Summary information and primary citation
- PDB-id
- 5wc9; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein
- Method
- X-ray (3.15 Å)
- Summary
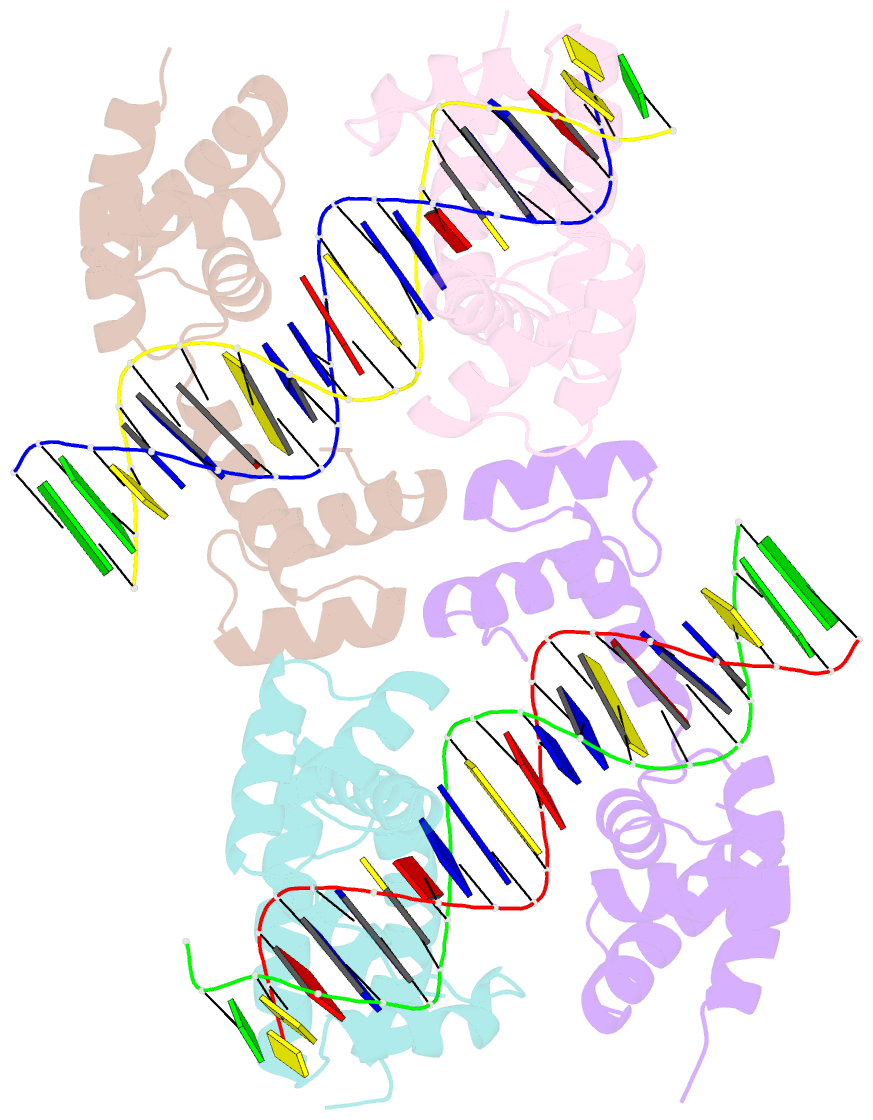
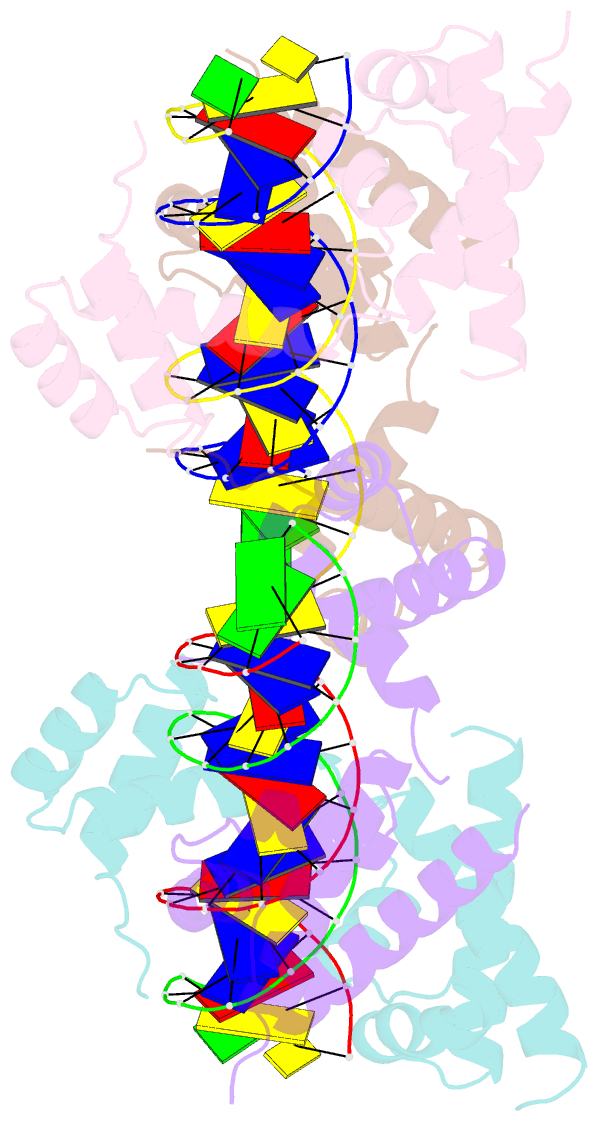
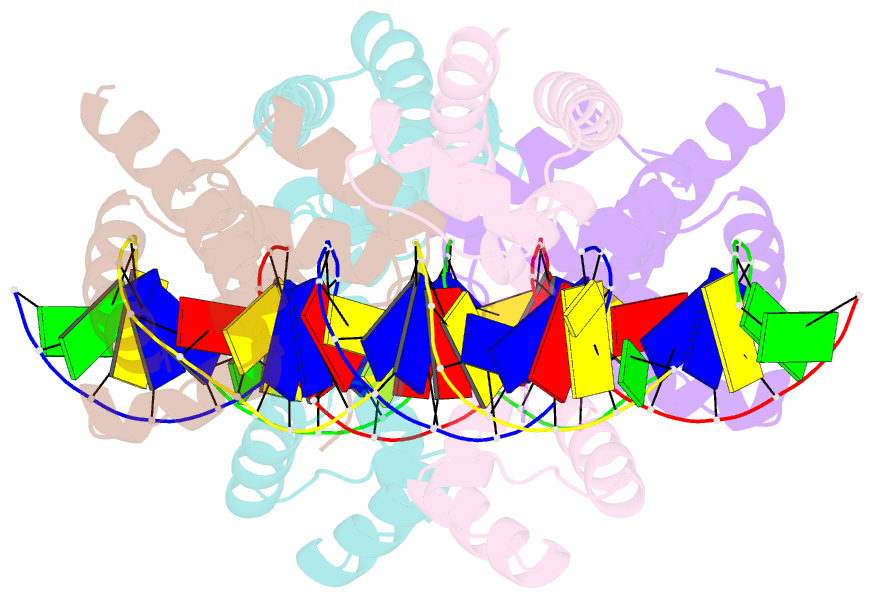
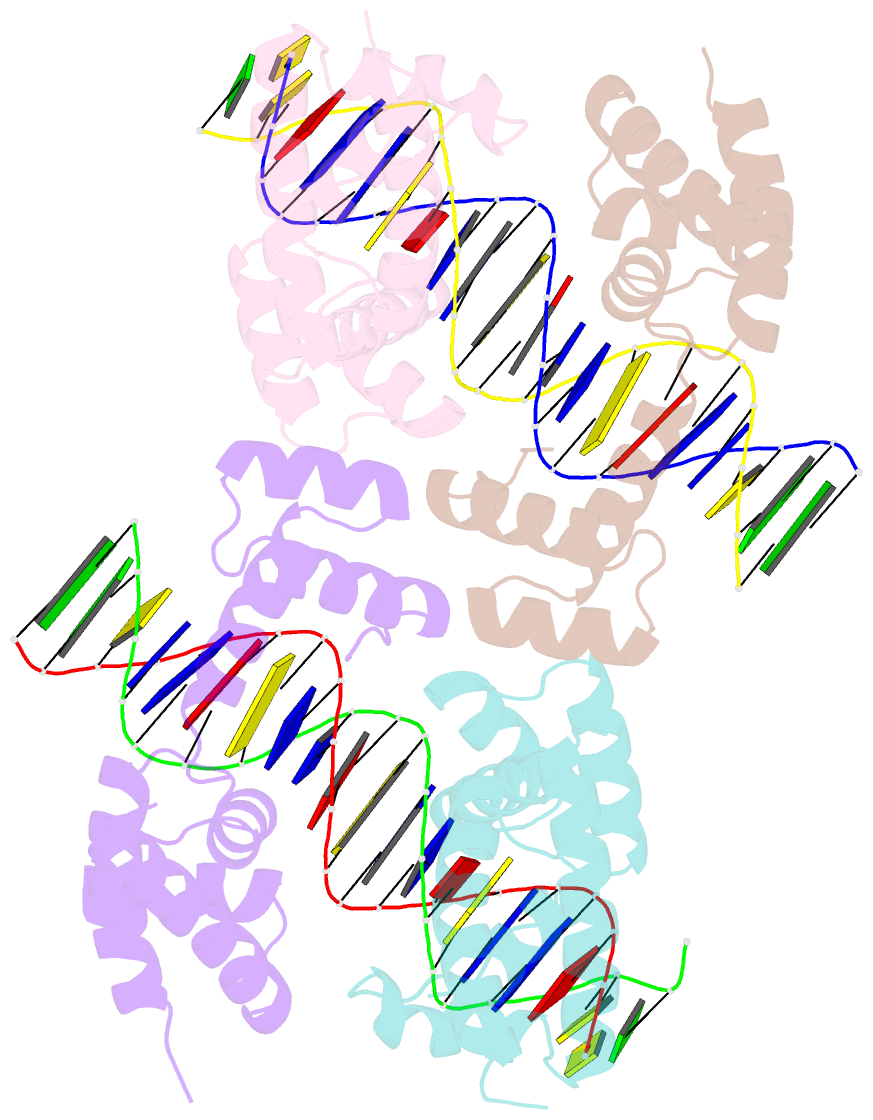
- Human pit-1 and 4xcatt DNA complex
- Reference
- Agarwal S, Cho TY (2018): "Biochemical and structural characterization of a novel cooperative binding mode by Pit-1 with CATT repeats in the macrophage migration inhibitory factor promoter." Nucleic Acids Res., 46, 929-941. doi: 10.1093/nar/gkx1183.
- Abstract
- Overexpression of the proinflammatory cytokine macrophage migration inhibitory factor (MIF) is linked to a number of autoimmune diseases and cancer. MIF production has been correlated to the number of CATT repeats in a microsatellite region upstream of the MIF gene. We have characterized the interaction of pituitary-specific positive transcription factor 1 (Pit-1) with a portion of the MIF promoter region flanking a microsatellite polymorphism (-794 CATT5-8). Using fluorescence anisotropy, we quantified tight complex formation between Pit-1 and an oligonucleotide consisting of eight consecutive CATT repeats (8xCATT) with an apparent Kd of 35 nM. Using competition experiments we found a 23 base pair oligonucleotide with 4xCATT repeats to be the minimum DNA sequence necessary for high affinity interaction with Pit-1. The stoichiometry of the Pit-1 DNA interaction was determined to be 2:1 and binding is cooperative in nature. We subsequently structurally characterized the complex and discovered a completely novel binding mode for Pit-1 in contrast to previously described Pit-1 complex structures. The affinity of Pit-1 for the CATT target sequence was found to be highly dependent on cooperativity. This work lays the groundwork for understanding transcriptional regulation of MIF and pursuing Pit-1 as a therapeutic target to treat MIF-mediated inflammatory disorders.