Summary information and primary citation
- PDB-id
- 5zw4; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein-RNA
- Method
- X-ray (1.7 Å)
- Summary
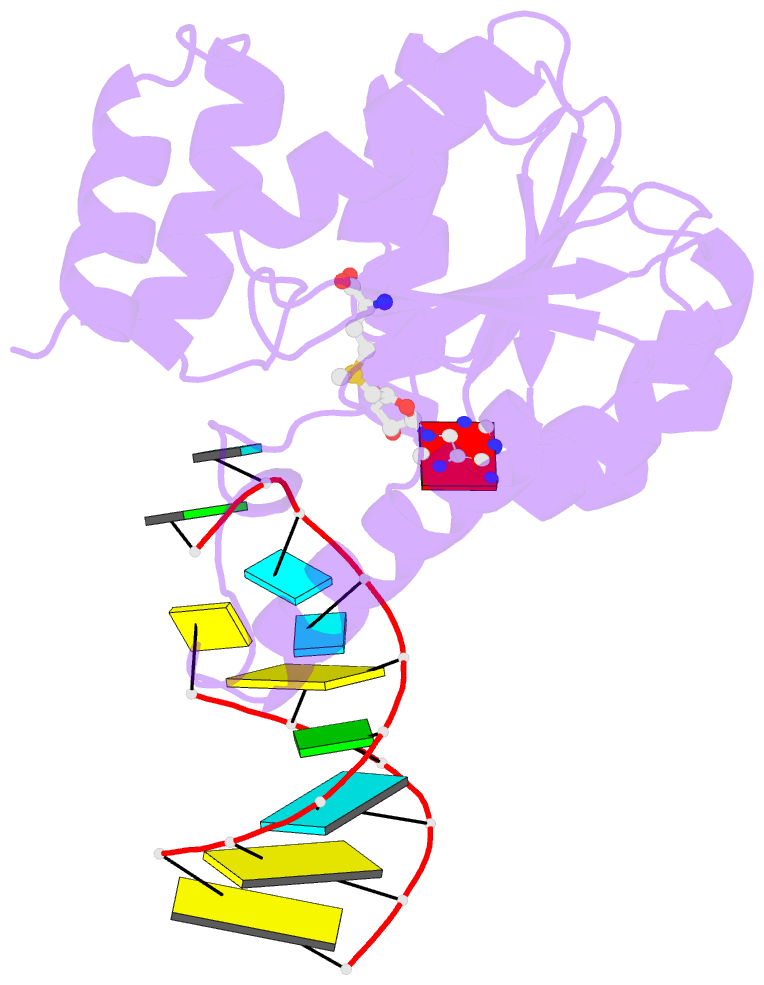
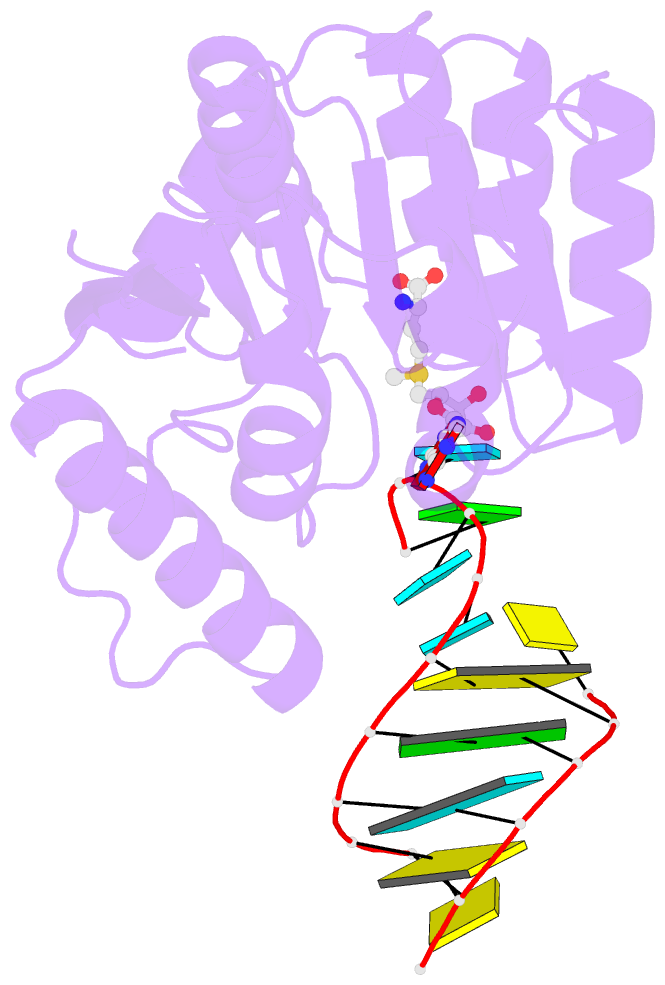
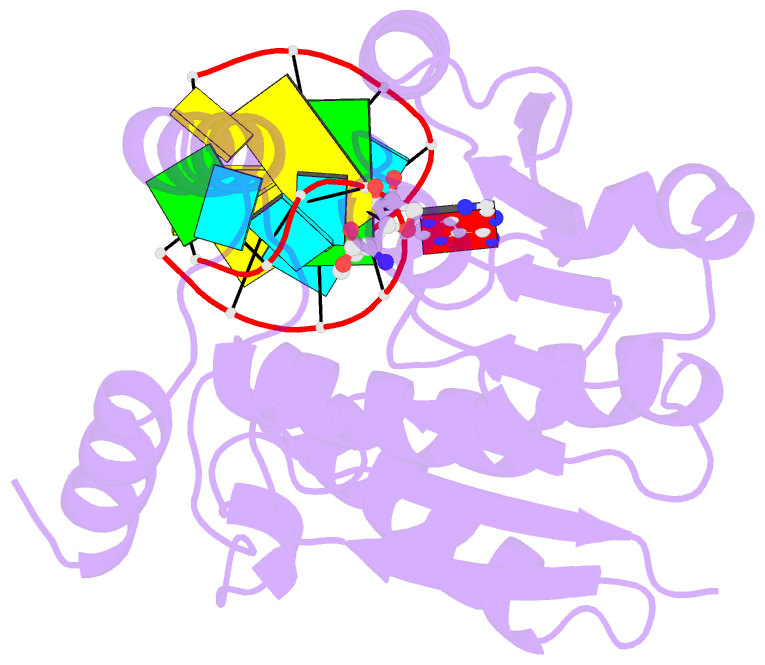
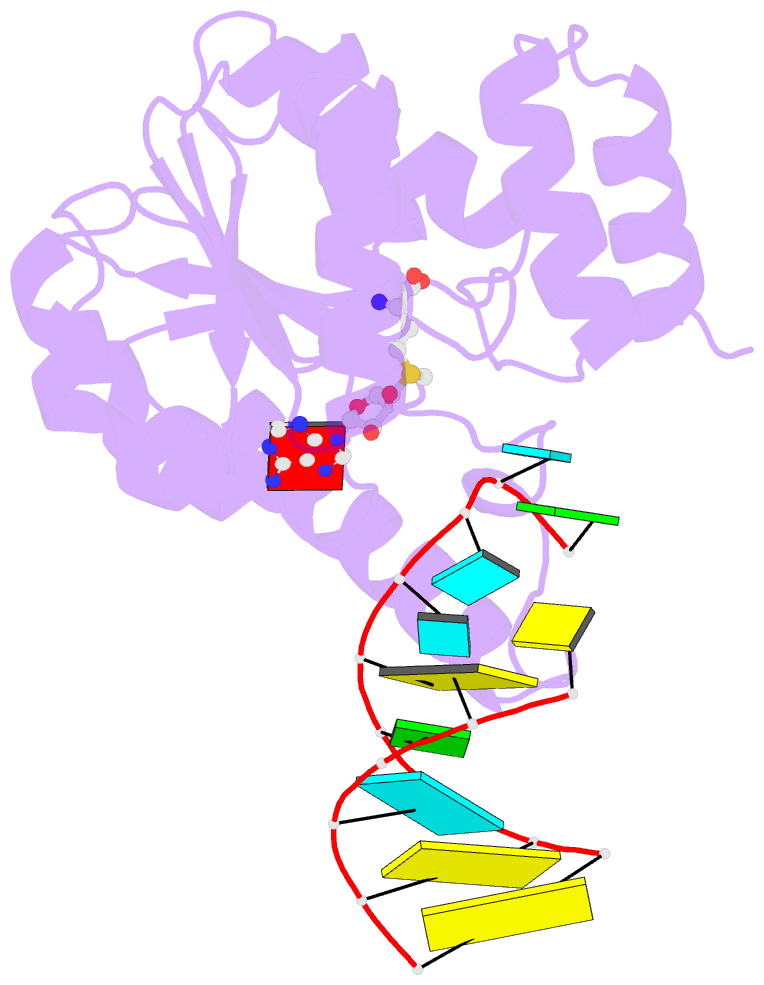
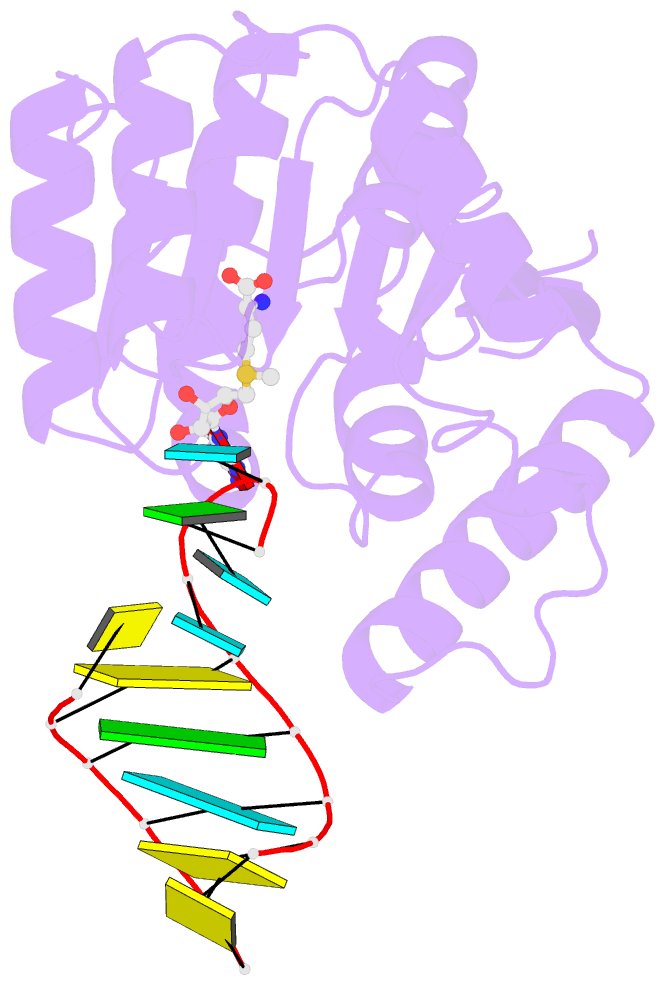
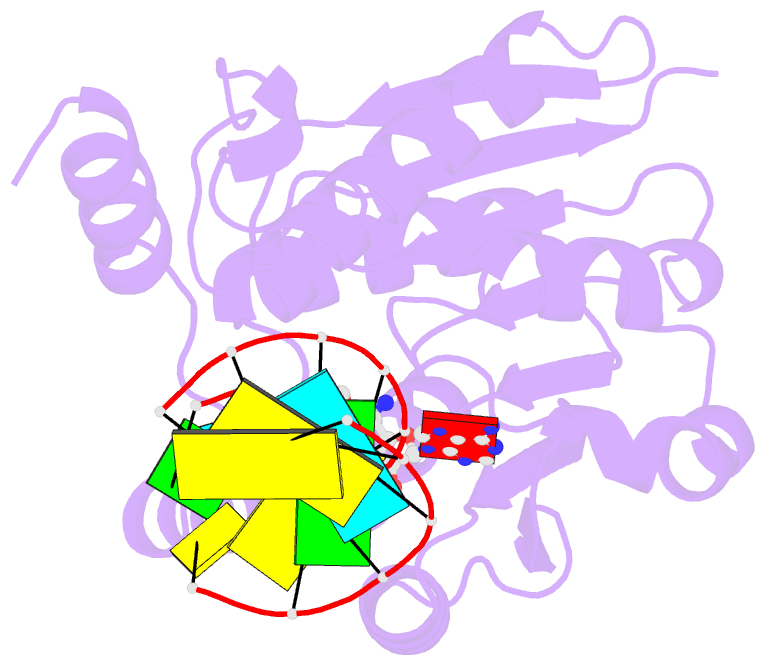
- Crystal structure of trna bound trmr
- Reference
- Ryu H, Grove TL, Almo SC, Kim J (2018): "Identification of a novel tRNA wobble uridine modifying activity in the biosynthesis of 5-methoxyuridine." Nucleic Acids Res., 46, 9160-9169. doi: 10.1093/nar/gky592.
- Abstract
- Derivatives of 5-hydroxyuridine (ho5U), such as 5-methoxyuridine (mo5U) and 5-oxyacetyluridine (cmo5U), are ubiquitous modifications of the wobble position of bacterial tRNA that are believed to enhance translational fidelity by the ribosome. In gram-negative bacteria, the last step in the biosynthesis of cmo5U from ho5U involves the unique metabolite carboxy S-adenosylmethionine (Cx-SAM) and the carboxymethyl transferase CmoB. However, the equivalent position in the tRNA of Gram-positive bacteria is instead mo5U, where the methyl group is derived from SAM and installed by an unknown methyltransferase. By utilizing a cmoB-deficient strain of Escherichia coli as a host and assaying for the formation of mo5U in total RNA isolates with methyltransferases of unknown function from Bacillus subtilis, we found that this modification is installed by the enzyme TrmR (formerly known as YrrM). Furthermore, X-ray crystal structures of TrmR with and without the anticodon stemloop of tRNAAla have been determined, which provide insight into both sequence and structure specificity in the interactions of TrmR with tRNA.