Summary information and primary citation
- PDB-id
- 5z9w; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- viral protein
- Method
- cryo-EM (3.6 Å)
- Summary
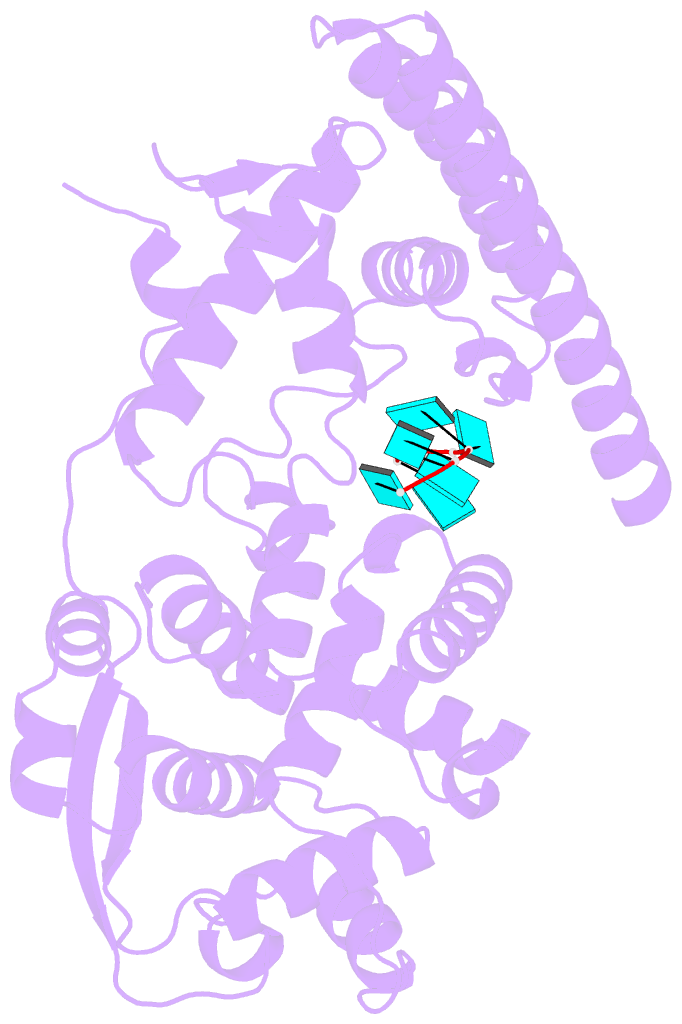
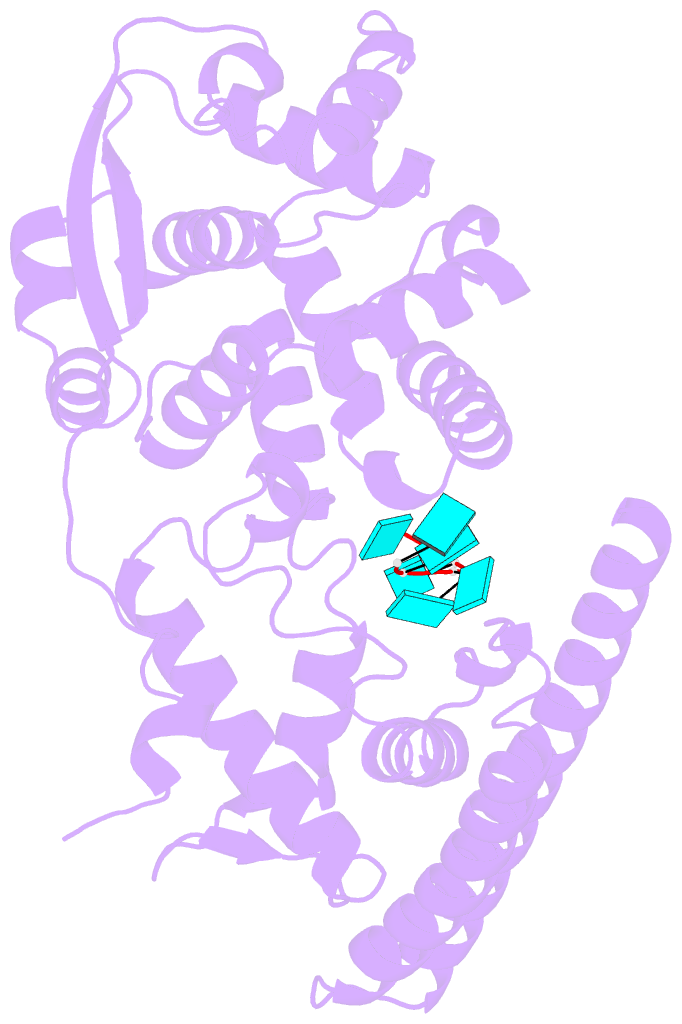
- Ebola virus nucleoprotein-RNA complex
- Reference
- Sugita Y, Matsunami H, Kawaoka Y, Noda T, Wolf M (2018): "Cryo-EM structure of the Ebola virus nucleoprotein-RNA complex at 3.6 angstrom resolution." Nature, 563, 137-140. doi: 10.1038/s41586-018-0630-0.
- Abstract
- Ebola virus causes haemorrhagic fever with a high fatality rate in humans and non-human primates. It belongs to the family Filoviridae in the order Mononegavirales, which are viruses that contain linear, non-segmented, negative-sense, single-stranded genomic RNA1,2. The enveloped, filamentous virion contains the nucleocapsid, consisting of the helical nucleoprotein-RNA complex, VP24, VP30, VP35 and viral polymerase1,3. The nucleoprotein-RNA complex acts as a scaffold for nucleocapsid formation and as a template for RNA replication and transcription by condensing RNA into the virion4,5. RNA binding and nucleoprotein oligomerization are synergistic and do not readily occur independently6. Although recent cryo-electron tomography studies have revealed the overall architecture of the nucleocapsid core4,5, there has been no high-resolution reconstruction of the nucleocapsid. Here we report the structure of a recombinant Ebola virus nucleoprotein-RNA complex expressed in mammalian cells without chemical fixation, at near-atomic resolution using single-particle cryo-electron microscopy. Our structure reveals how the Ebola virus nucleocapsid core encapsidates its viral genome, its sequence-independent coordination with RNA by nucleoprotein, and the dynamic transition between the RNA-free and RNA-bound states. It provides direct structural evidence for the role of the N terminus of nucleoprotein in subunit oligomerization, and for the hydrophobic and electrostatic interactions that lead to the formation of the helical assembly. The structure is validated as representative of the native biological assembly of the nucleocapsid core by consistent dimensions and symmetry with the full virion5. The atomic model provides a detailed mechanistic basis for understanding nucleocapsid assembly and highlights key structural features that may serve as targets for anti-viral drug development.