Summary information and primary citation
- PDB-id
- 6bek; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein-DNA
- Method
- X-ray (1.7 Å)
- Summary
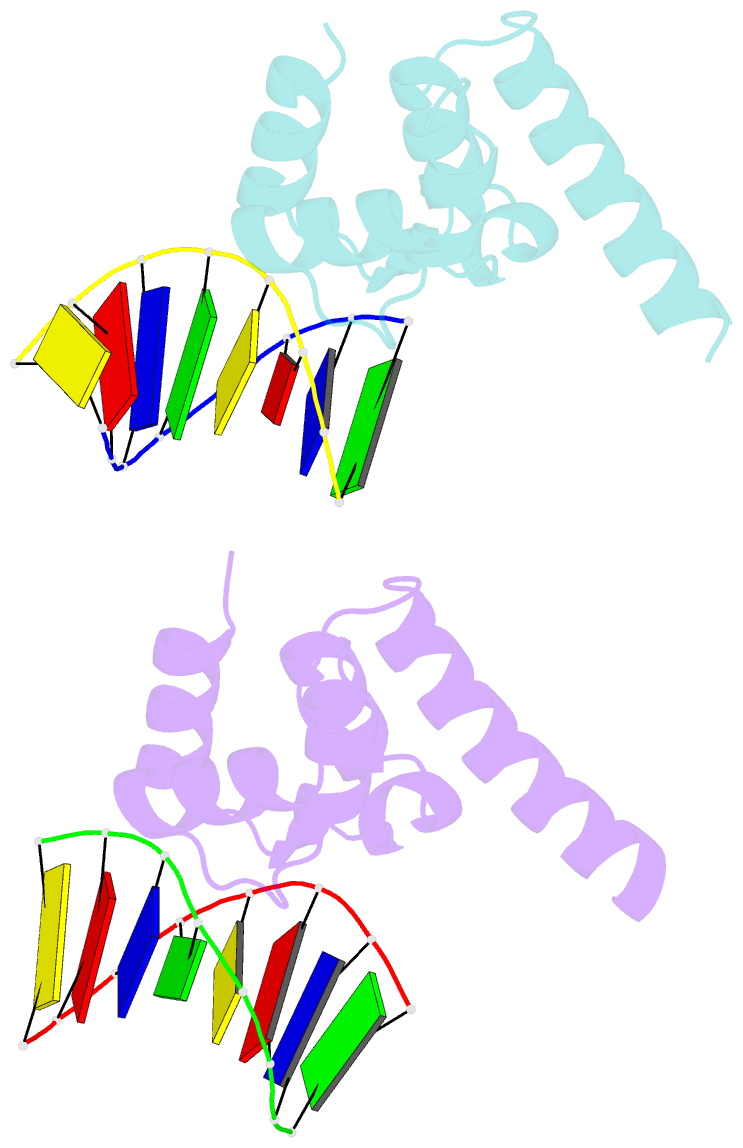
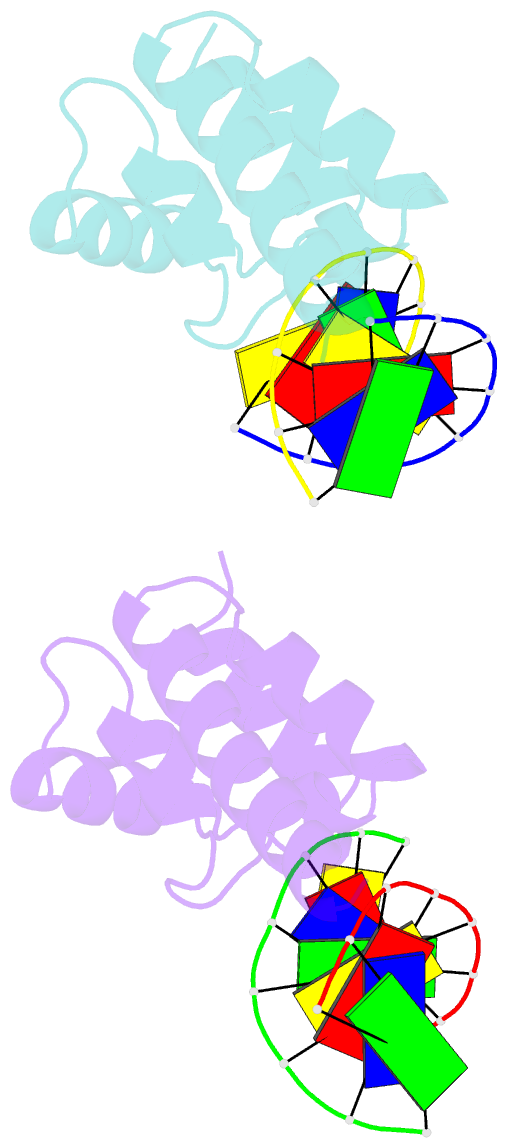
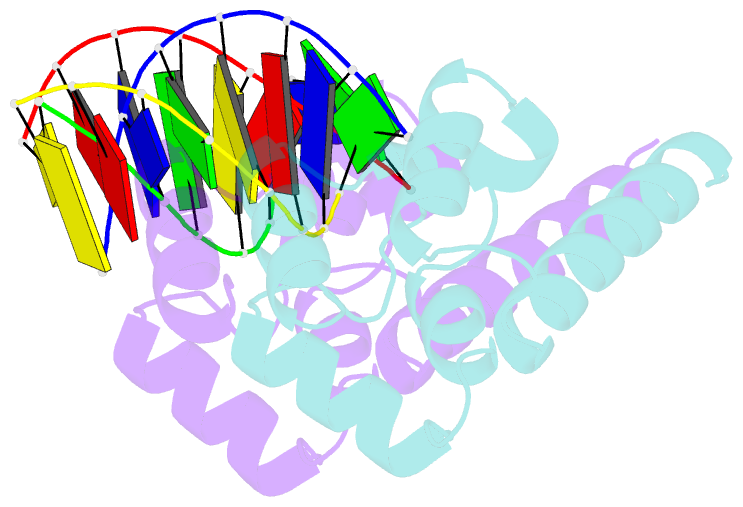
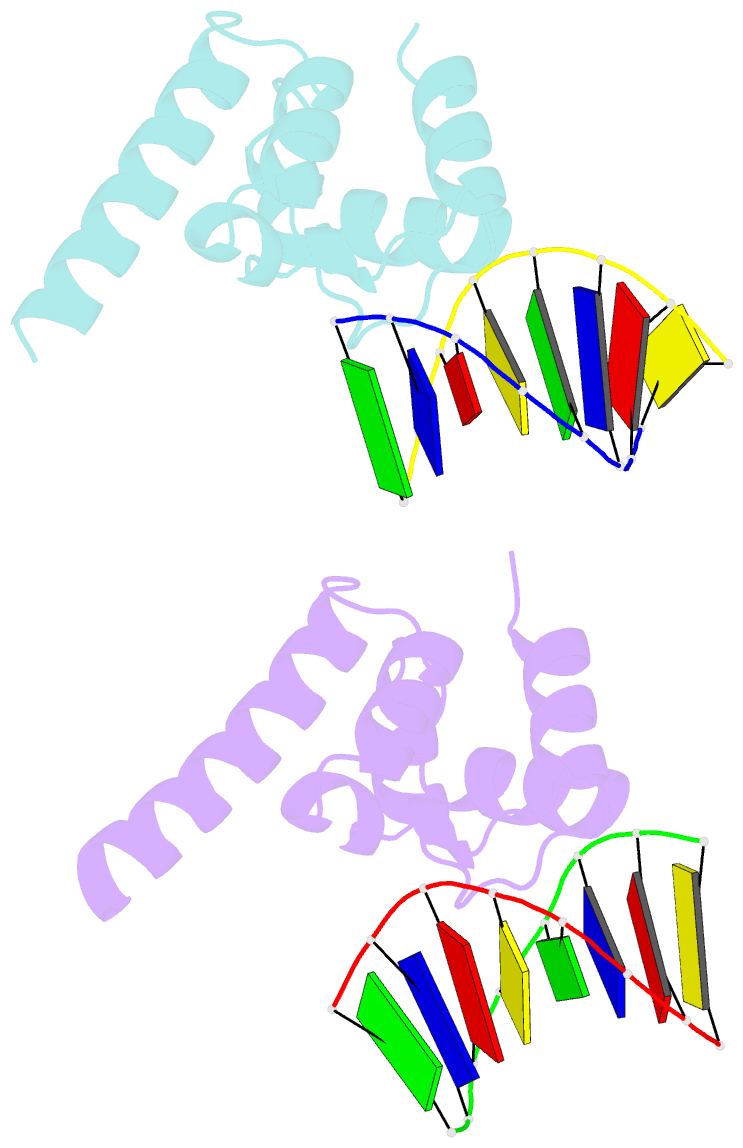
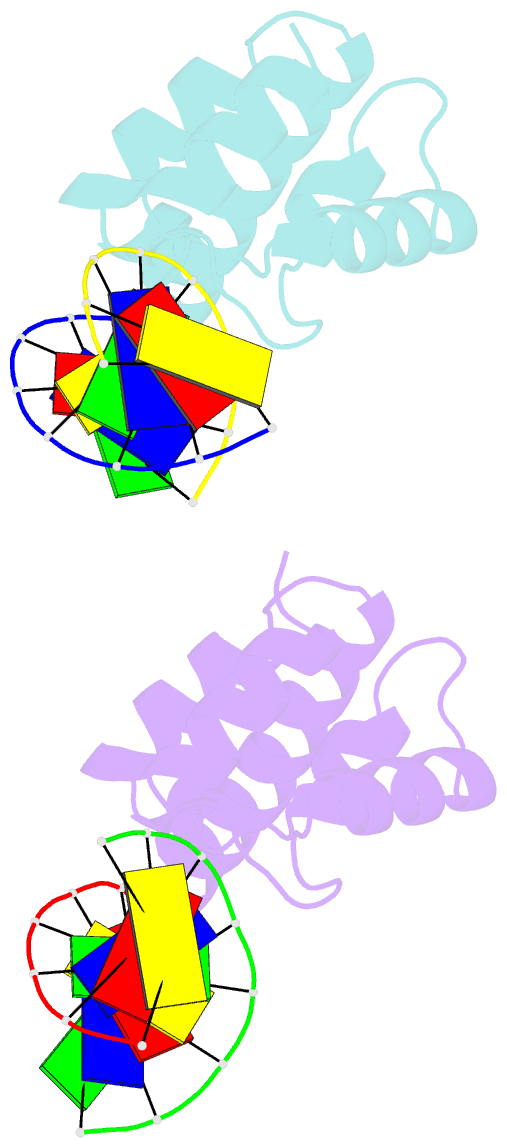
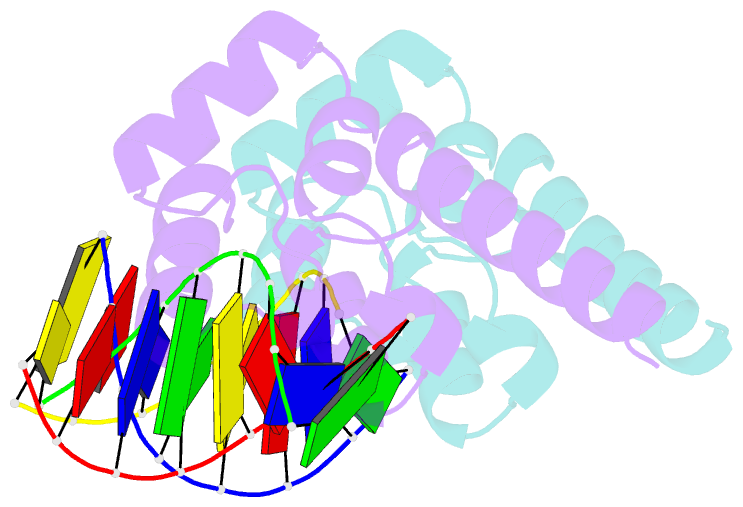
- Structure of sihf bound to an 8bp palindromic DNA
- Reference
- Nanji T, Gehrke EJ, Shen Y, Gloyd M, Zhang X, Firby CD, Huynh A, Razi A, Ortega J, Elliot MA, Guarne A (2019): "Streptomyces IHF uses multiple interfaces to bind DNA." Biochim Biophys Acta Gen Subj, 1863, 129405. doi: 10.1016/j.bbagen.2019.07.014.
- Abstract
- Background: Nucleoid associated proteins (NAPs) are essential for chromosome condensation in bacterial cells. Despite being a diverse group, NAPs share two common traits: they are small, oligomeric proteins and their oligomeric state is critical for DNA condensation. Streptomyces coelicolor IHF (sIHF) is an actinobacterial-specific nucleoid-associated protein that despite its name, shares neither sequence nor structural homology with the well-characterized Escherichia coli IHF. Like E. coli IHF, sIHF is needed for efficient nucleoid condensation, morphological development and antibiotic production in S. coelicolor.
Methods: Using a combination of crystallography, small-angle X-ray scattering, electron microscopy and structure-guided functional assays, we characterized how sIHF binds and remodels DNA.
Results: The structure of sIHF bound to DNA revealed two DNA-binding elements on opposite surfaces of the helix bundle. Using structure-guided functional assays, we identified an additional surface that drives DNA binding in solution. Binding by each element is necessary for both normal development and antibiotic production in vivo, while in vitro, they act collectively to restrain negative supercoils.
Conclusions: The cleft defined by the N-terminal and the helix bundle of sIHF drives DNA binding, but the two additional surfaces identified on the crystal structure are necessary to stabilize binding, remodel DNA and maintain wild-type levels of antibiotic production. We propose a model describing how the multiple DNA-binding elements enable oligomerization-independent nucleoid condensation.