Summary information and primary citation
- PDB-id
- 6mcf; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription
- Method
- NMR
- Summary
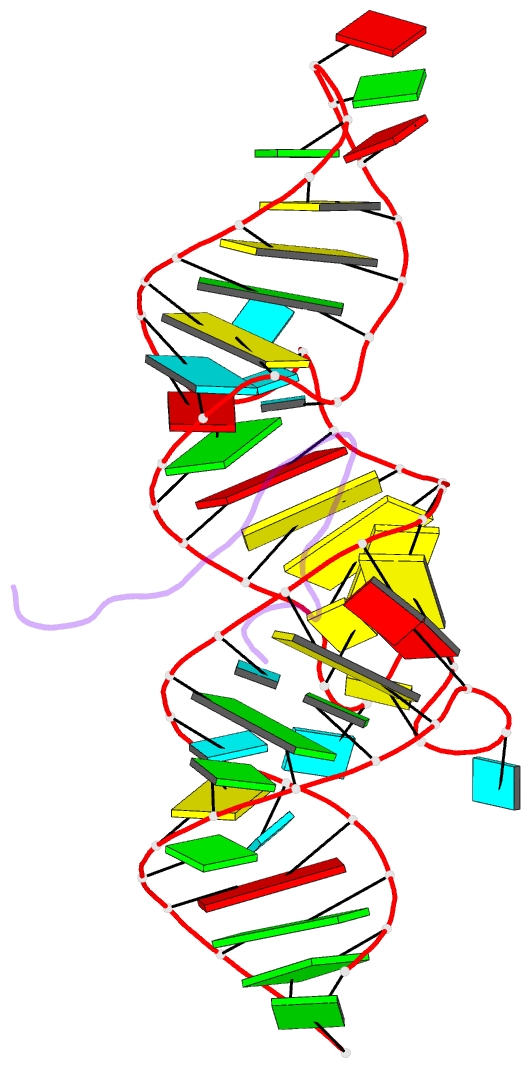
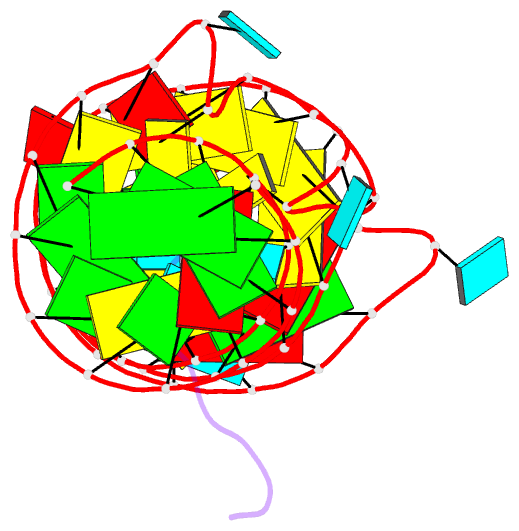
- Solution structure of 7sk stem-loop 1 with hiv-1 tat RNA binding domain
- Reference
- Pham VV, Salguero C, Khan SN, Meagher JL, Brown WC, Humbert N, de Rocquigny H, Smith JL, D'Souza VM (2018): "HIV-1 Tat interactions with cellular 7SK and viral TAR RNAs identifies dual structural mimicry." Nat Commun, 9, 4266. doi: 10.1038/s41467-018-06591-6.
- Abstract
- The HIV Tat protein competes with the 7SK:HEXIM interaction to hijack pTEFb from 7SK snRNP and recruit it to the TAR motif on stalled viral transcripts. Here we solve structures of 7SK stemloop-1 and TAR in complex with Tat's RNA binding domain (RBD) to gain insights into this process. We find that 7SK is peppered with arginine sandwich motifs (ASM)-three classical and one with a pseudo configuration. Despite having similar RBDs, the presence of an additional arginine, R52, confers Tat the ability to remodel the pseudo configuration, required for HEXIM binding, into a classical sandwich, thus displacing HEXIM. Tat also uses R52 to remodel the TAR bulge into an ASM whose structure is identical to that of the remodeled ASM in 7SK. Together, our structures reveal a dual structural mimicry wherein viral Tat and TAR have co-opted structural motifs present in cellular HEXIM and 7SK for productive transcription of its genome.