Summary information and primary citation
- PDB-id
- 6h5s; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- viral protein
- Method
- cryo-EM (3.3 Å)
- Summary
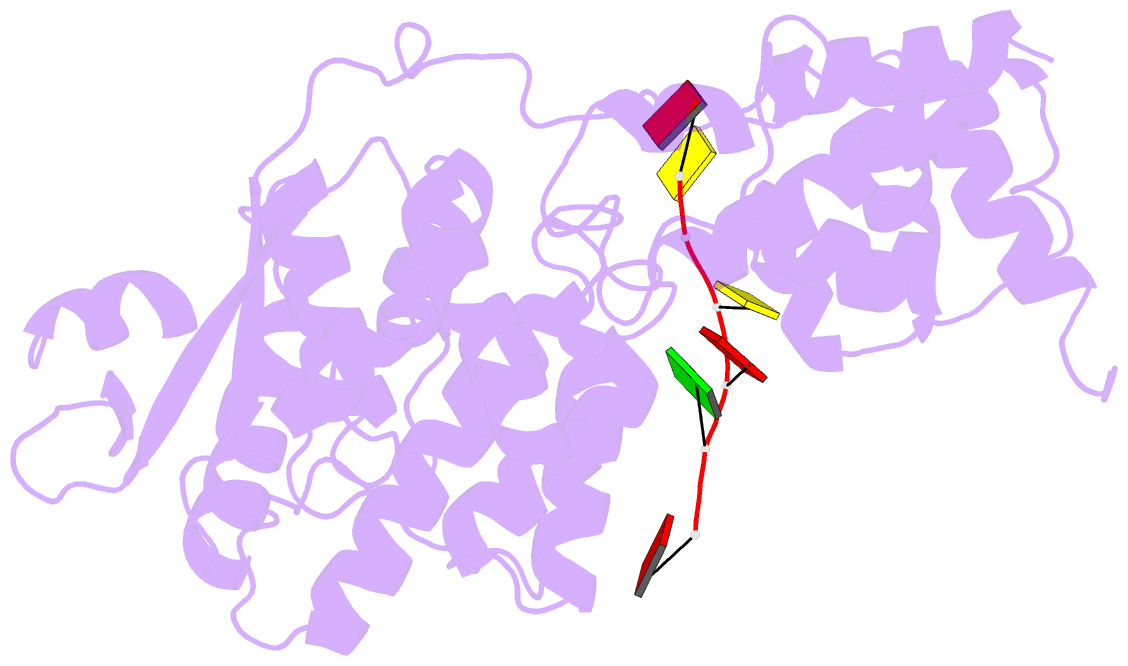
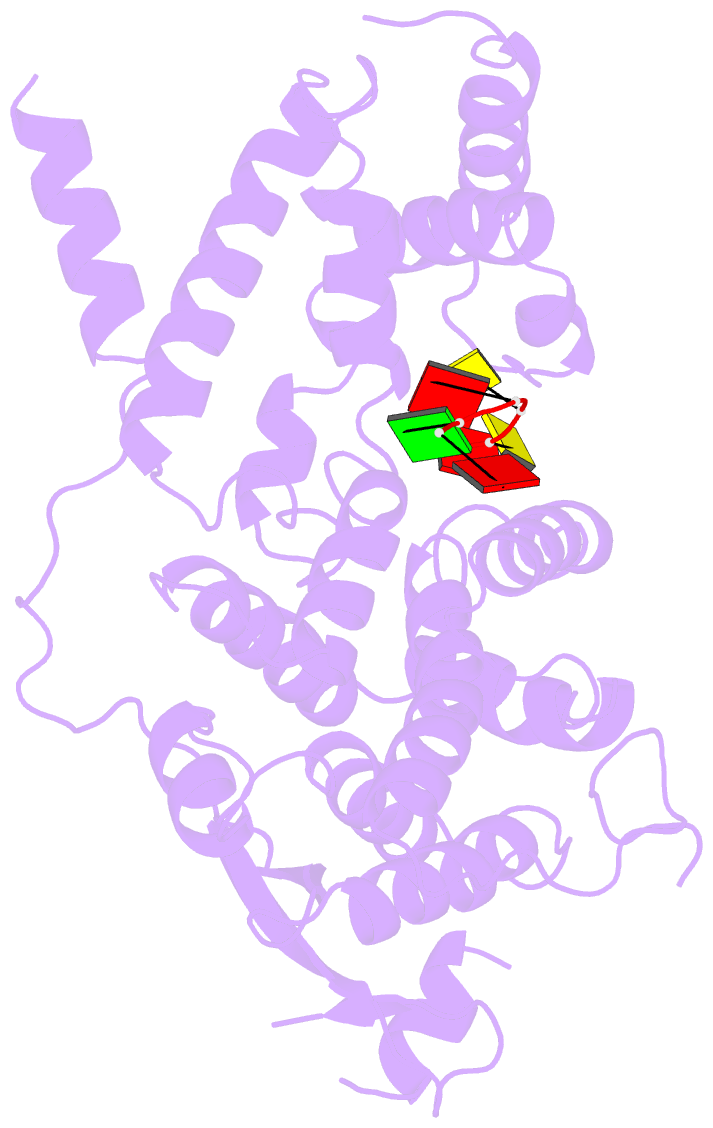
- cryo-EM map of in vitro assembled measles virus n into nucleocapsid-like particles (nclps) bound to viral genomic 5-prime RNA hexamers.
- Reference
- Desfosses A, Milles S, Jensen MR, Guseva S, Colletier JP, Maurin D, Schoehn G, Gutsche I, Ruigrok RWH, Blackledge M (2019): "Assembly and cryo-EM structures of RNA-specific measles virus nucleocapsids provide mechanistic insight into paramyxoviral replication." Proc.Natl.Acad.Sci.USA, 116, 4256-4264. doi: 10.1073/pnas.1816417116.
- Abstract
- Assembly of paramyxoviral nucleocapsids on the RNA genome is an essential step in the viral cycle. The structural basis of this process has remained obscure due to the inability to control encapsidation. We used a recently developed approach to assemble measles virus nucleocapsid-like particles on specific sequences of RNA hexamers (poly-Adenine and viral genomic 5') in vitro, and determined their cryoelectron microscopy maps to 3.3-Å resolution. The structures unambiguously determine 5' and 3' binding sites and thereby the binding-register of viral genomic RNA within nucleocapsids. This observation reveals that the 3' end of the genome is largely exposed in fully assembled measles nucleocapsids. In particular, the final three nucleotides of the genome are rendered accessible to the RNA-dependent RNA polymerase complex, possibly enabling efficient RNA processing. The structures also reveal local and global conformational changes in the nucleoprotein upon assembly, in particular involving helix α6 and helix α13 that form edges of the RNA binding groove. Disorder is observed in the bound RNA, localized at one of the two backbone conformational switch sites. The high-resolution structure allowed us to identify putative nucleobase interaction sites in the RNA-binding groove, whose impact on assembly kinetics was measured using real-time NMR. Mutation of one of these sites, R195, whose sidechain stabilizes both backbone and base of a bound nucleic acid, is thereby shown to be essential for nucleocapsid-like particle assembly.