Summary information and primary citation
- PDB-id
- 6kdn; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transferase-DNA
- Method
- X-ray (2.303 Å)
- Summary
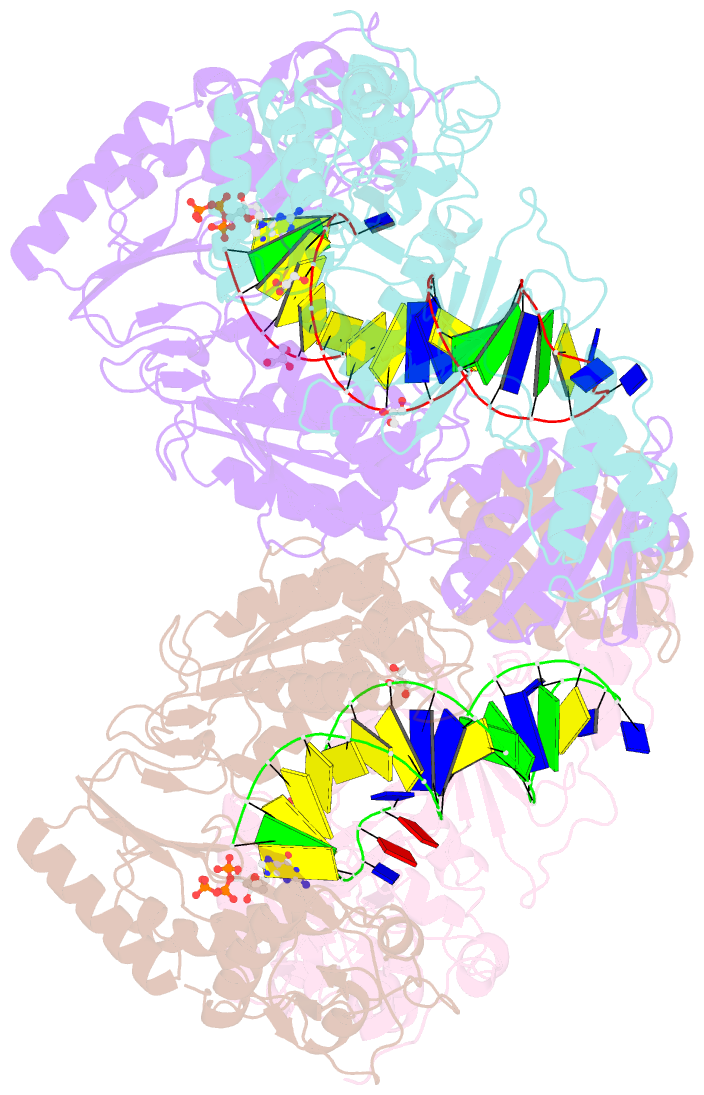
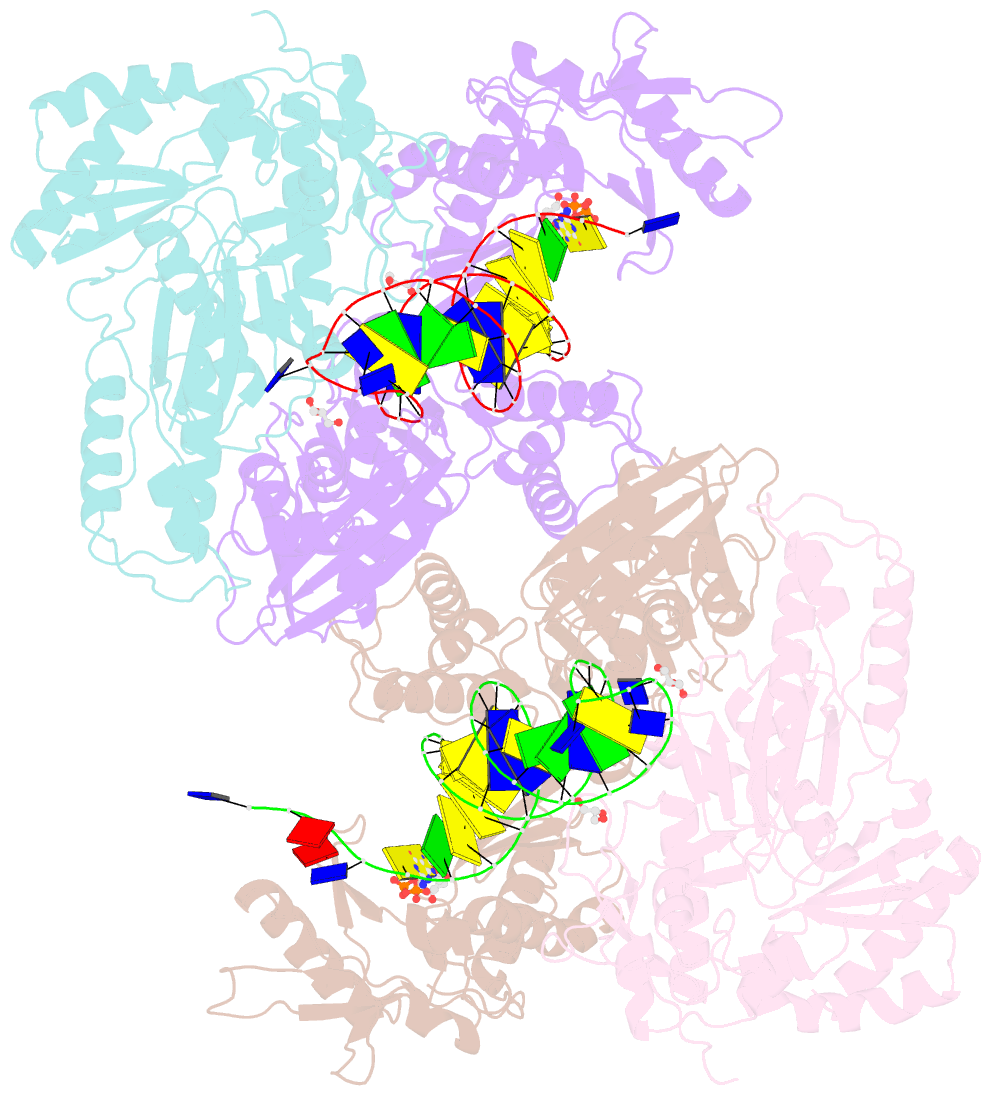
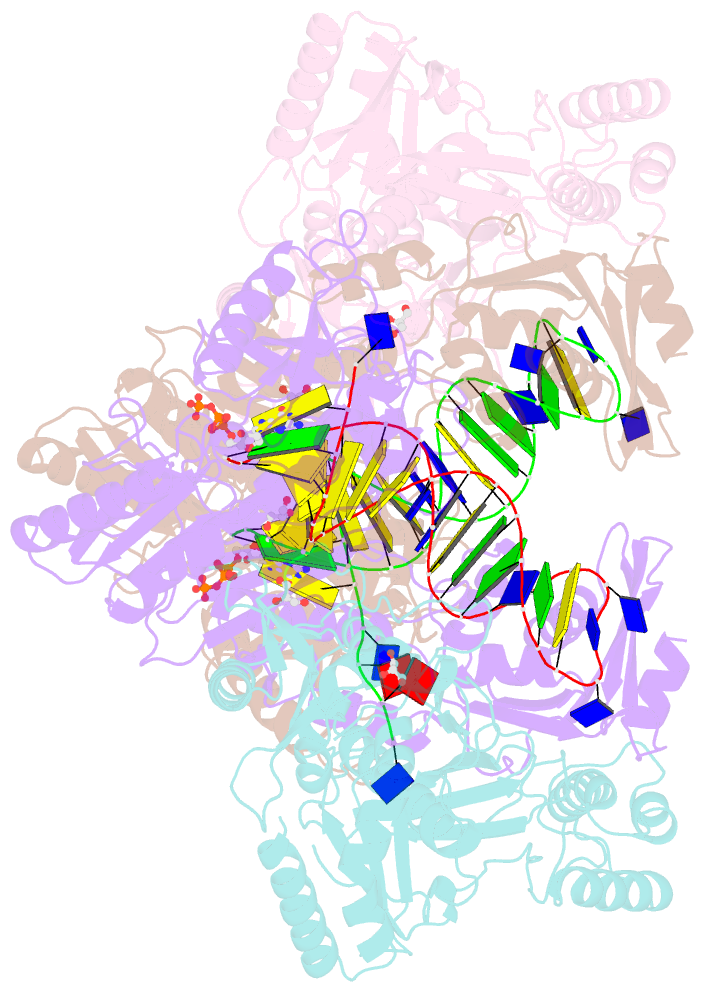
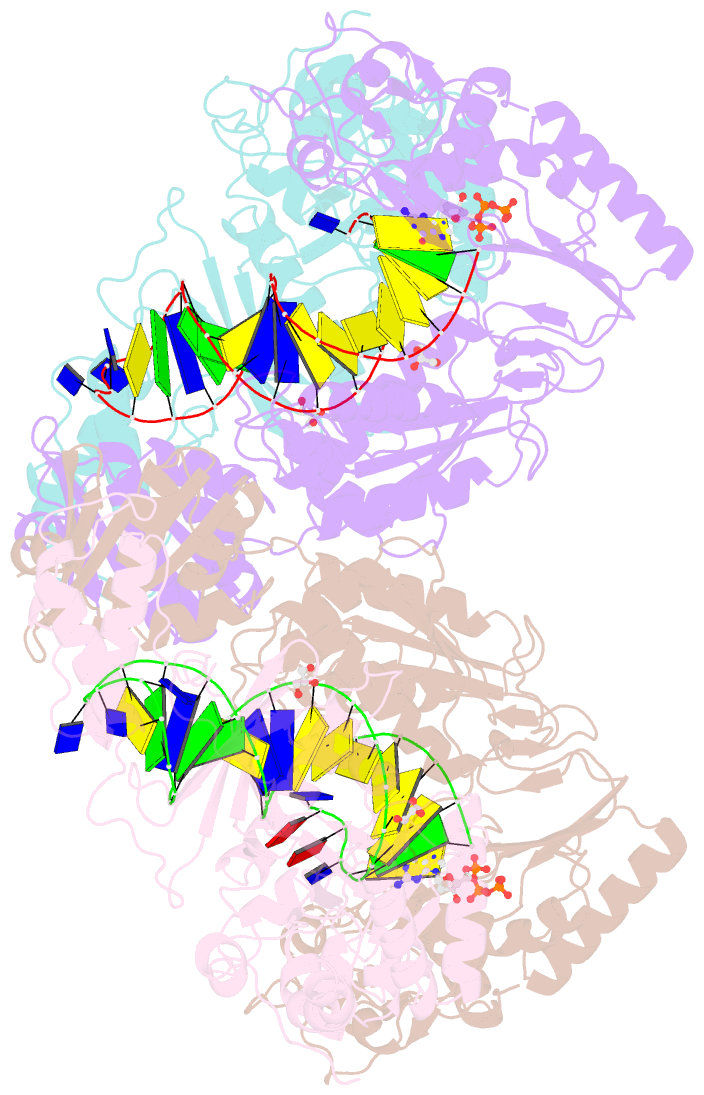
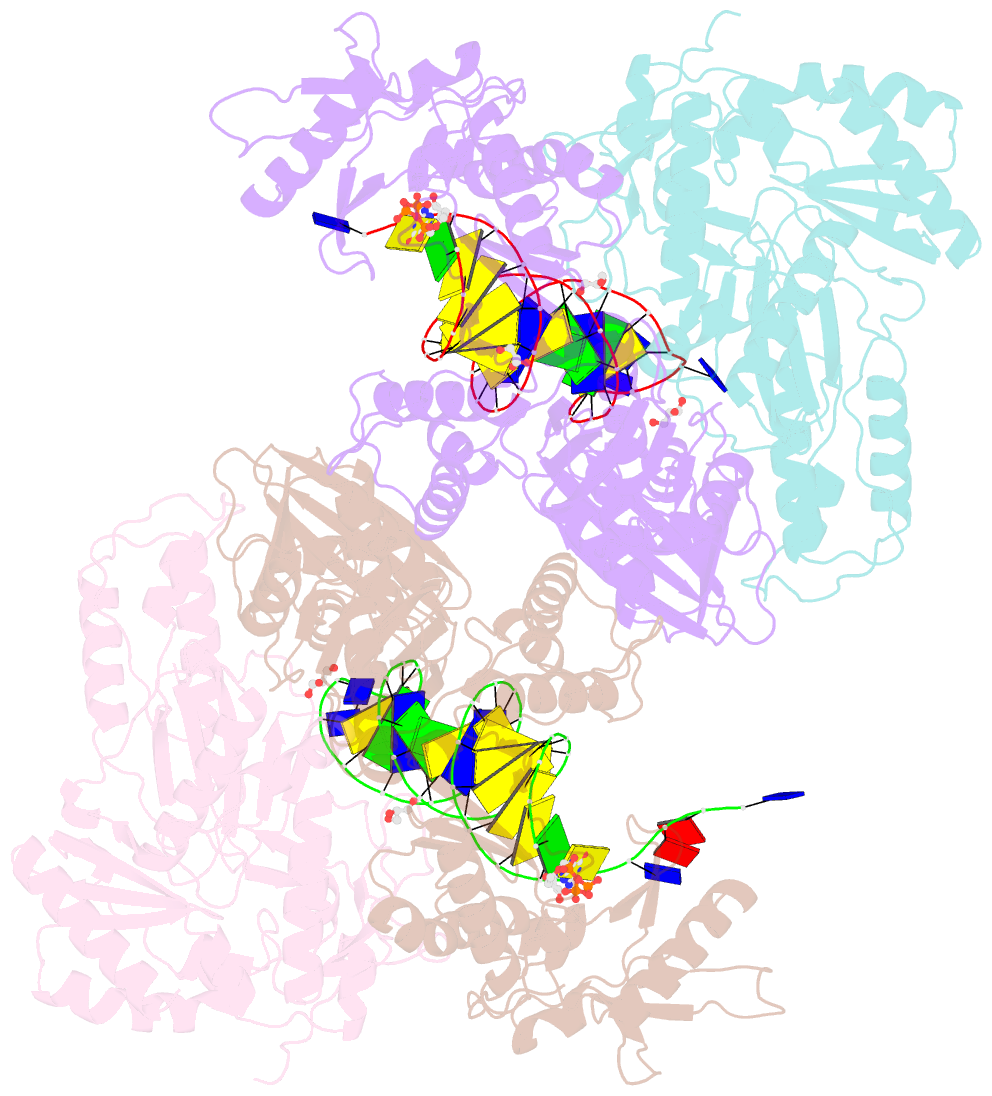
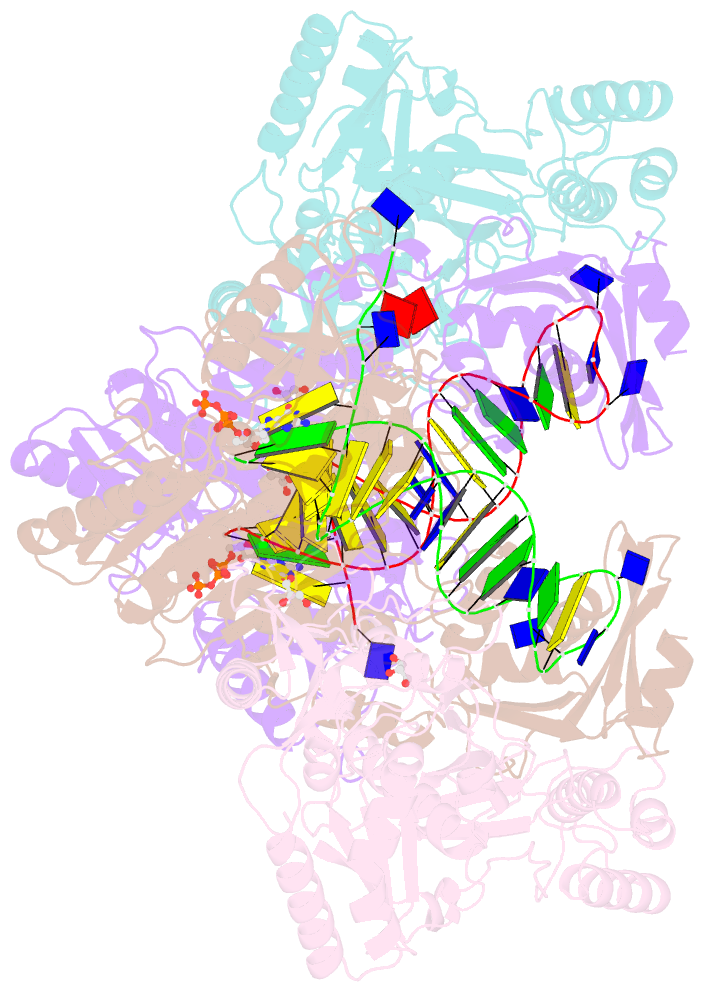
- Hiv-1 reverse transcriptase with q151m-y115f-f116y:DNA:dgtp ternary complex
- Reference
- Yasutake Y, Hattori SI, Tamura N, Matsuda K, Kohgo S, Maeda K, Mitsuya H (2020): "Structural features in common of HBV and HIV-1 resistance against chirally-distinct nucleoside analogues entecavir and lamivudine." Sci Rep, 10, 3021. doi: 10.1038/s41598-020-59775-w.
- Abstract
- Chronic hepatitis B virus (HBV) infection is a major public health problem that affects millions of people worldwide. Nucleoside analogue reverse transcriptase (RT) inhibitors, such as entecavir (ETV) and lamivudine (3TC), serve as crucial anti-HBV drugs. However, structural studies of HBV RT have been hampered due to its unexpectedly poor solubility. Here, we show that human immunodeficiency virus type-1 (HIV-1) with HBV-associated amino acid substitutions Y115F/F116Y/Q151M in its RT (HIVY115F/F116Y/Q151M) is highly susceptible to ETV and 3TC. Additionally, we experimentally simulated previously reported ETV/3TC resistance for HBV using HIVY115F/F116Y/Q151M with F160M/M184V (L180M/M204V in HBV RT) substituted. We determined crystal structures for HIV-1 RTY115F/F116Y/Q151M:DNA complexed with 3TC-triphosphate (3TC-TP)/ETV-triphosphate (ETV-TP)/dCTP/dGTP. These structures revealed an atypically tight binding conformation of 3TC-TP, where the Met184 side-chain is pushed away by the oxathiolane of 3TC-TP and exocyclic methylene of ETV-TP. Structural analysis of RTY115F/F116Y/Q151M/F160M/M184V:DNA:3TC-TP also demonstrated that the loosely bound 3TC-TP is misaligned at the active site to prevent a steric clash with the side chain γ-methyl of Val184. These findings shed light on the common structural mechanism of HBV and HIV-1 resistance to 3TC and ETV and should aid in the design of new agents to overcome drug resistance to 3TC and ETV.