Summary information and primary citation
- PDB-id
- 6knc; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- replication-DNA
- Method
- cryo-EM (9.3 Å)
- Summary
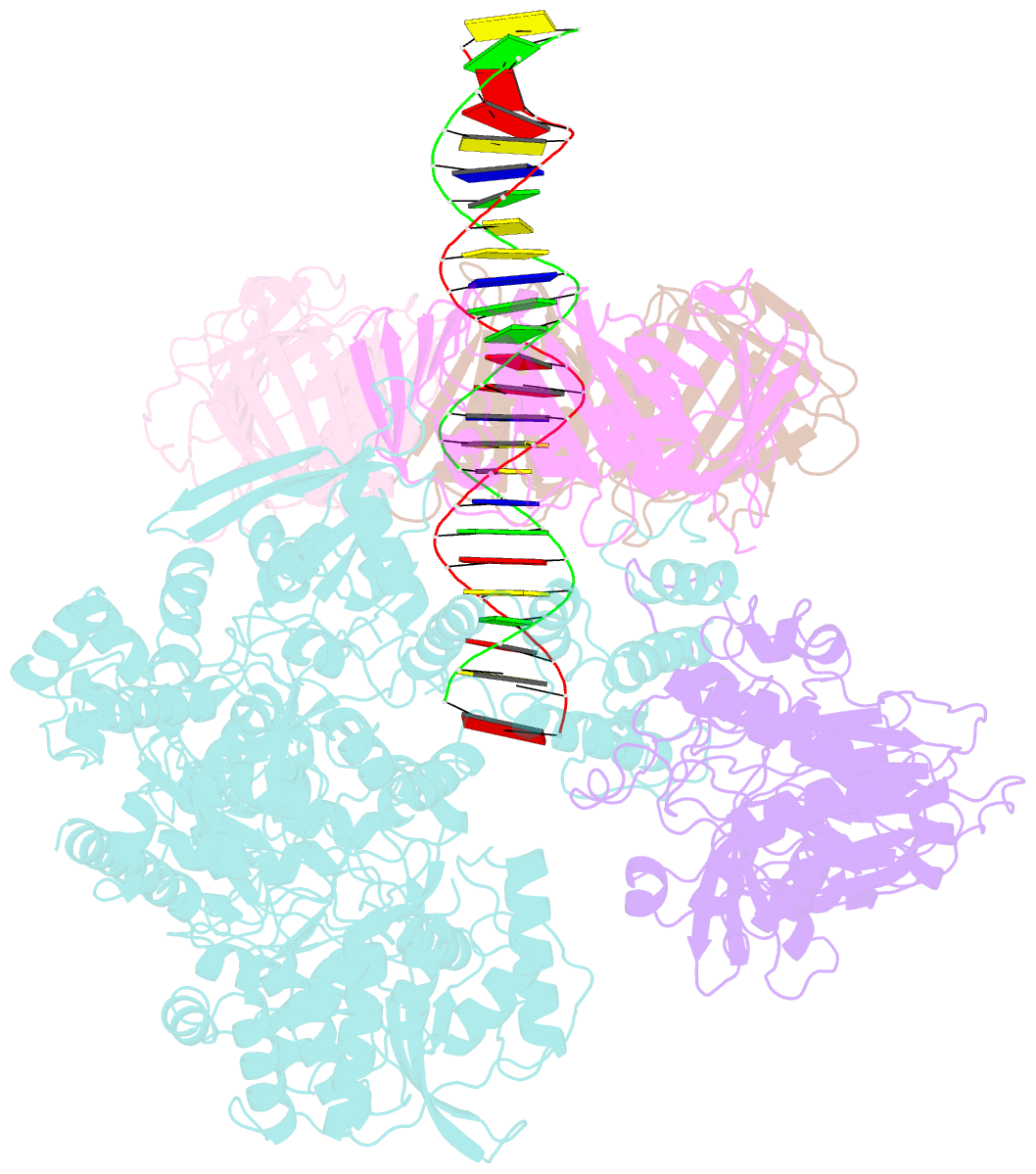
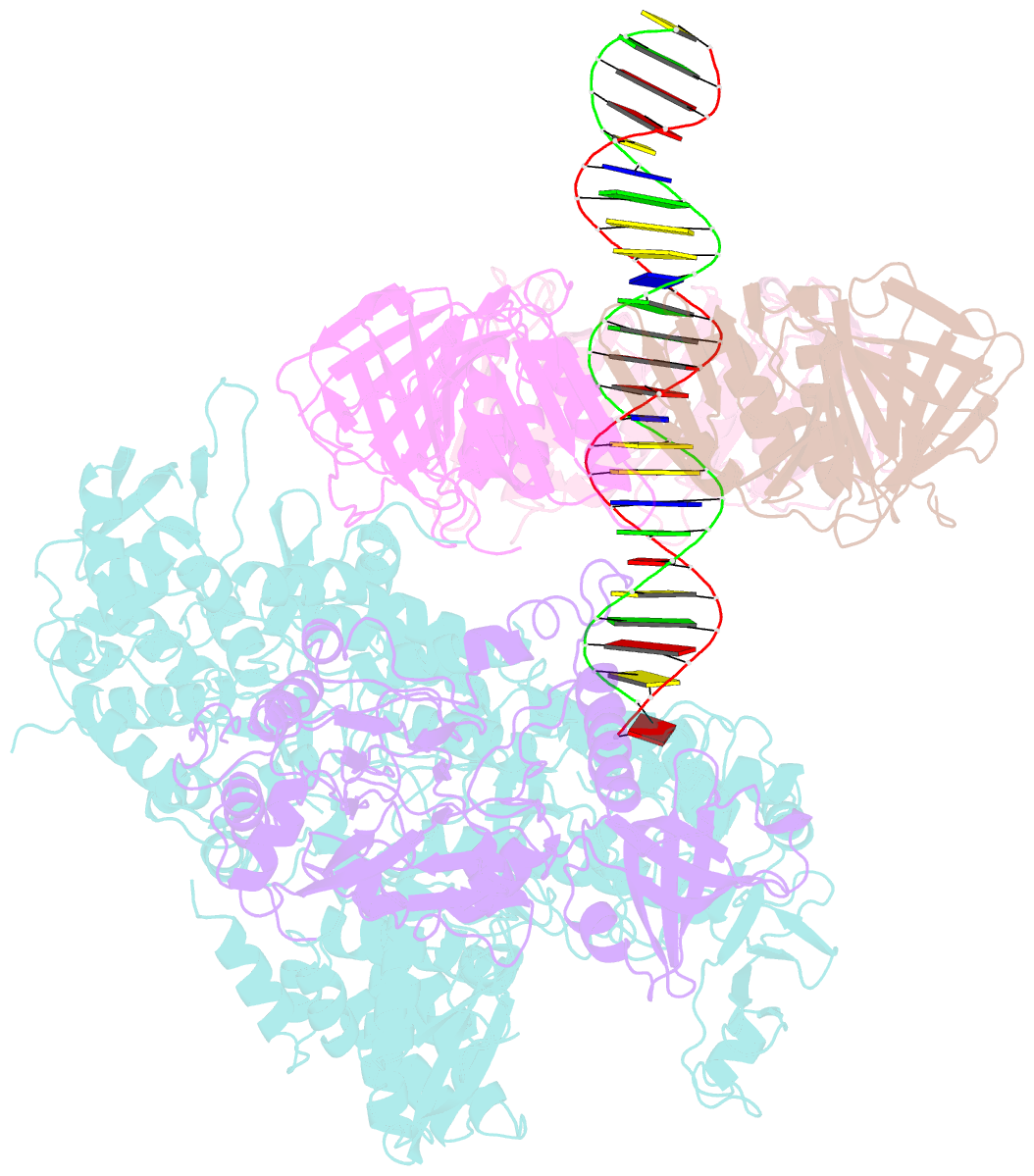
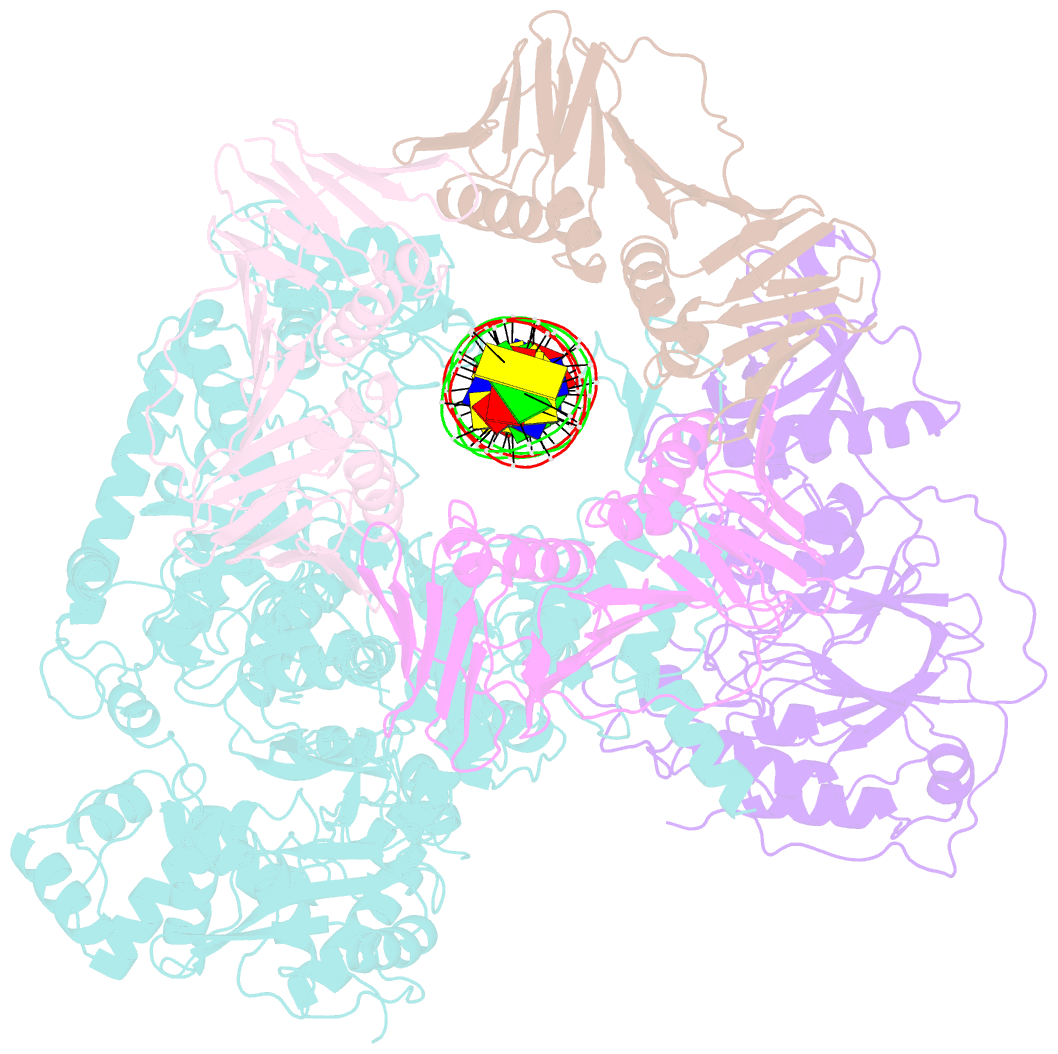
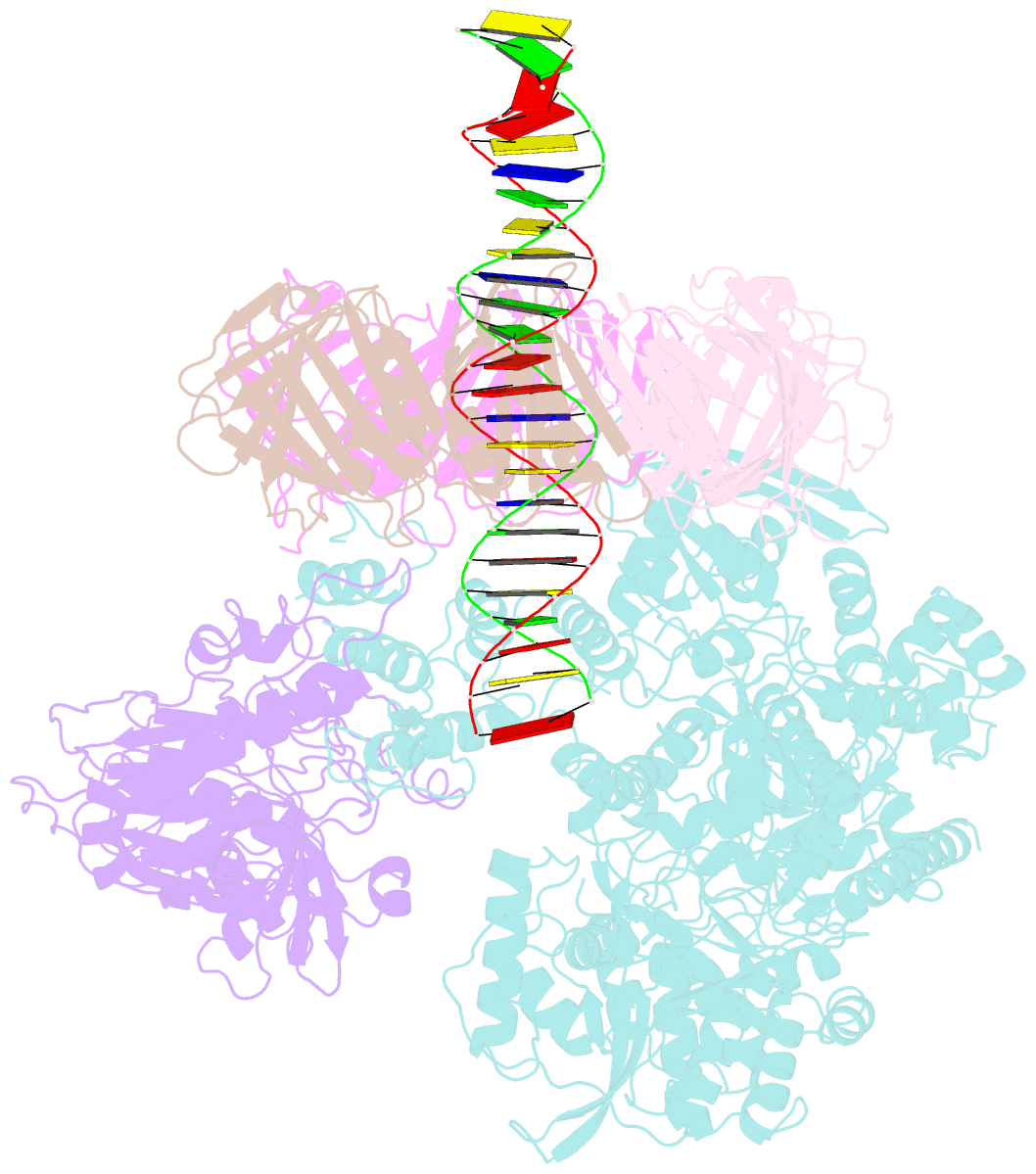
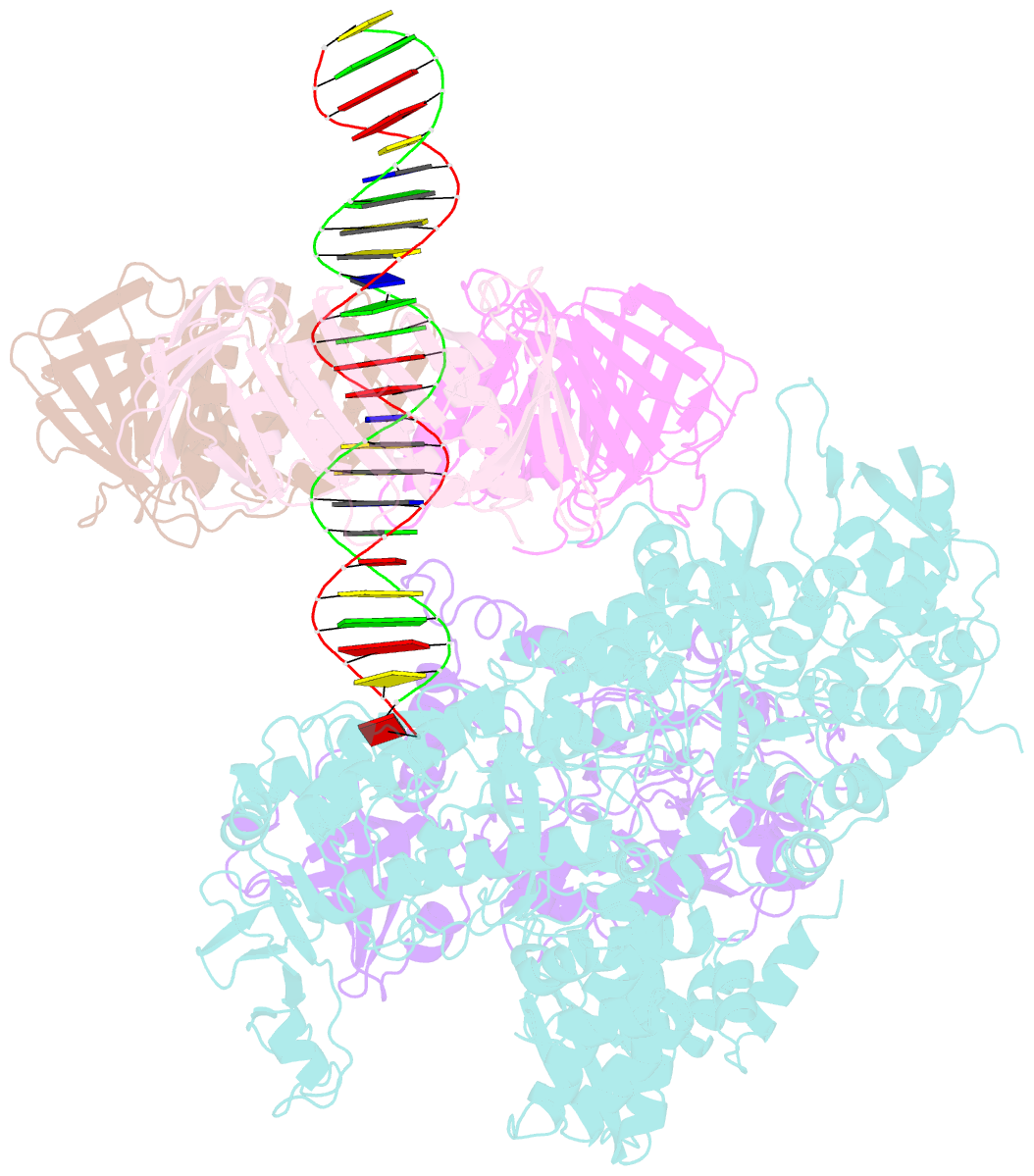
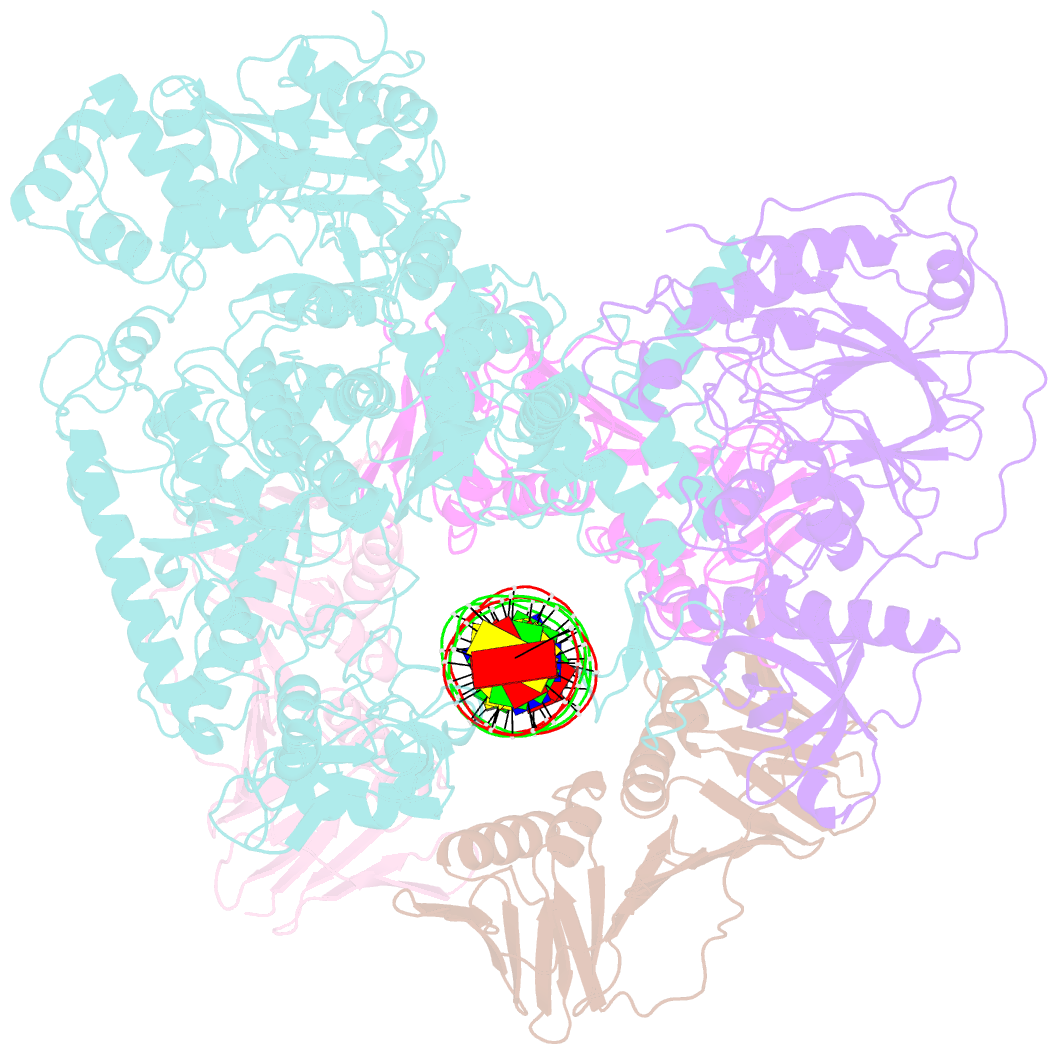
- Pold-pcna-DNA (form b)
- Reference
- Mayanagi K, Oki K, Miyazaki N, Ishino S, Yamagami T, Morikawa K, Iwasaki K, Kohda D, Shirai T, Ishino Y (2020): "Two conformations of DNA polymerase D-PCNA-DNA, an archaeal replisome complex, revealed by cryo-electron microscopy." Bmc Biol., 18, 152. doi: 10.1186/s12915-020-00889-y.
- Abstract
- Background: DNA polymerase D (PolD) is the representative member of the D family of DNA polymerases. It is an archaea-specific DNA polymerase required for replication and unrelated to other known DNA polymerases. PolD consists of a heterodimer of two subunits, DP1 and DP2, which contain catalytic sites for 3'-5' editing exonuclease and DNA polymerase activities, respectively, with both proteins being mutually required for the full activities of each enzyme. However, the processivity of the replicase holoenzyme has additionally been shown to be enhanced by the clamp molecule proliferating cell nuclear antigen (PCNA), making it crucial to elucidate the interaction between PolD and PCNA on a structural level for a full understanding of its functional relevance. We present here the 3D structure of a PolD-PCNA-DNA complex from Thermococcus kodakarensis using single-particle cryo-electron microscopy (EM).
Results: Two distinct forms of the PolD-PCNA-DNA complex were identified by 3D classification analysis. Fitting the reported crystal structures of truncated forms of DP1 and DP2 from Pyrococcus abyssi onto our EM map showed the 3D atomic structural model of PolD-PCNA-DNA. In addition to the canonical interaction between PCNA and PolD via PIP (PCNA-interacting protein)-box motif, we found a new contact point consisting of a glutamate residue at position 171 in a β-hairpin of PCNA, which mediates interactions with DP1 and DP2. The DNA synthesis activity of a mutant PolD with disruption of the E171-mediated PCNA interaction was not stimulated by PCNA in vitro.
Conclusions: Based on our analyses, we propose that glutamate residues at position 171 in each subunit of the PCNA homotrimer ring can function as hooks to lock PolD conformation on PCNA for conversion of its activity. This hook function of the clamp molecule may be conserved in the three domains of life.