Summary information and primary citation
- PDB-id
- 6nme; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- unknown function-RNA
- Method
- cryo-EM (5.67 Å)
- Summary
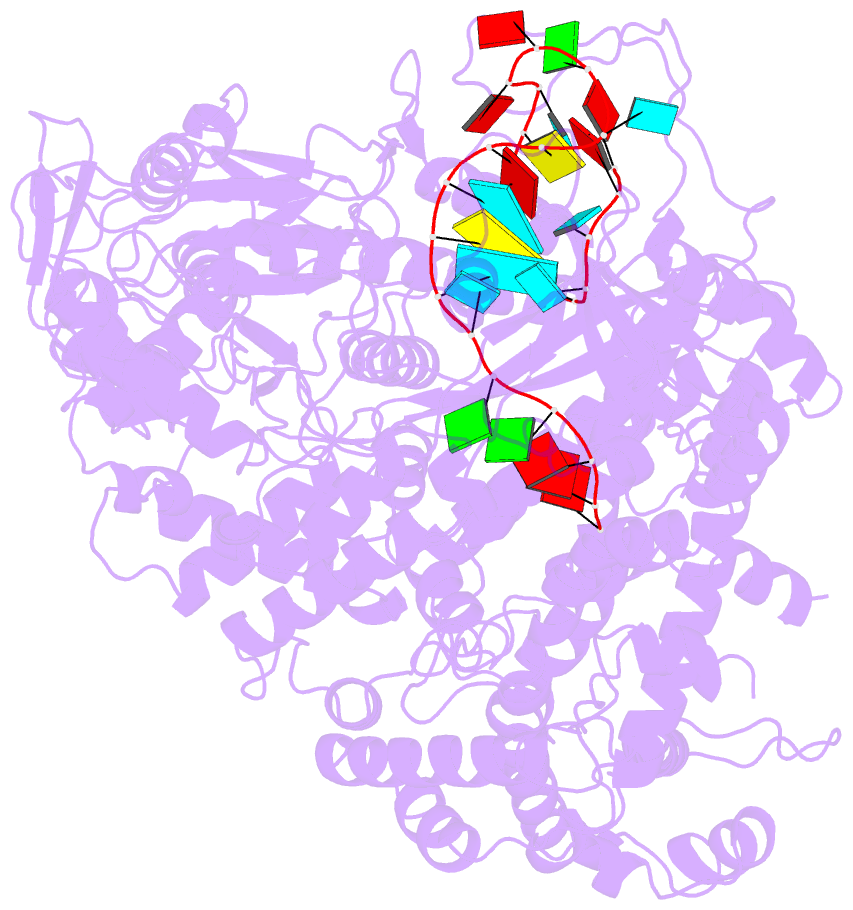
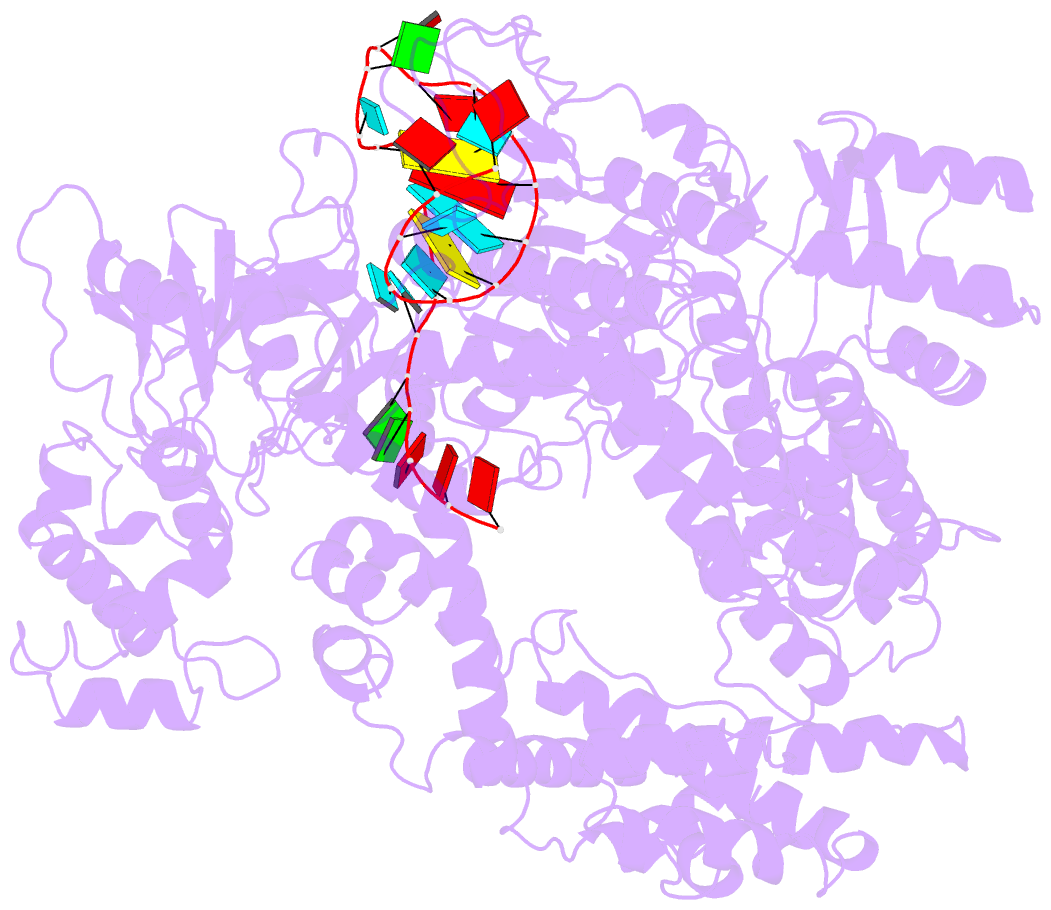
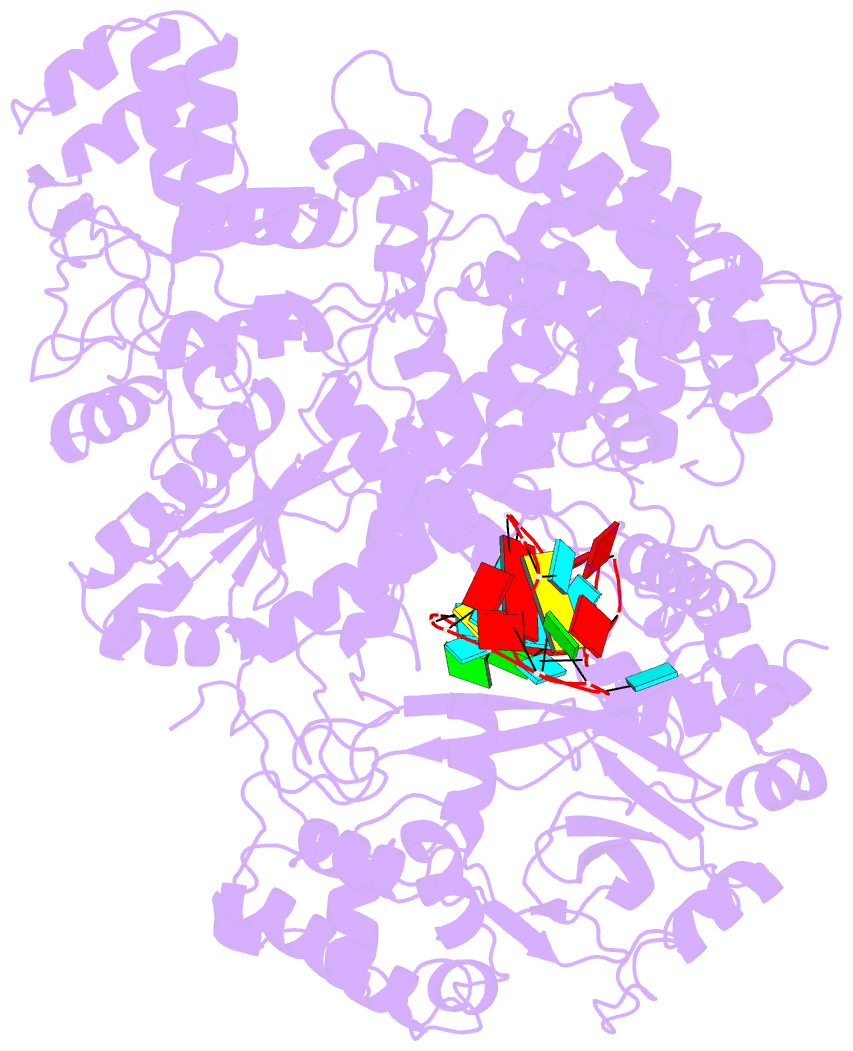
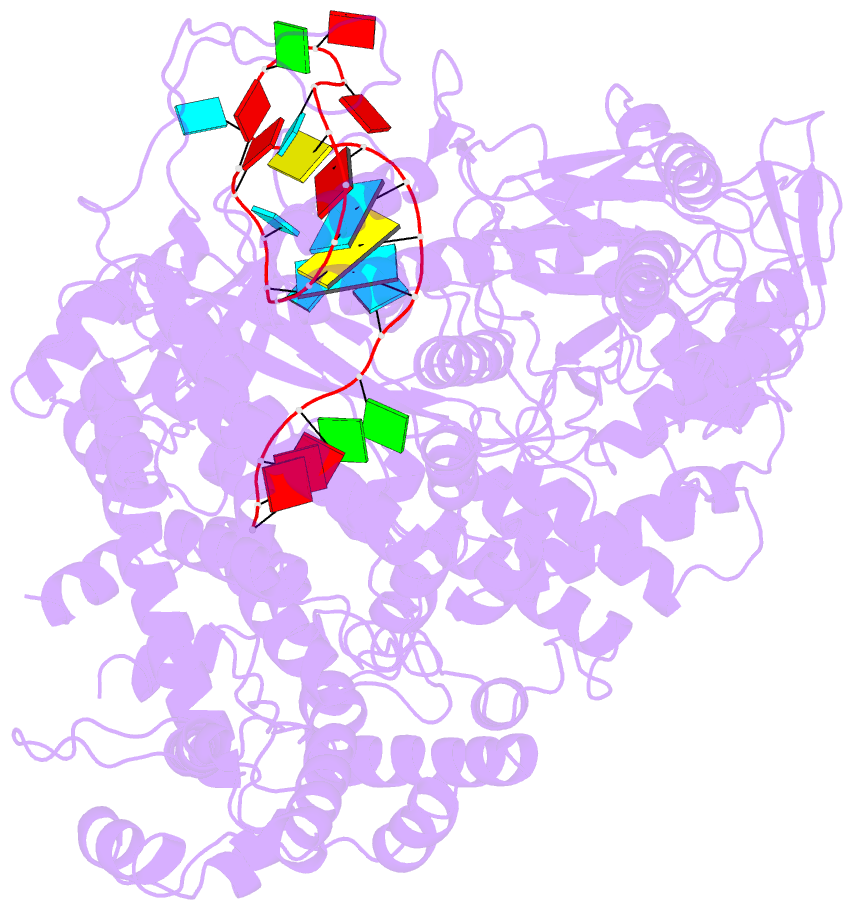
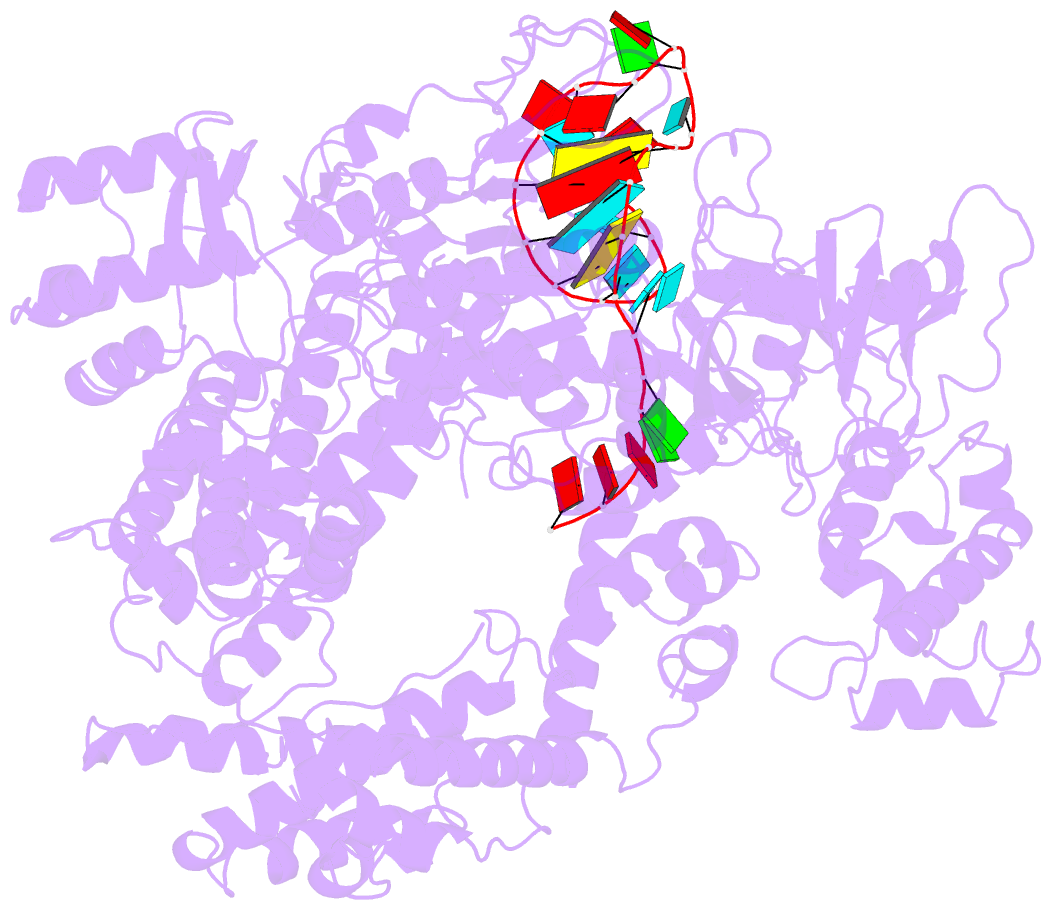
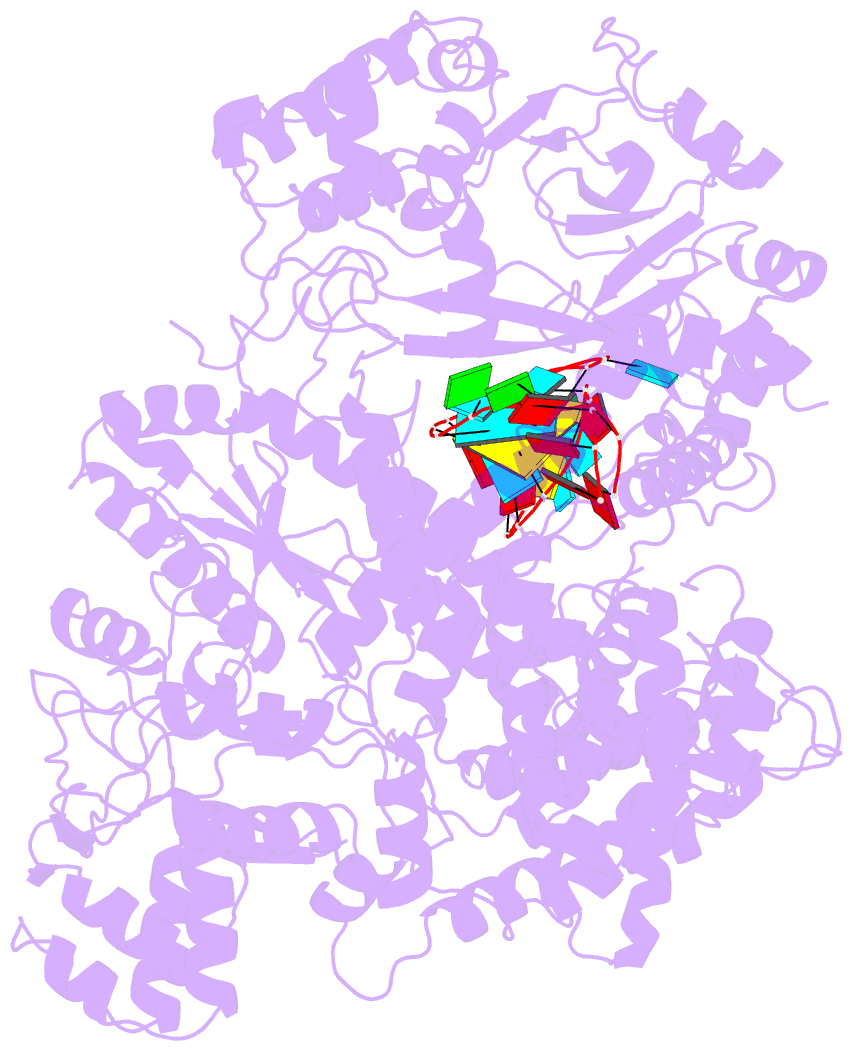
- Structure of lbcas12a-crrna
- Reference
- Zhang H, Li Z, Daczkowski CM, Gabel C, Mesecar AD, Chang L (2019): "Structural Basis for the Inhibition of CRISPR-Cas12a by Anti-CRISPR Proteins." Cell Host Microbe, 25, 815. doi: 10.1016/j.chom.2019.05.004.
- Abstract
- CRISPR-Cas12a (Cpf1), a type V CRISPR-associated nuclease, provides bacterial immunity against bacteriophages and plasmids but also serves as a tool for genome editing. Foreign nucleic acids are integrated into the CRISPR locus, prompting transcription of CRISPR RNAs (crRNAs) that guide Cas12a cleavage of foreign complementary DNA. However, mobile genetic elements counteract Cas12a with inhibitors, notably type V-A anti-CRISPRs (AcrVAs). We present cryoelectron microscopy structures of Cas12a-crRNA bound to AcrVA1 and AcrVA4 at 3.5 and 3.3 Å resolutions, respectively. AcrVA1 is sandwiched between the recognition (REC) and nuclease (NUC) lobes of Cas12a and inserts into the binding pocket for the protospacer-adjacent motif (PAM), a short DNA sequence guiding Cas12a targeting. AcrVA1 cleaves crRNA in a Cas12a-dependent manner, inactivating Cas12a-crRNA complexes. The AcrVA4 dimer is anchored around the crRNA pseudoknot of Cas12a-crRNA, preventing required conformational changes for crRNA-DNA heteroduplex formation. These results uncover molecular mechanisms for CRISPR-Cas12a inhibition, providing insights into bacteria-phage dynamics.