Summary information and primary citation
- PDB-id
- 6iro; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- DNA binding protein-DNA
- Method
- cryo-EM (3.4 Å)
- Summary
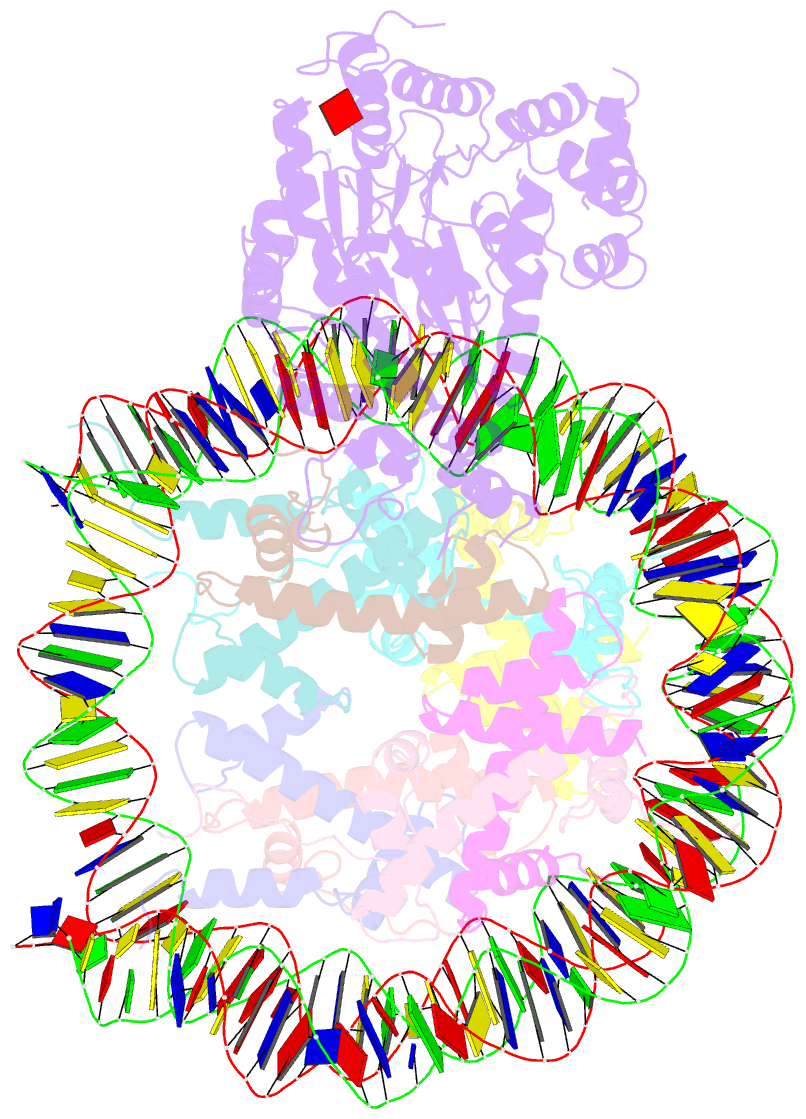
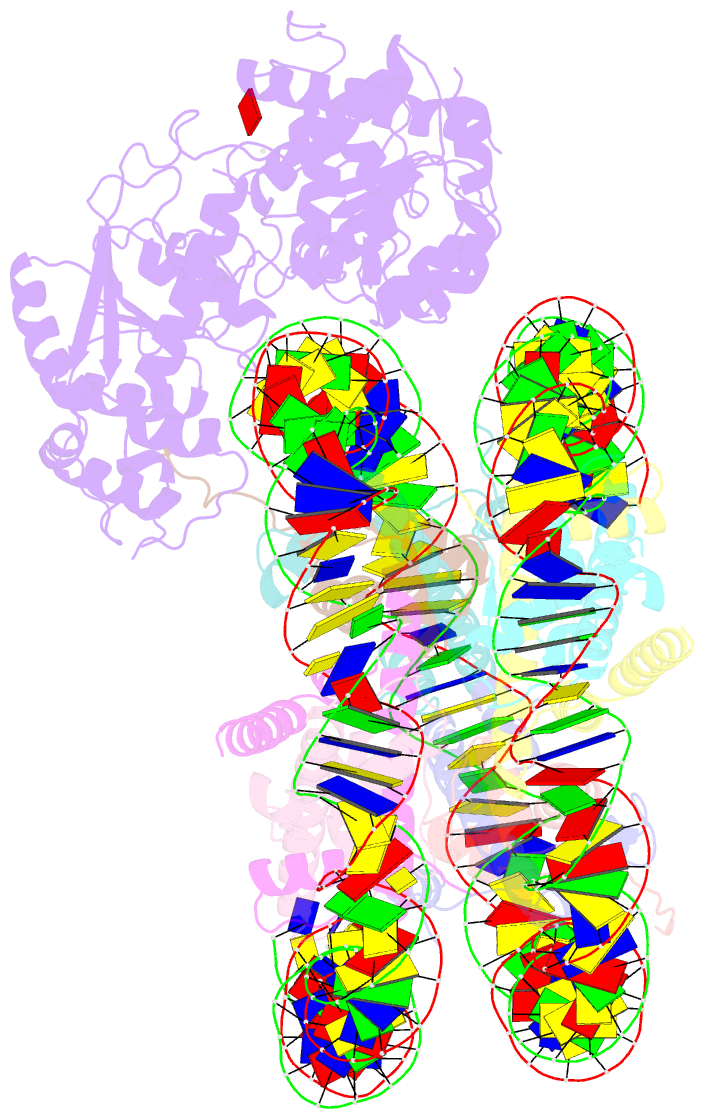
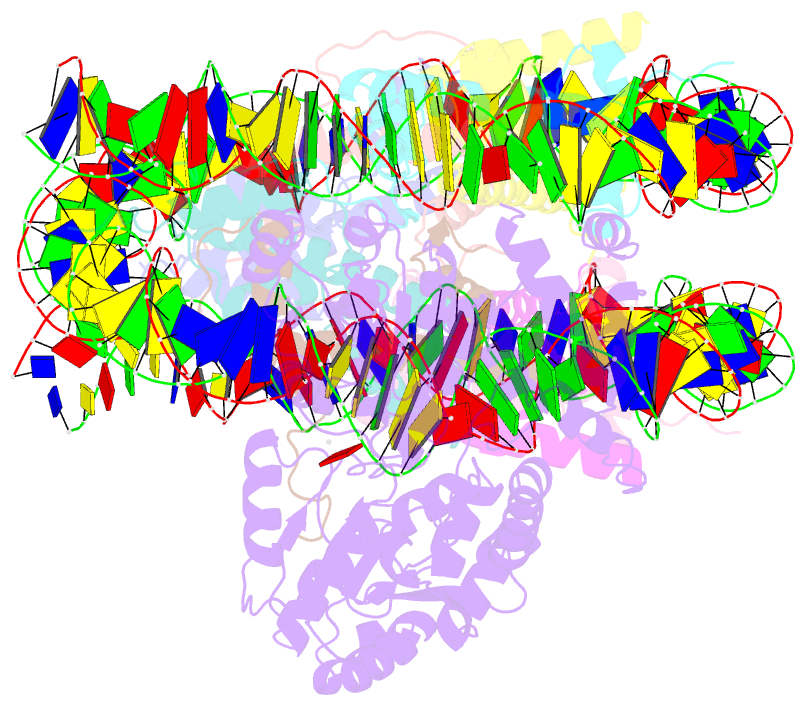
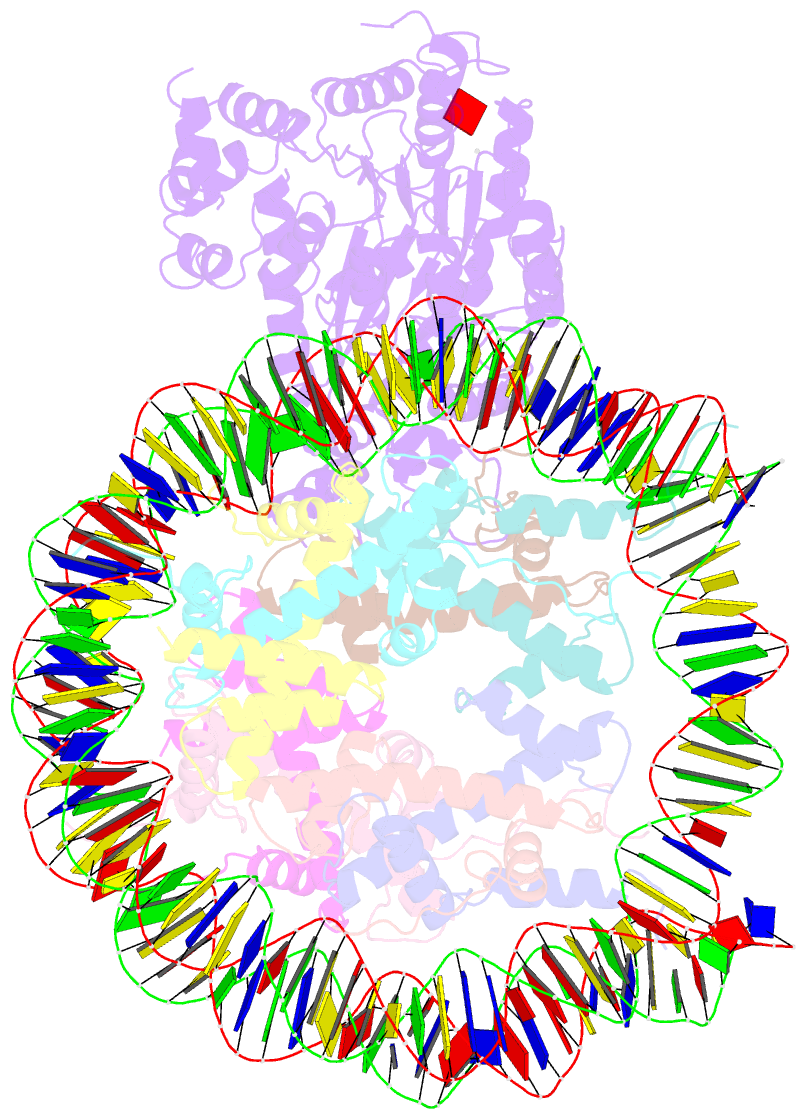
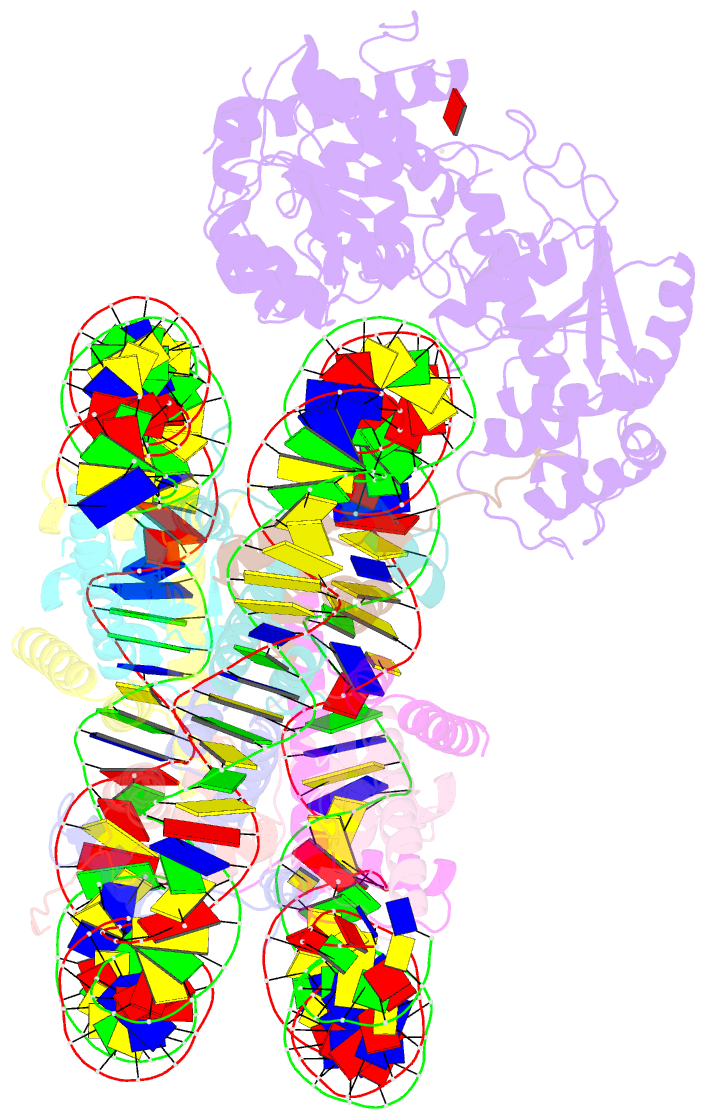
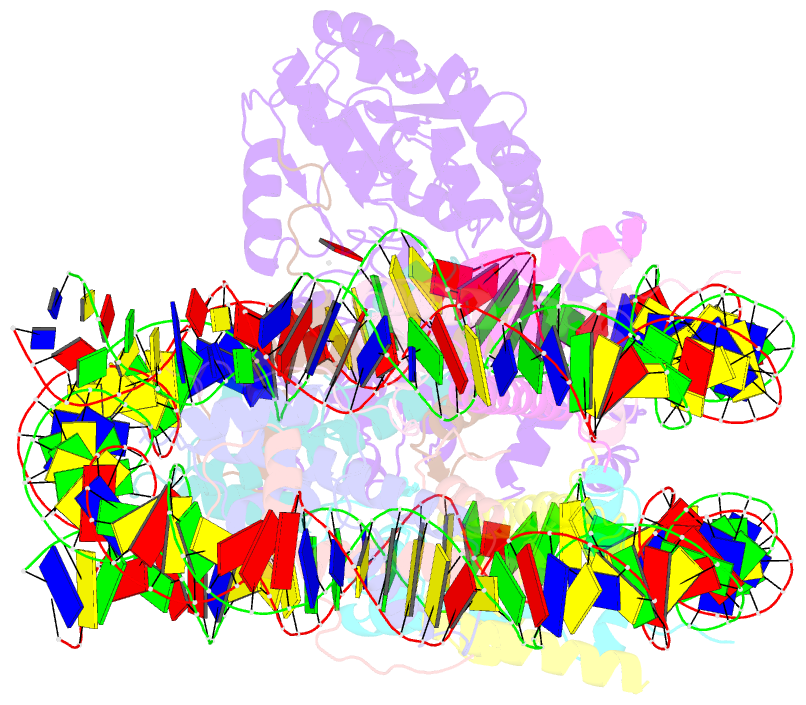
- The crosslinked complex of iswi-nucleosome in the adp-bound state
- Reference
- Yan L, Wu H, Li X, Gao N, Chen Z (2019): "Structures of the ISWI-nucleosome complex reveal a conserved mechanism of chromatin remodeling." Nat. Struct. Mol. Biol., 26, 258-266. doi: 10.1038/s41594-019-0199-9.
- Abstract
- Chromatin remodelers are diverse enzymes, and different models have been proposed to explain how these proteins work. Here we report the 3.3 Å-resolution cryogenic electron microscopy (cryo-EM) structures of Saccharomyces cerevisiae ISWI (ISW1) in complex with the nucleosome in adenosine diphosphate (ADP)-bound and ADP-BeFx-bound states. The data show that after nucleosome binding, ISW1 is activated by substantial rearrangement of the catalytic domains, with the regulatory AutoN domain packing the first RecA-like core and the NegC domain being disordered. The high-resolution structure reveals local DNA distortion and translocation induced by ISW1 in the ADP-bound state, which is essentially identical to that induced by the Snf2 chromatin remodeler, suggesting a common mechanism of DNA translocation. The histone core remains largely unperturbed, and prevention of histone distortion by crosslinking did not inhibit the activity of yeast ISW1 or its human homolog. Together, our findings suggest a general mechanism of chromatin remodeling involving local DNA distortion without notable histone deformation.