Summary information and primary citation
- PDB-id
- 6rax; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydolase
- Method
- cryo-EM (3.99 Å)
- Summary
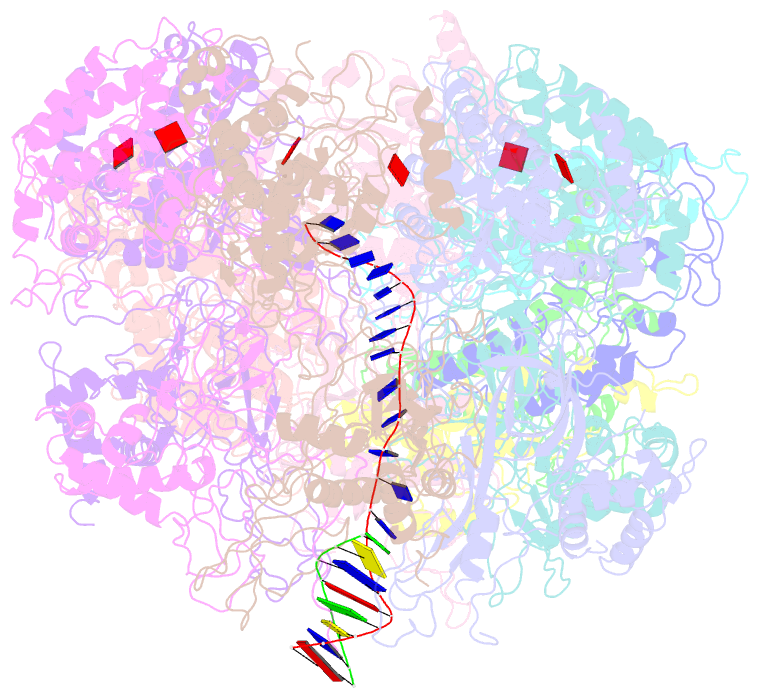
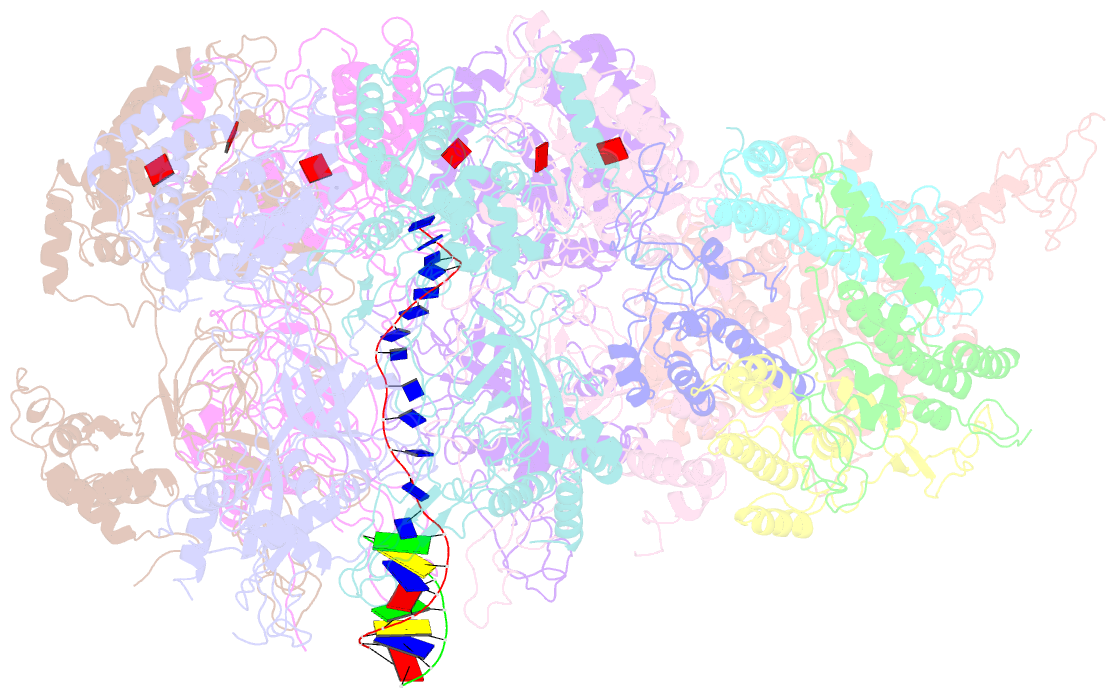
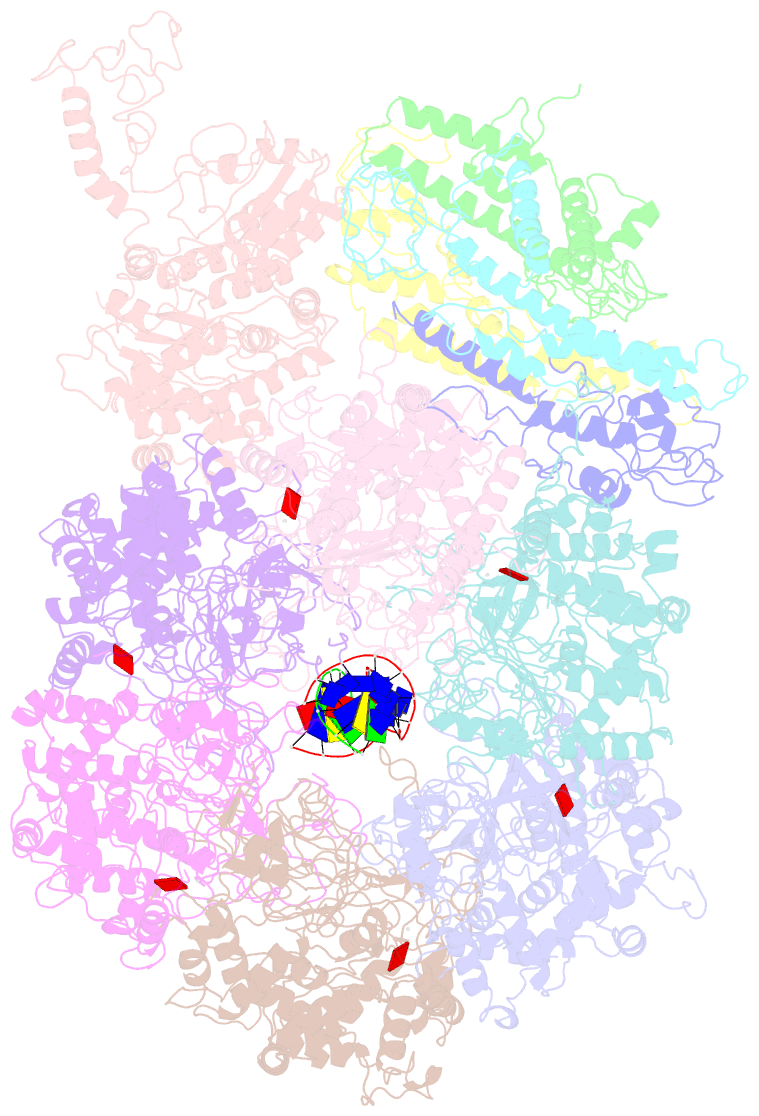
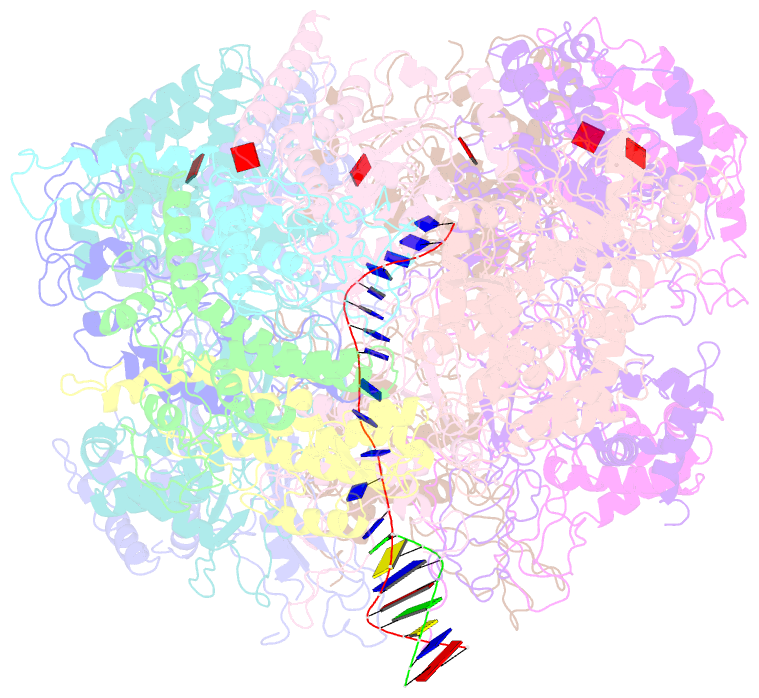
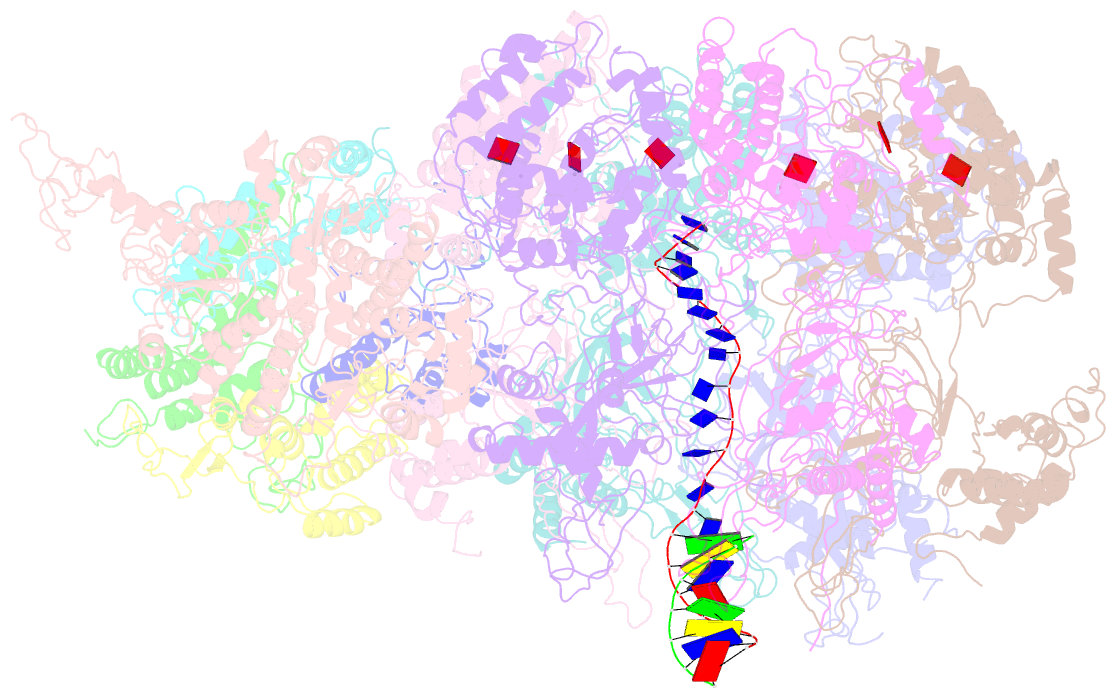
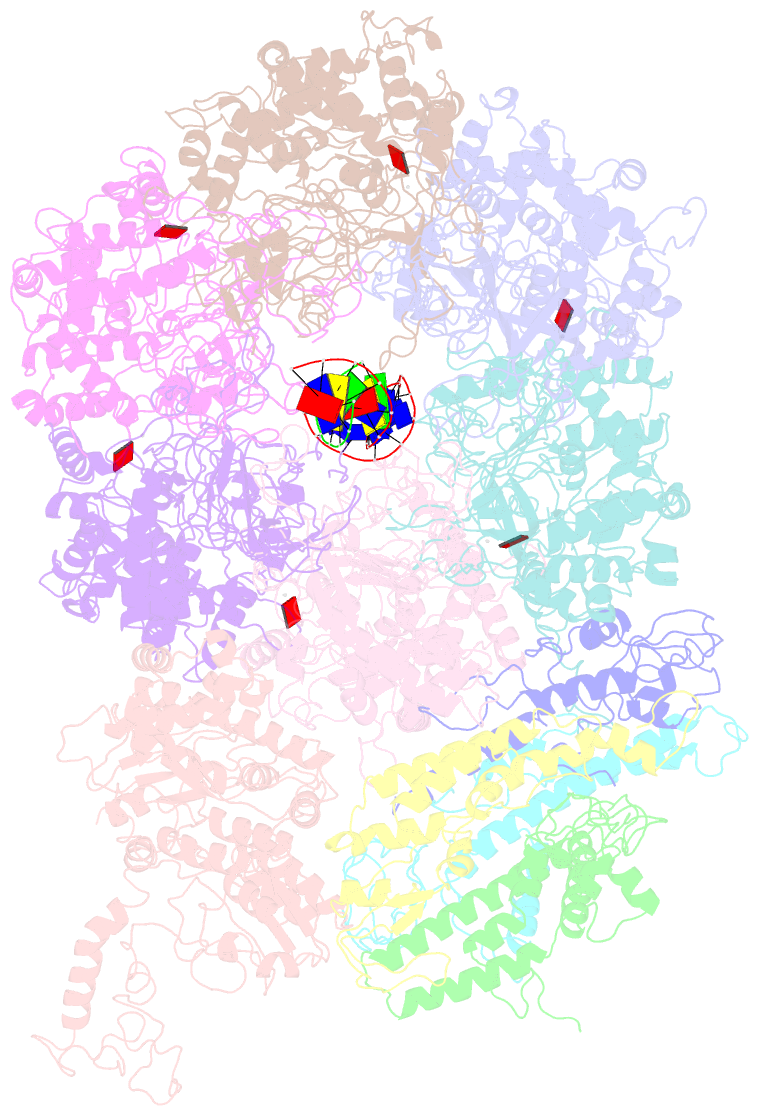
- D. melanogaster cmg-DNA, state 1b
- Reference
- Eickhoff P, Kose HB, Martino F, Petojevic T, Abid Ali F, Locke J, Tamberg N, Nans A, Berger JM, Botchan MR, Yardimci H, Costa A (2019): "Molecular Basis for ATP-Hydrolysis-Driven DNA Translocation by the CMG Helicase of the Eukaryotic Replisome." Cell Rep, 28, 2673-2688.e8. doi: 10.1016/j.celrep.2019.07.104.
- Abstract
- In the eukaryotic replisome, DNA unwinding by the Cdc45-MCM-Go-Ichi-Ni-San (GINS) (CMG) helicase requires a hexameric ring-shaped ATPase named minichromosome maintenance (MCM), which spools single-stranded DNA through its central channel. Not all six ATPase sites are required for unwinding; however, the helicase mechanism is unknown. We imaged ATP-hydrolysis-driven translocation of the CMG using cryo-electron microscopy (cryo-EM) and found that the six MCM subunits engage DNA using four neighboring protomers at a time, with ATP binding promoting DNA engagement. Morphing between different helicase states leads us to suggest a non-symmetric hand-over-hand rotary mechanism, explaining the asymmetric requirements of ATPase function around the MCM ring of the CMG. By imaging of a higher-order replisome assembly, we find that the Mrc1-Csm3-Tof1 fork-stabilization complex strengthens the interaction between parental duplex DNA and the CMG at the fork, which might support the coupling between DNA translocation and fork unwinding.