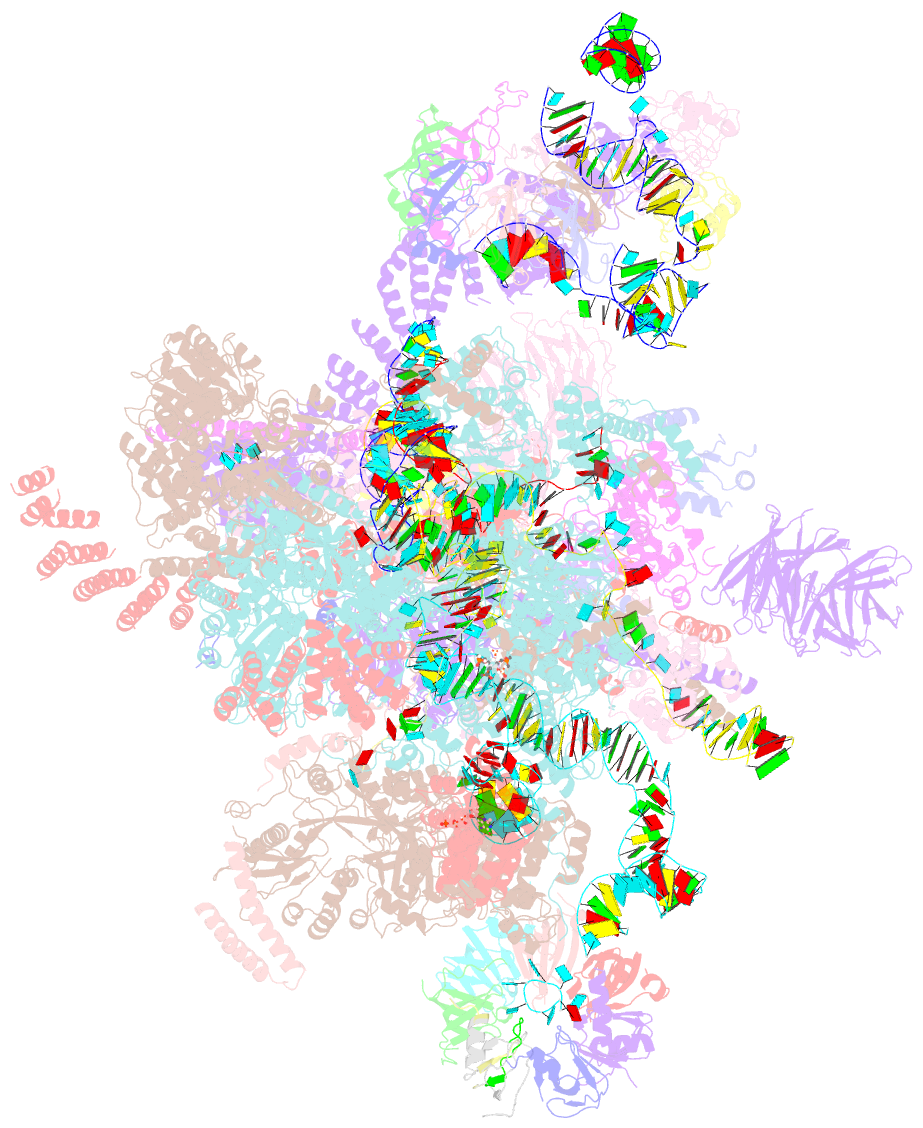
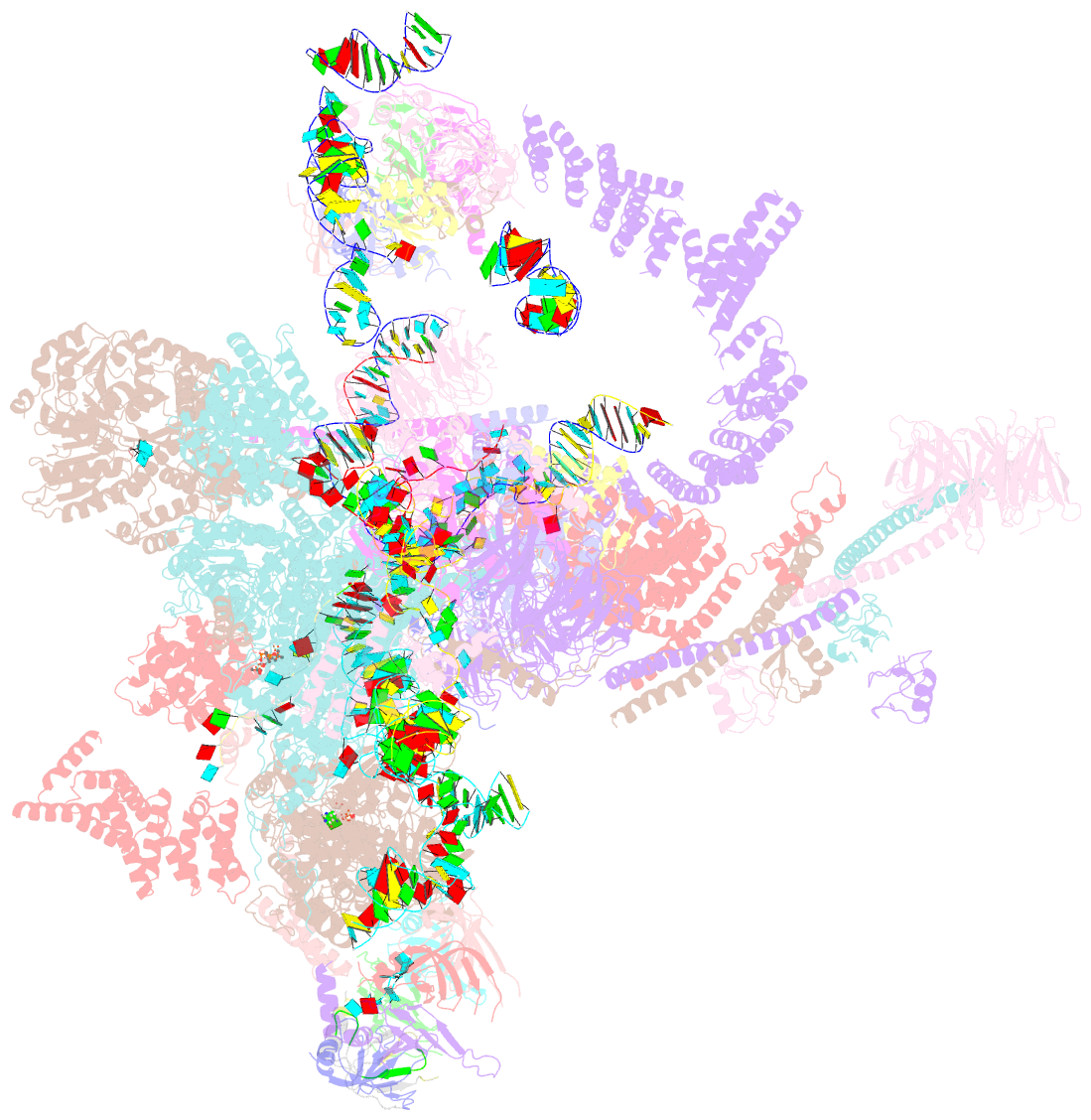
- Structure of a spliceosome remodeled for exon ligation
- Reference
- Fica SM, Oubridge C, Galej WP, Wilkinson ME, Bai XC, Newman AJ, Nagai K (2017): "Structure of a spliceosome remodelled for exon ligation." Nature, 542, 377-380. doi: 10.1038/nature21078.
- Abstract
- The spliceosome excises introns from pre-mRNAs in two sequential transesterifications-branching and exon ligation-catalysed at a single catalytic metal site in U6 small nuclear RNA (snRNA). Recently reported structures of the spliceosomal C complex with the cleaved 5' exon and lariat-3'-exon bound to the catalytic centre revealed that branching-specific factors such as Cwc25 lock the branch helix into position for nucleophilic attack of the branch adenosine at the 5' splice site. Furthermore, the ATPase Prp16 is positioned to bind and translocate the intron downstream of the branch point to destabilize branching-specific factors and release the branch helix from the active site. Here we present, at 3.8 Å resolution, the cryo-electron microscopy structure of a Saccharomyces cerevisiae spliceosome stalled after Prp16-mediated remodelling but before exon ligation. While the U6 snRNA catalytic core remains firmly held in the active site cavity of Prp8 by proteins common to both steps, the branch helix has rotated by 75° compared to the C complex and is stabilized in a new position by Prp17, Cef1 and the reoriented Prp8 RNase H-like domain. This rotation of the branch helix removes the branch adenosine from the catalytic core, creates a space for 3' exon docking, and restructures the pairing of the 5' splice site with the U6 snRNA ACAGAGA region. Slu7 and Prp18, which promote exon ligation, bind together to the Prp8 RNase H-like domain. The ATPase Prp22, bound to Prp8 in place of Prp16, could interact with the 3' exon, suggesting a possible basis for mRNA release after exon ligation. Together with the structure of the C complex, our structure of the C* complex reveals the two major conformations of the spliceosome during the catalytic stages of splicing.