Summary information and primary citation
- PDB-id
- 6ifo; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein-RNA
- Method
- X-ray (3.313 Å)
- Summary
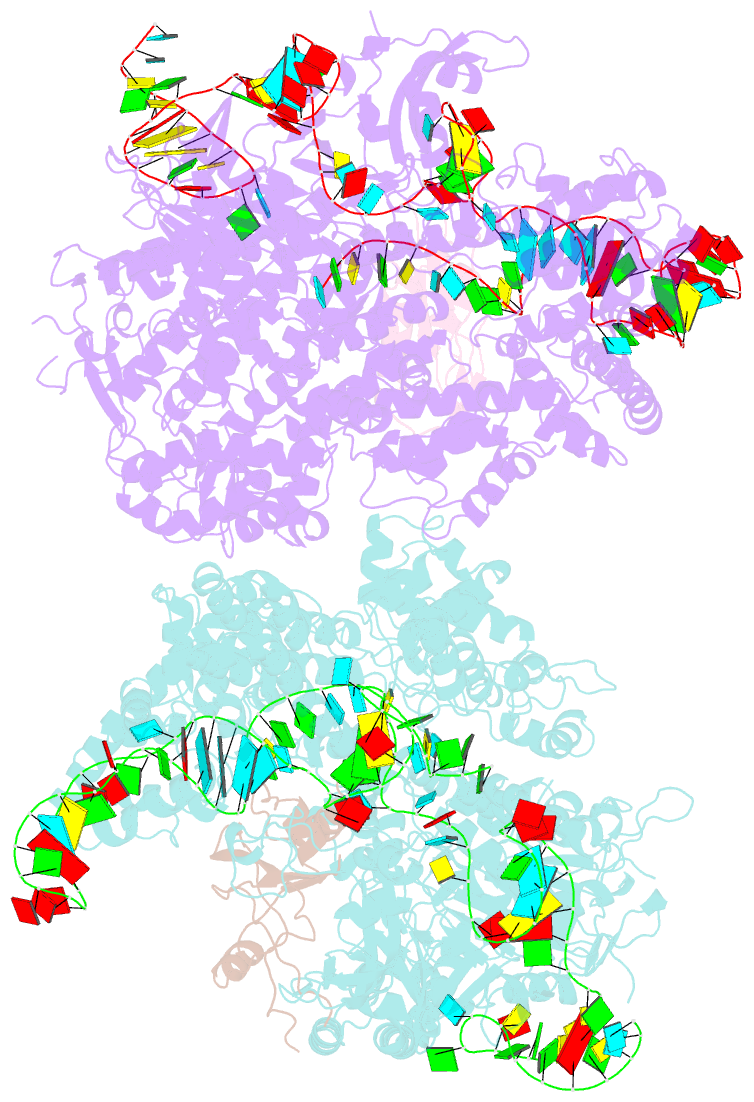
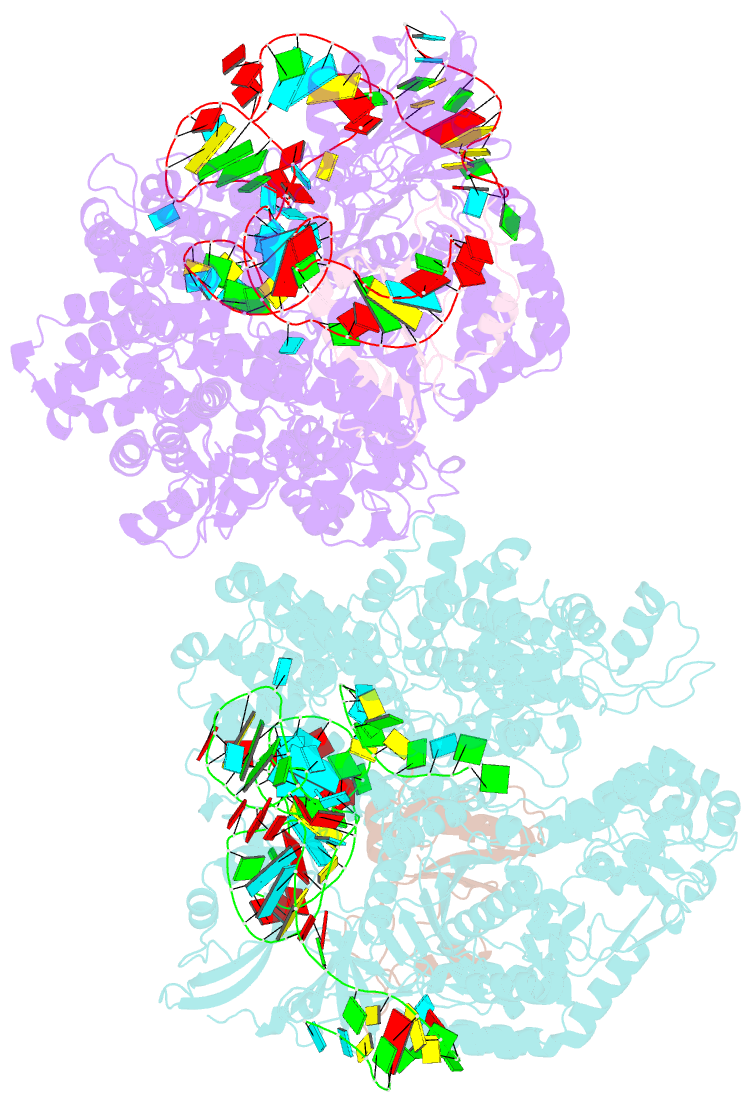
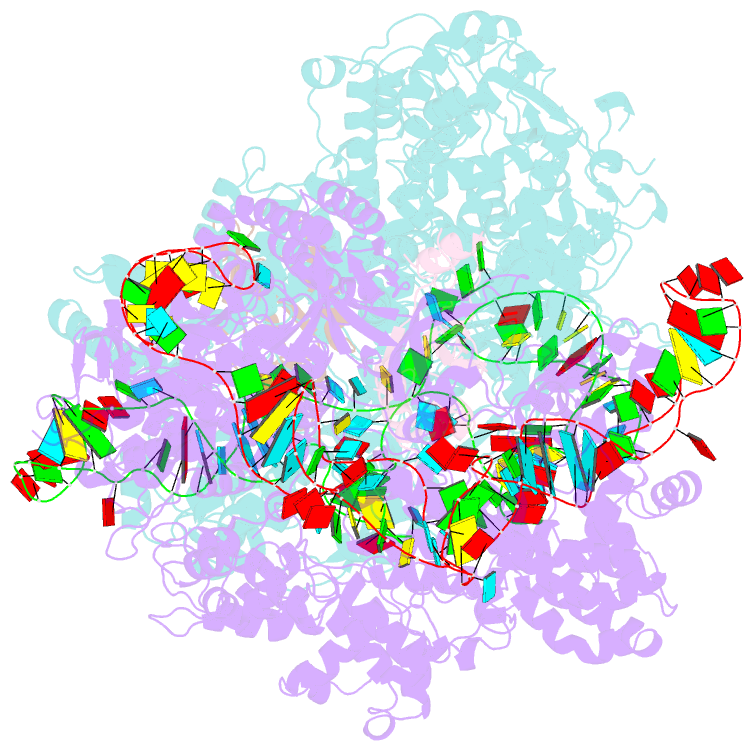
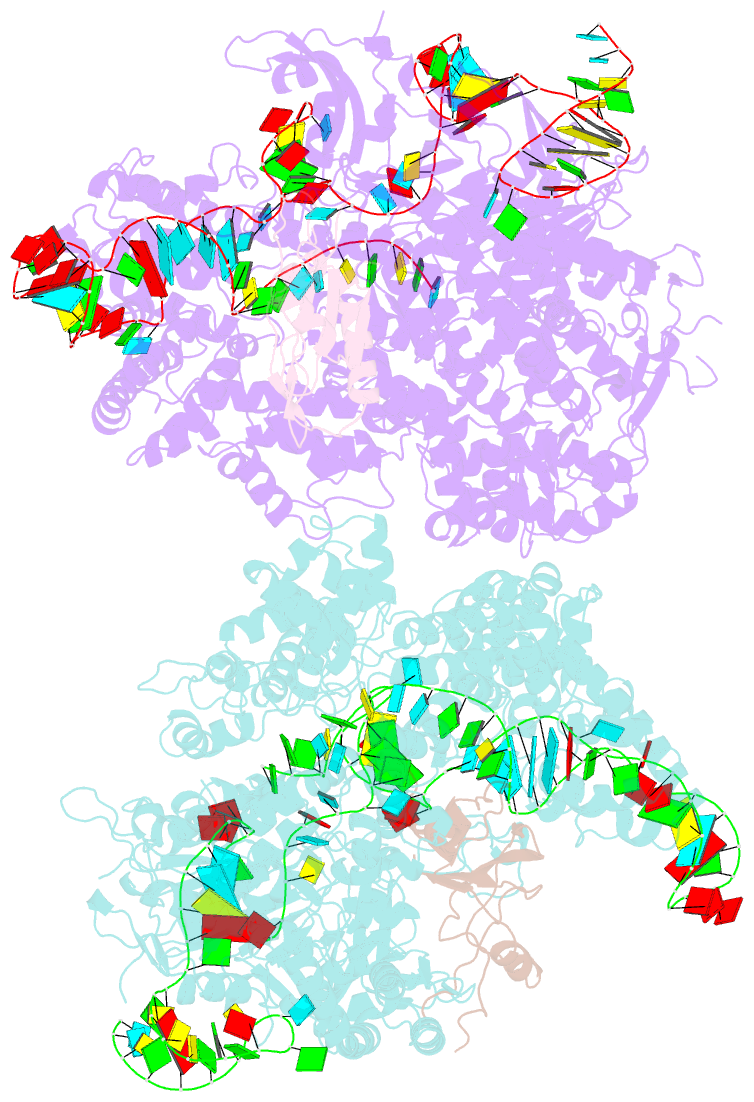
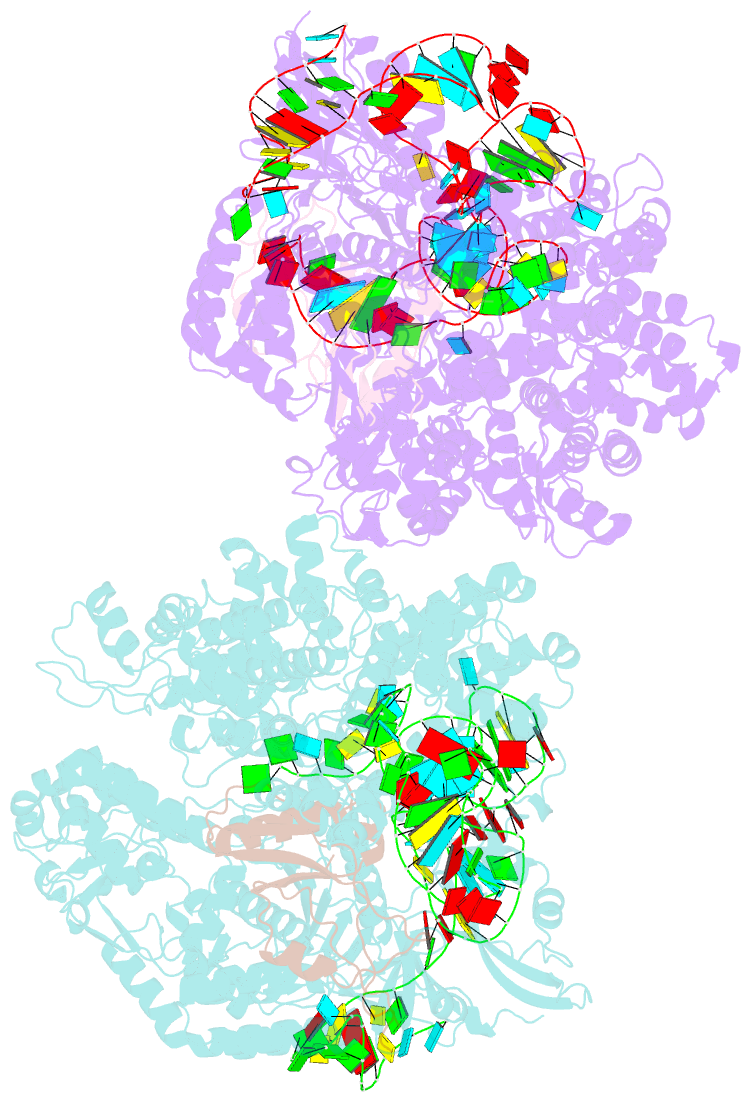
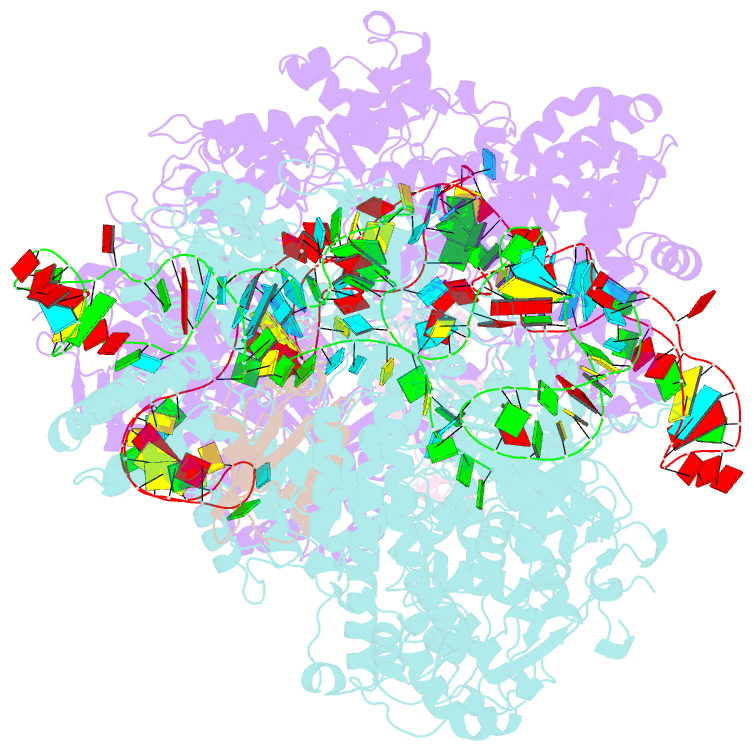
- Crystal structure of acriia2-spycas9-sgrna ternary complex
- Reference
- Liu L, Yin M, Wang M, Wang Y (2019): "Phage AcrIIA2 DNA Mimicry: Structural Basis of the CRISPR and Anti-CRISPR Arms Race." Mol. Cell, 73, 611-620.e3. doi: 10.1016/j.molcel.2018.11.011.
- Abstract
- CRISPR-Cas (clustered regularly interspaced short palindromic repeats-CRISPR-associated proteins) systems provide prokaryotic cells with adaptive immunity against invading bacteriophages. Bacteriophages counteract bacterial responses by encoding anti-CRISPR inhibitor proteins (Acr). However, the structural basis for their inhibitory actions remains largely unknown. Here, we report the crystal structure of the AcrIIA2-SpyCas9-sgRNA (single-guide RNA) complex at 3.3 Å resolution. We show that AcrIIA2 binds SpyCas9 at a position similar to the target DNA binding region. More specifically, AcrIIA2 interacts with the protospacer adjacent motif (PAM) recognition residues of Cas9, preventing target double-stranded DNA (dsDNA) detection. Thus, phage-encoded AcrIIA2 appears to act as a DNA mimic that blocks subsequent dsDNA binding by virtue of its highly acidic residues, disabling bacterial Cas9 by competing with target dsDNA binding with a binding motif distinct from AcrIIA4. Our study provides a more detailed mechanistic understanding of AcrIIA2-mediated inhibition of SpyCas9, the most widely used genome-editing tool, opening new avenues for improved regulatory precision during genome editing.