Summary information and primary citation
- PDB-id
- 5msg; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- viral protein
- Method
- X-ray (3.8 Å)
- Summary
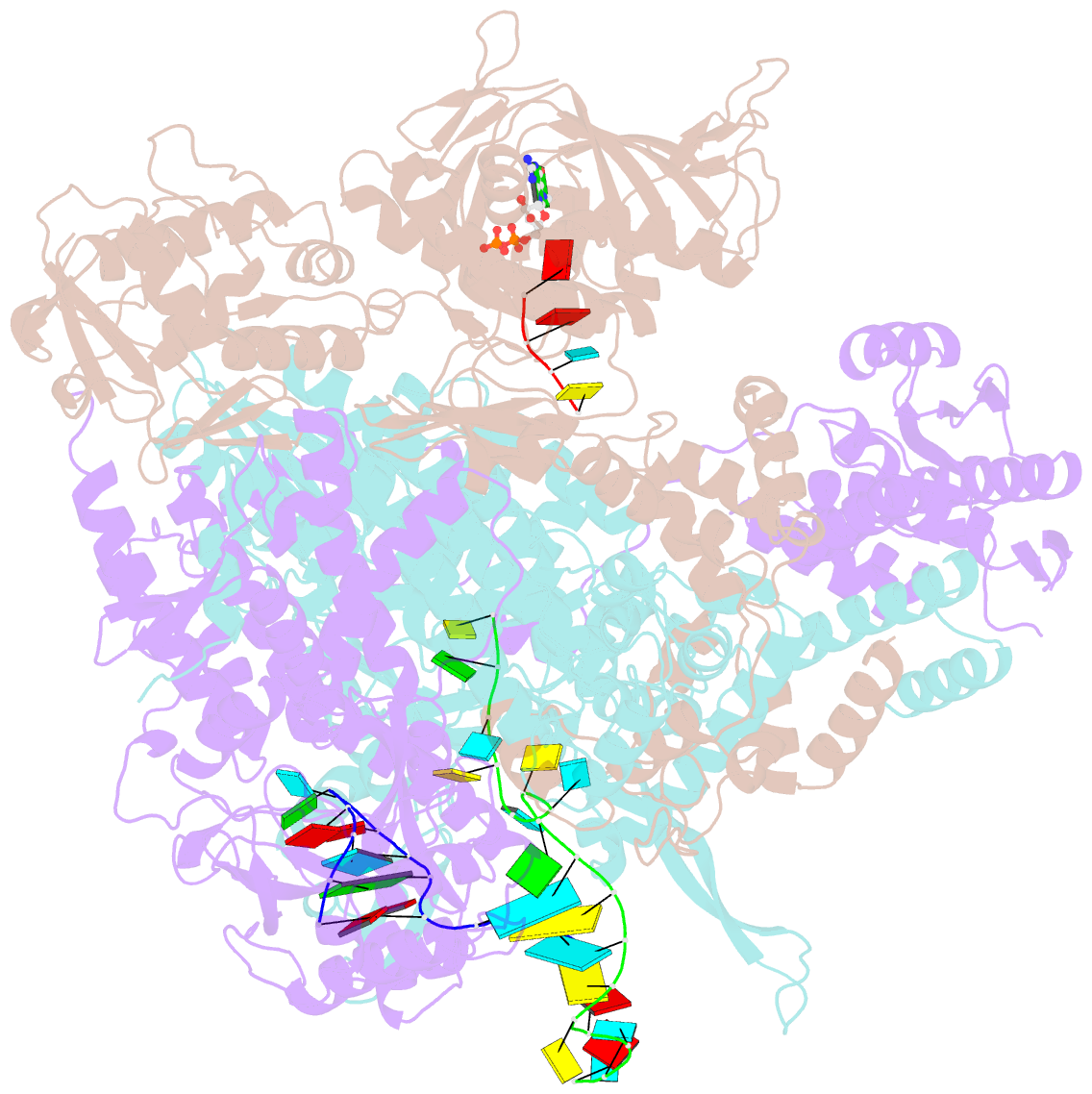
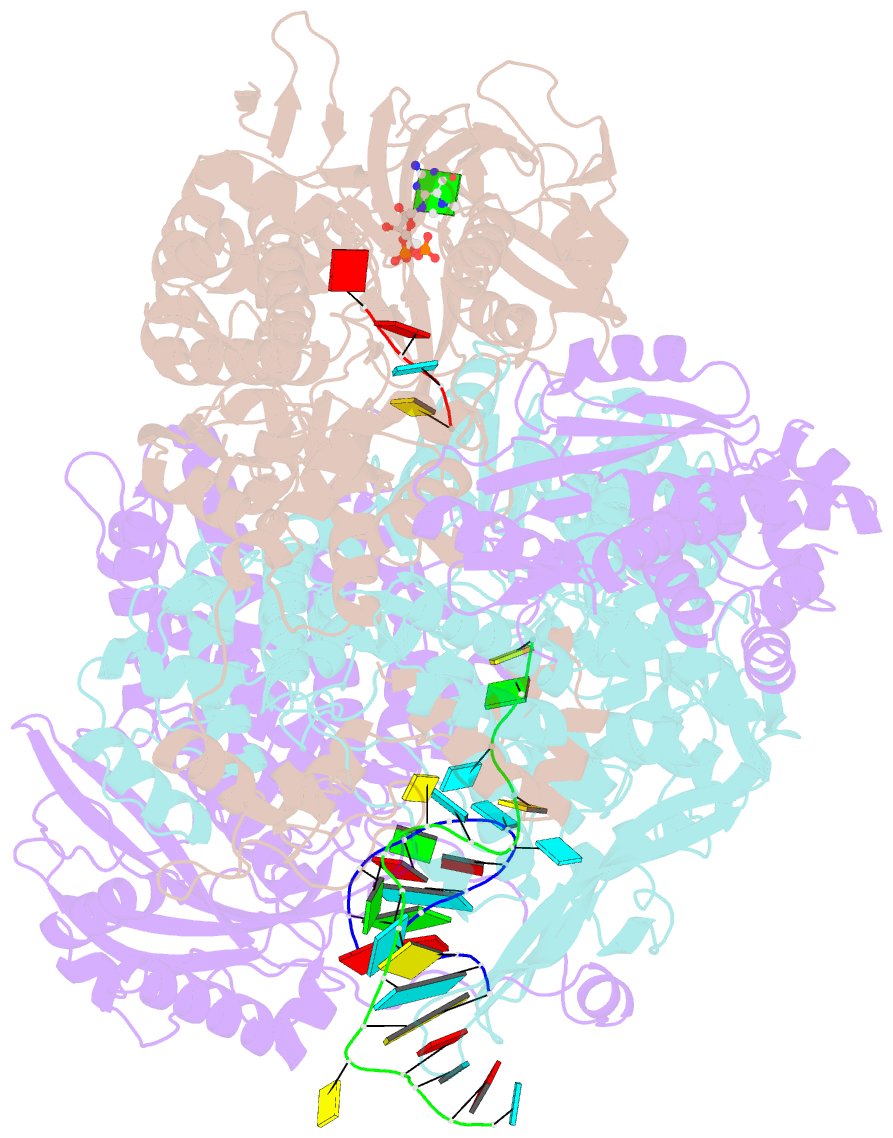
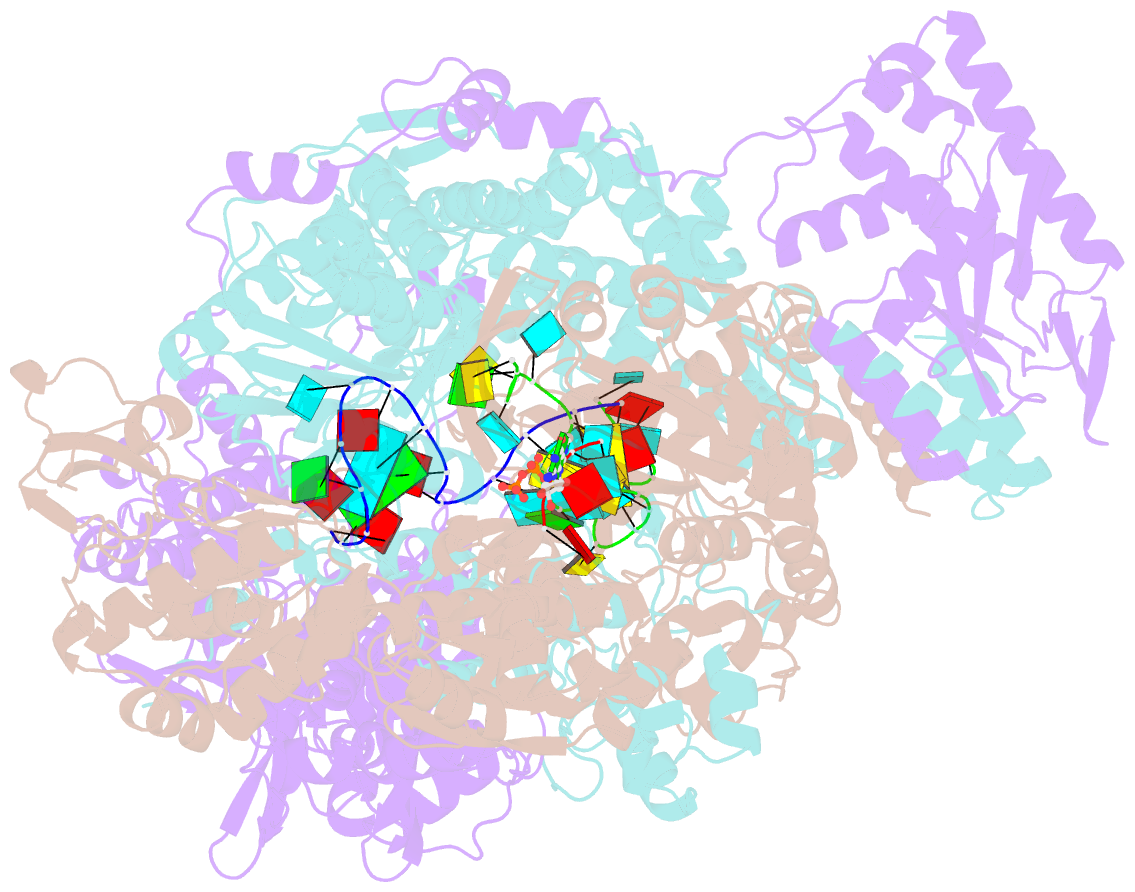
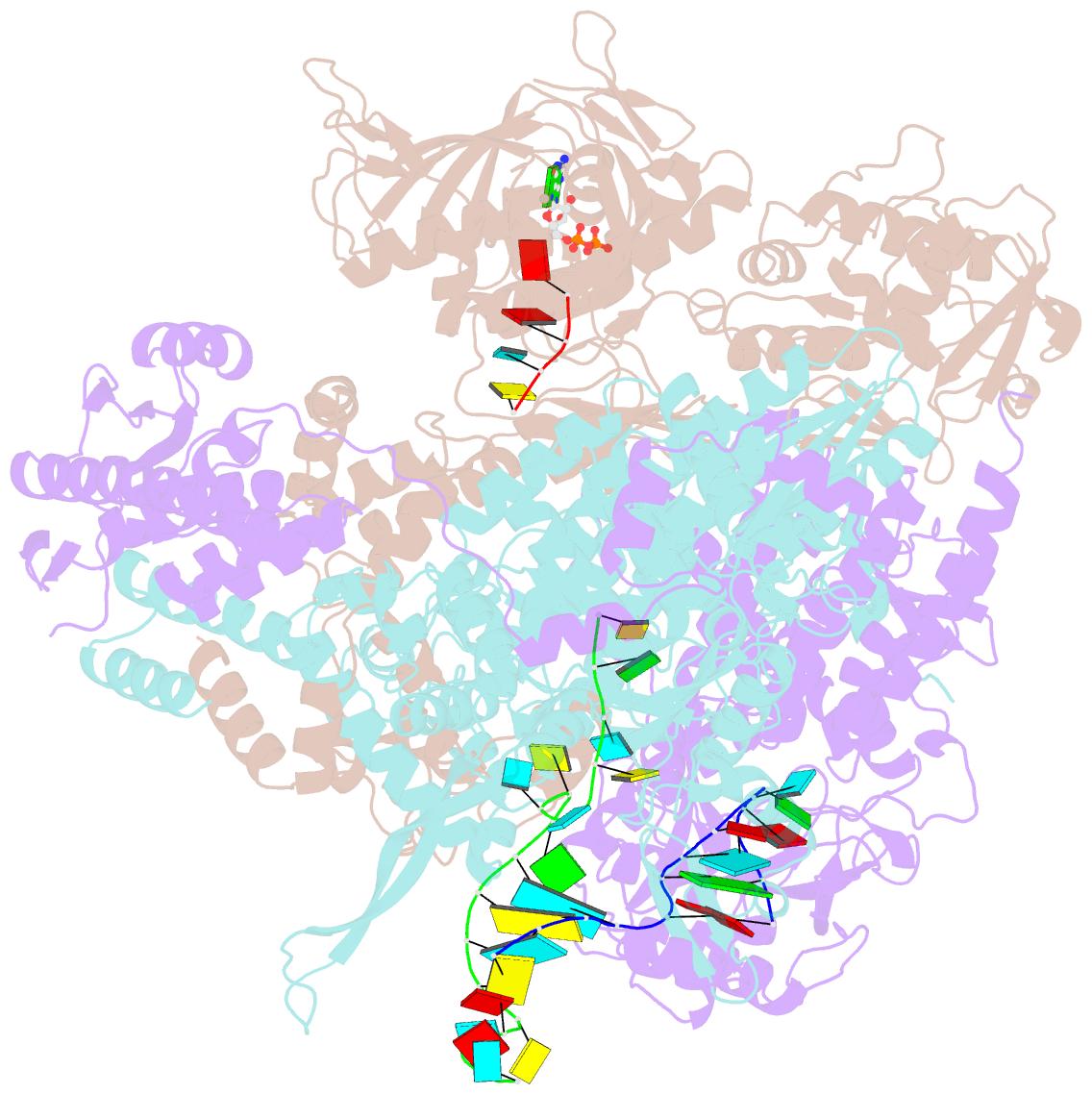
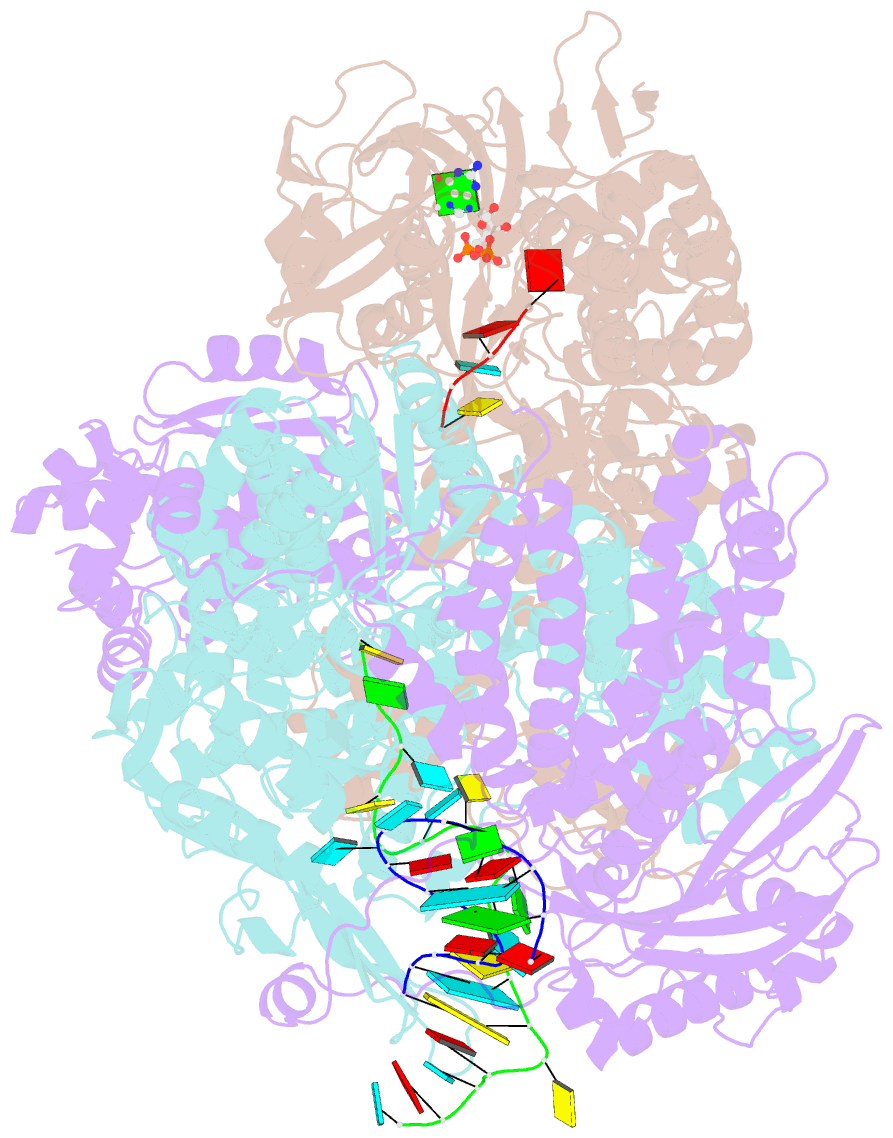
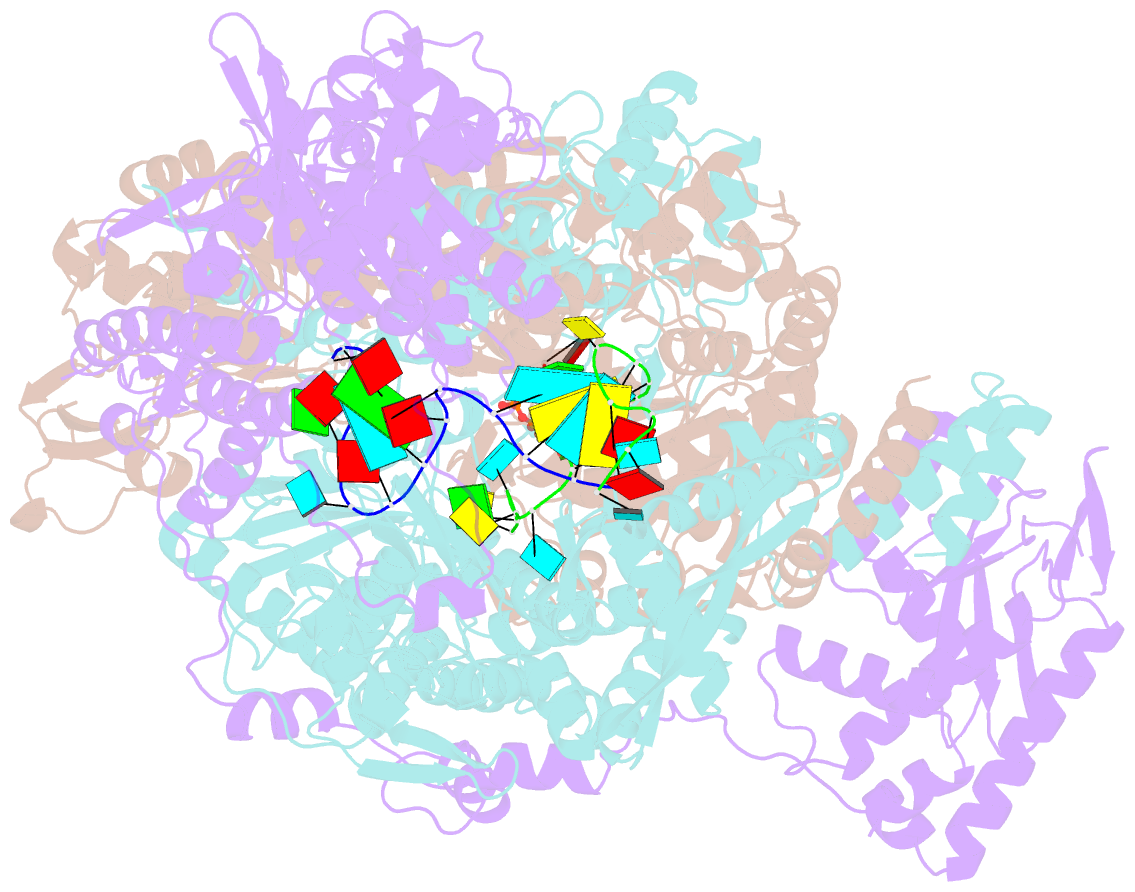
- Influenza b polymerase bound to vrna promoter and capped RNA primer
- Reference
- Reich S, Guilligay D, Cusack S (2017): "An in vitro fluorescence based study of initiation of RNA synthesis by influenza B polymerase." Nucleic Acids Res., 45, 3353-3368. doi: 10.1093/nar/gkx043.
- Abstract
- Influenza polymerase replicates, via a complementary RNA intermediate (cRNA), and transcribes the eight viral RNA (vRNA) genome segments. To initiate RNA synthesis it is bound to the conserved 5΄ and 3΄ extremities of the vRNA or cRNA (the 'promoter'). 5΄-3΄ base-pairing in the distal promoter region is essential to position the template RNA at the polymerase active site, as shown by a new crystal structure with the 3΄ end threading through the template entry tunnel. We develop fluorescence polarization assays to quantify initiation of cap-primed (transcription) or unprimed (replication) RNA synthesis by recombinant influenza B polymerase bound to the vRNA or cRNA promoter. The rate-limiting step is formation of a primed initiation complex with minimally ApG required to stabilize the 3΄ end of the template within the active-site. Polymerase bound to the vRNA promoter initiates RNA synthesis terminally, while the cRNA promoter directs internal initiation at a significantly lower rate. Progression to elongation requires breaking the promoter 5΄-3΄ base-pairing region and favourable compensation by the emerging template-product base-pairs. The RNA synthesis assay is adaptable to high-throughput screening for polymerase inhibitors. In a pilot study, we find that initiation at the cRNA promoter is unusually susceptible to inhibition by 2΄F-2΄dNTPs.