Summary information and primary citation
- PDB-id
- 5v05; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydrolase-DNA
- Method
- X-ray (2.902 Å)
- Summary
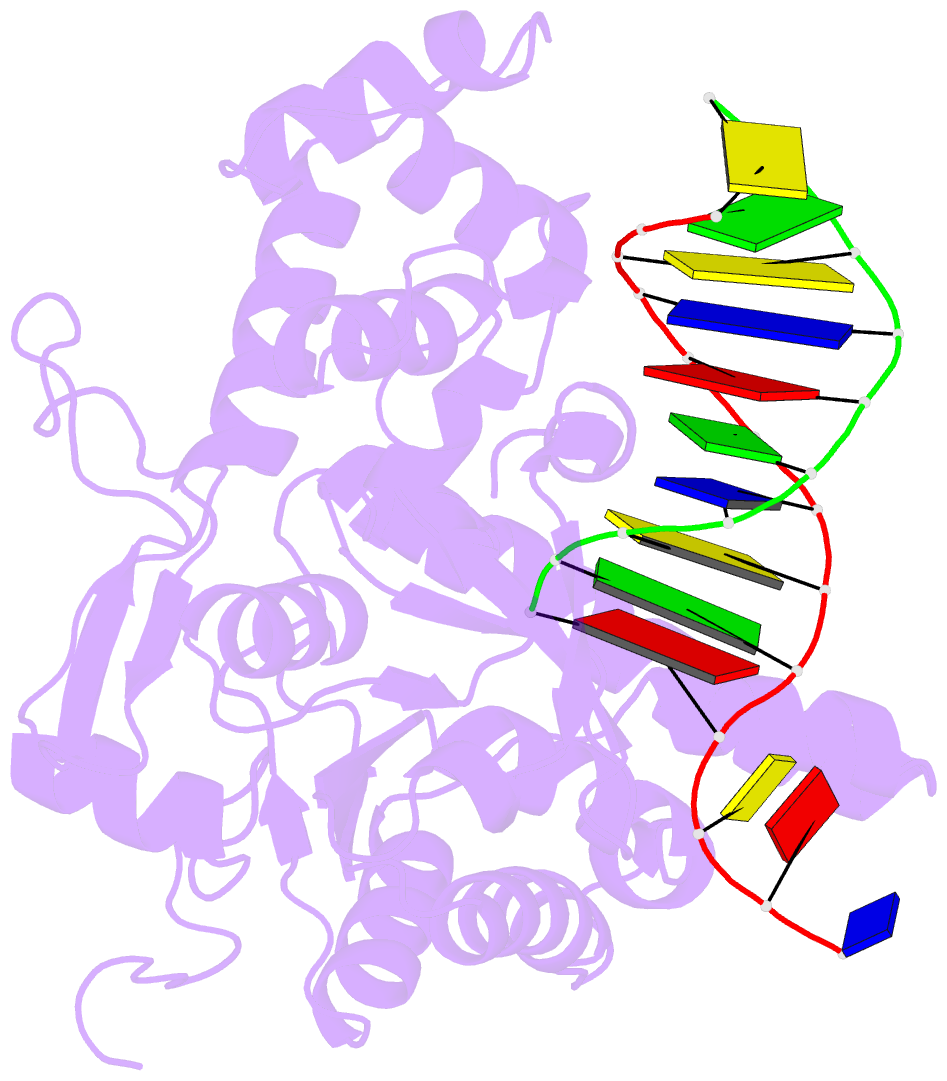
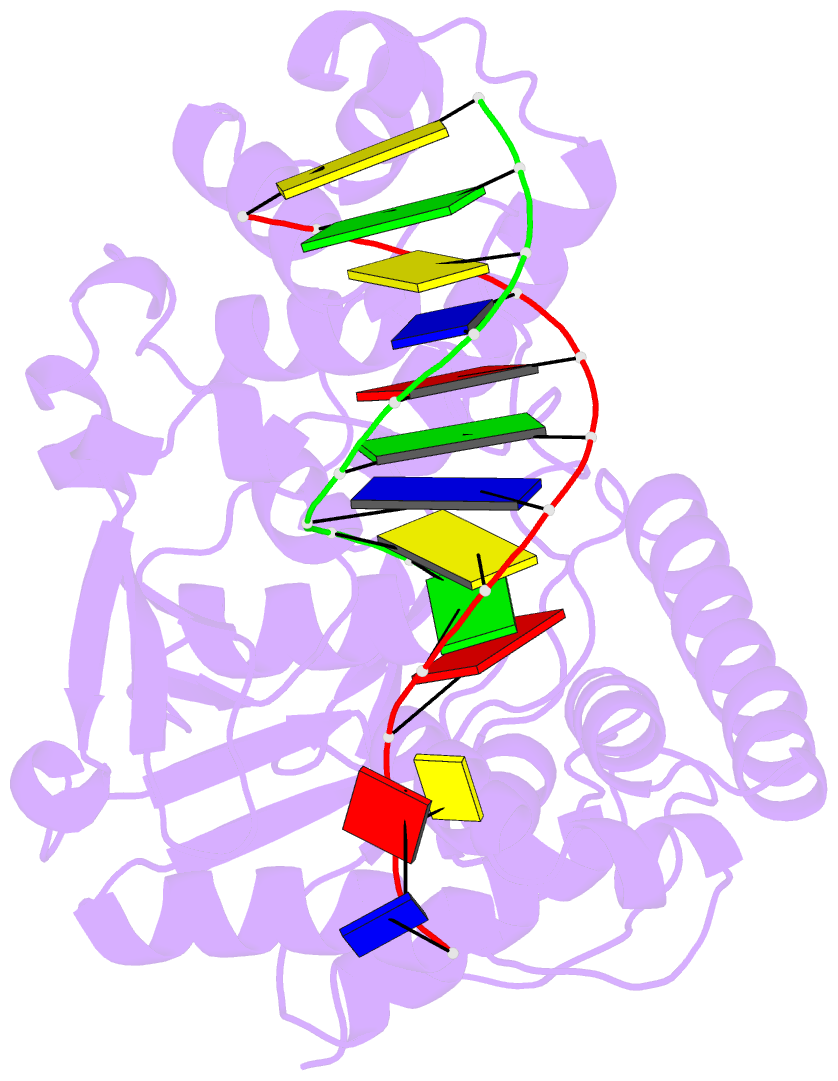
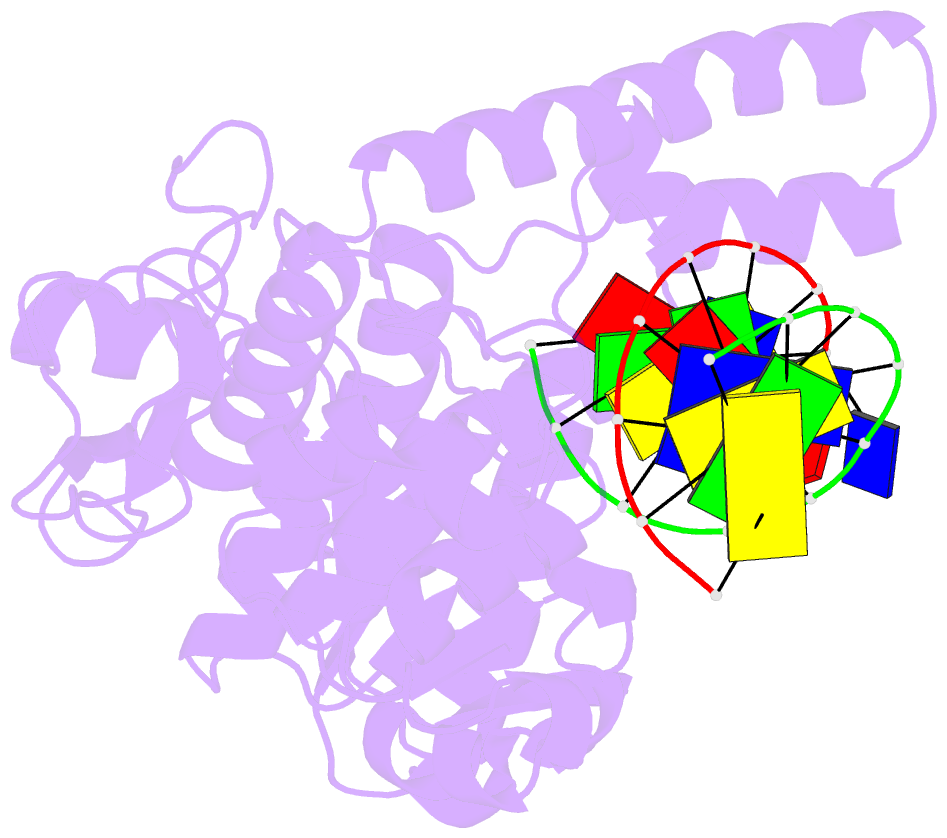
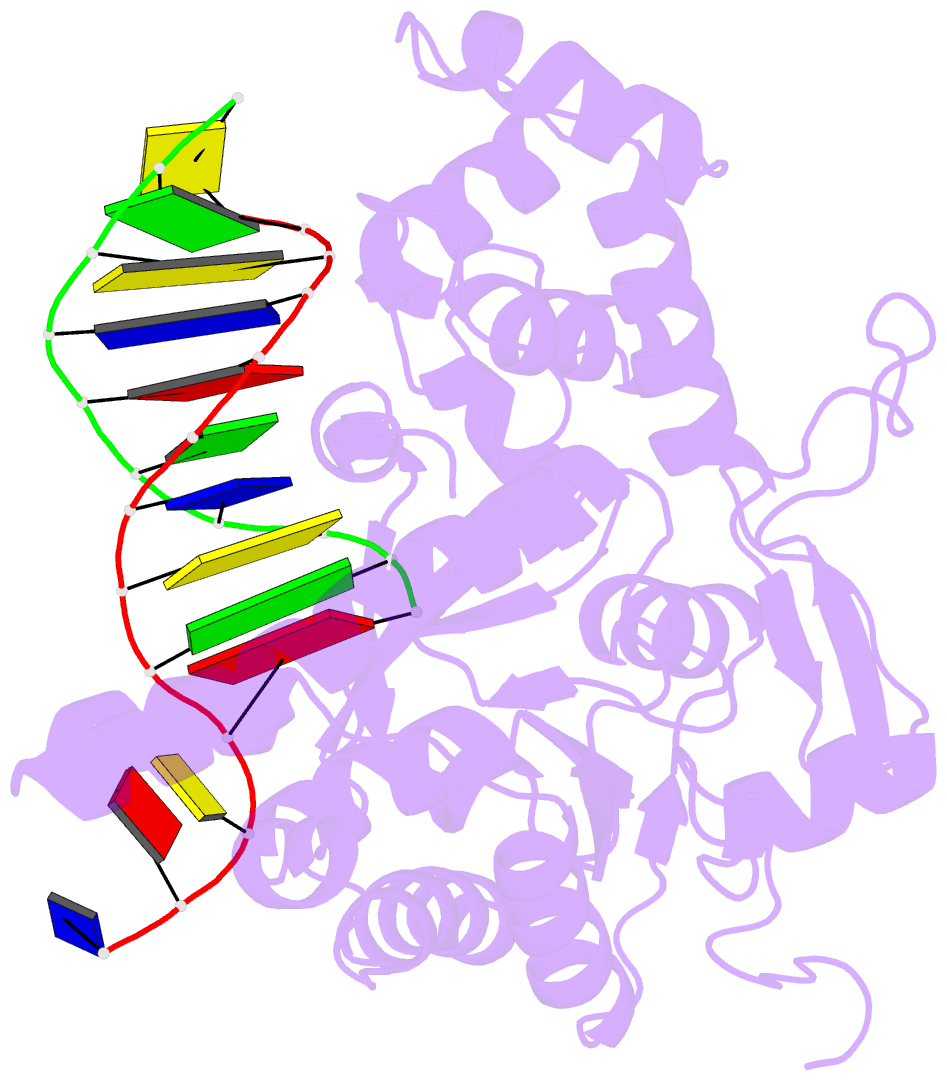
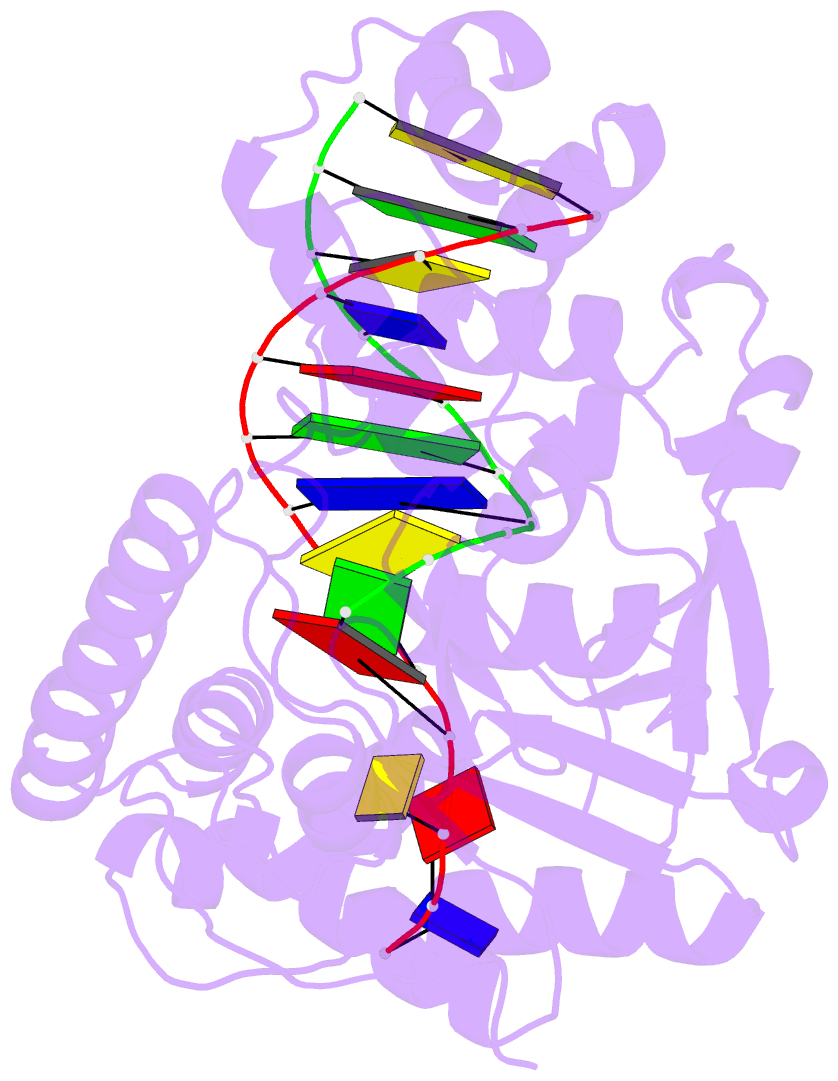
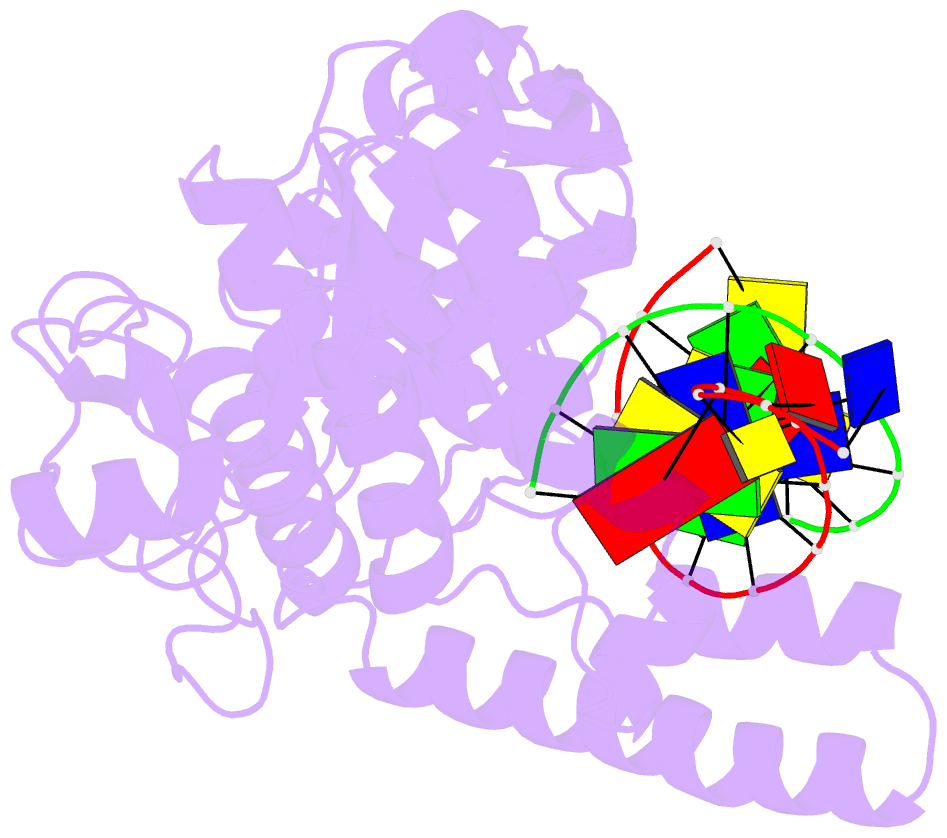
- Crystal structure of human exonuclease 1 exo1 (wt) in complex with 5' recessed-end DNA (riii)
- Reference
- Shi Y, Hellinga HW, Beese LS (2017): "Interplay of catalysis, fidelity, threading, and processivity in the exo- and endonucleolytic reactions of human exonuclease I." Proc. Natl. Acad. Sci. U.S.A., 114, 6010-6015. doi: 10.1073/pnas.1704845114.
- Abstract
- Human exonuclease 1 (hExo1) is a member of the RAD2/XPG structure-specific 5'-nuclease superfamily. Its dominant, processive 5'-3' exonuclease and secondary 5'-flap endonuclease activities participate in various DNA repair, recombination, and replication processes. A single active site processes both recessed ends and 5'-flap substrates. By initiating enzyme reactions in crystals, we have trapped hExo1 reaction intermediates that reveal structures of these substrates before and after their exo- and endonucleolytic cleavage, as well as structures of uncleaved, unthreaded, and partially threaded 5' flaps. Their distinctive 5' ends are accommodated by a small, mobile arch in the active site that binds recessed ends at its base and threads 5' flaps through a narrow aperture within its interior. A sequence of successive, interlocking conformational changes guides the two substrate types into a shared reaction mechanism that catalyzes their cleavage by an elaborated variant of the two-metal, in-line hydrolysis mechanism. Coupling of substrate-dependent arch motions to transition-state stabilization suppresses inappropriate or premature cleavage, enhancing processing fidelity. The striking reduction in flap conformational entropy is catalyzed, in part, by arch motions and transient binding interactions between the flap and unprocessed DNA strand. At the end of the observed reaction sequence, hExo1 resets without relinquishing DNA binding, suggesting a structural basis for its processivity.