Summary information and primary citation
- PDB-id
- 6s48; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- hydrolase
- Method
- X-ray (1.9 Å)
- Summary
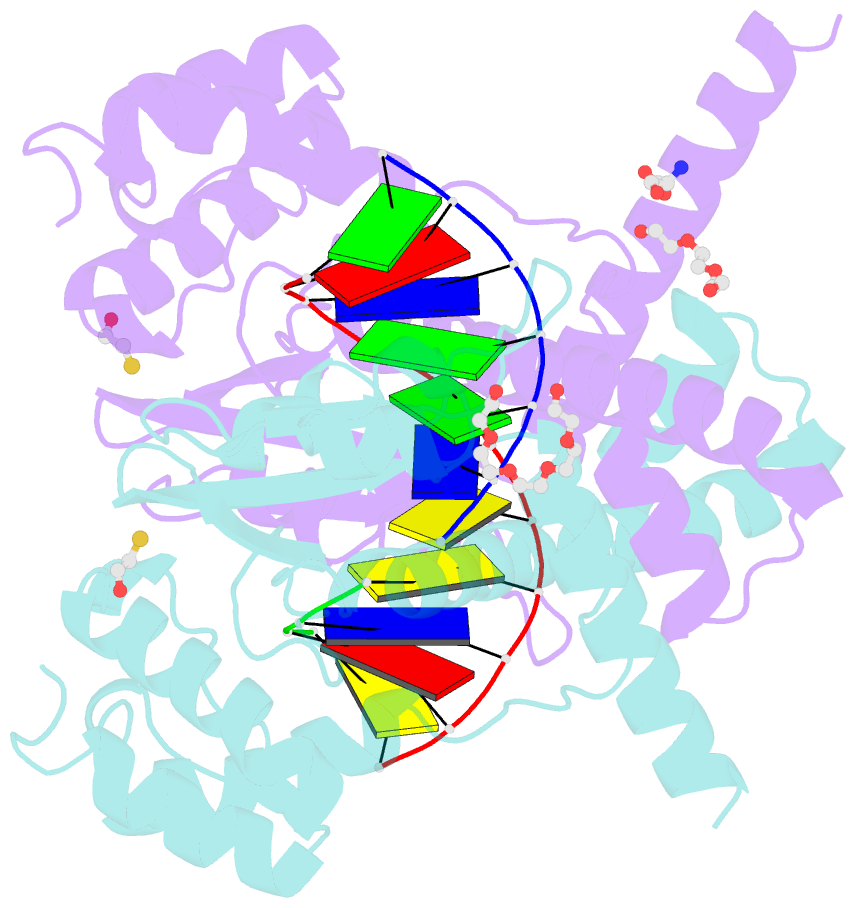
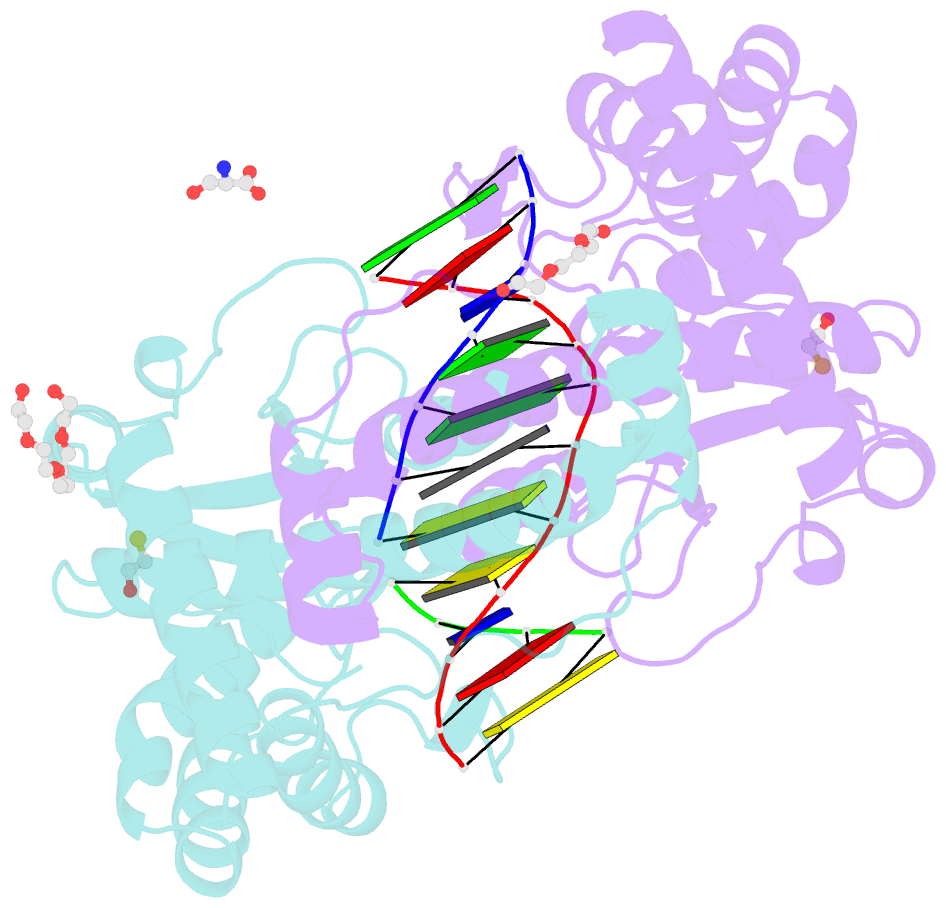
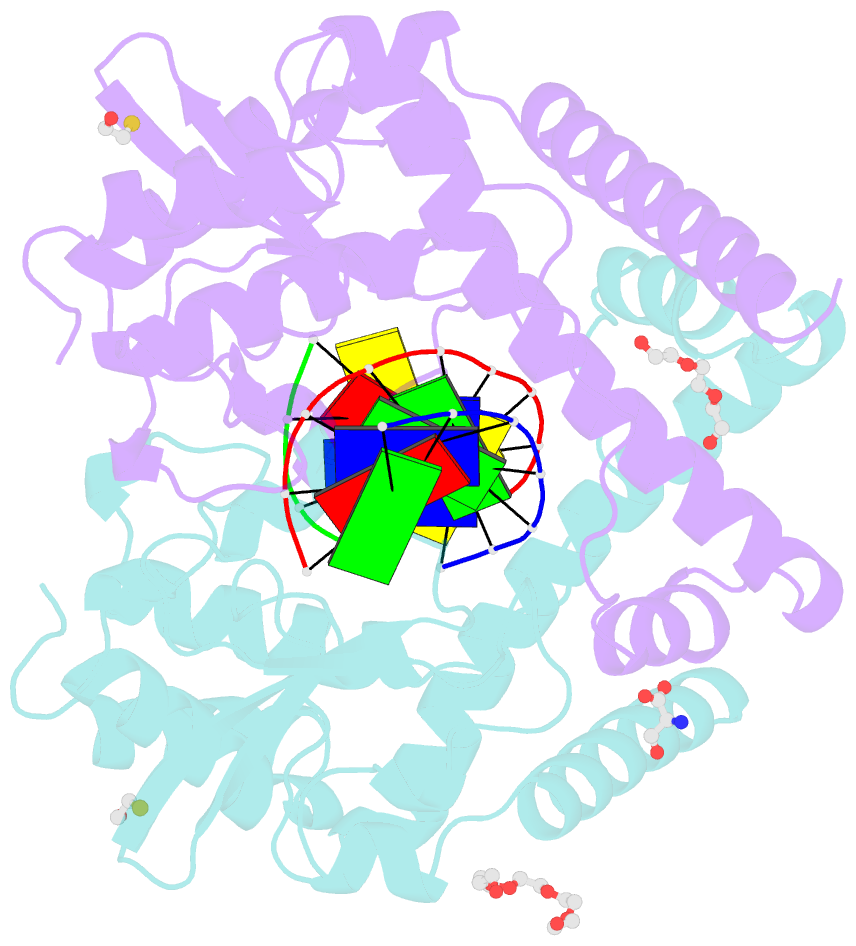
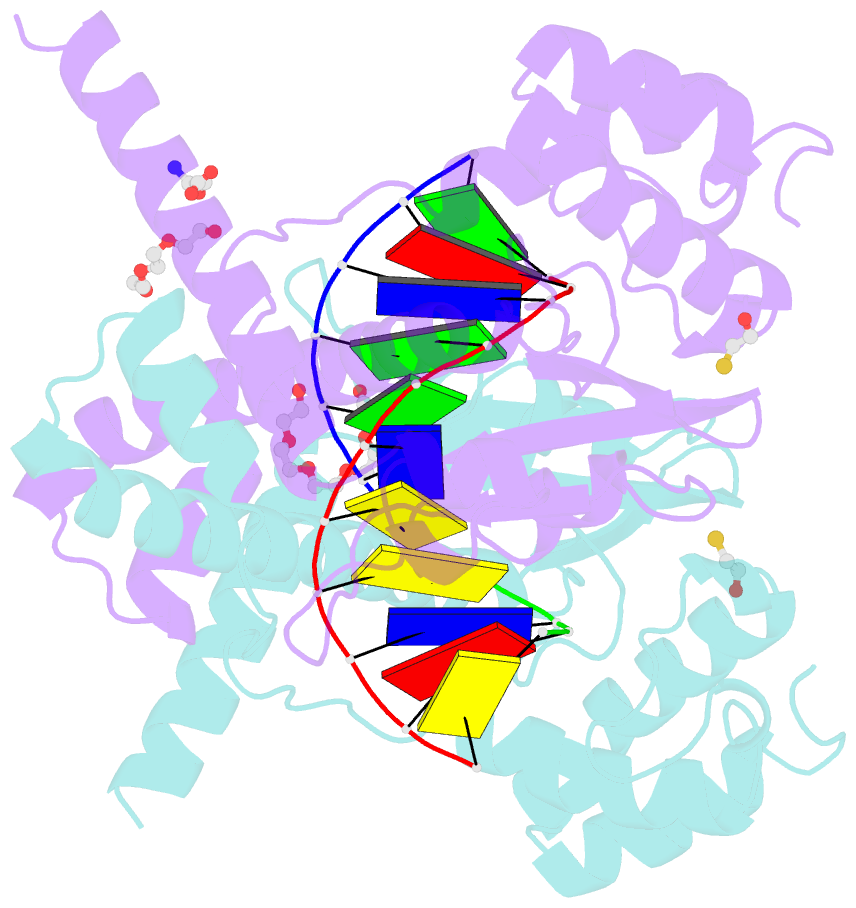
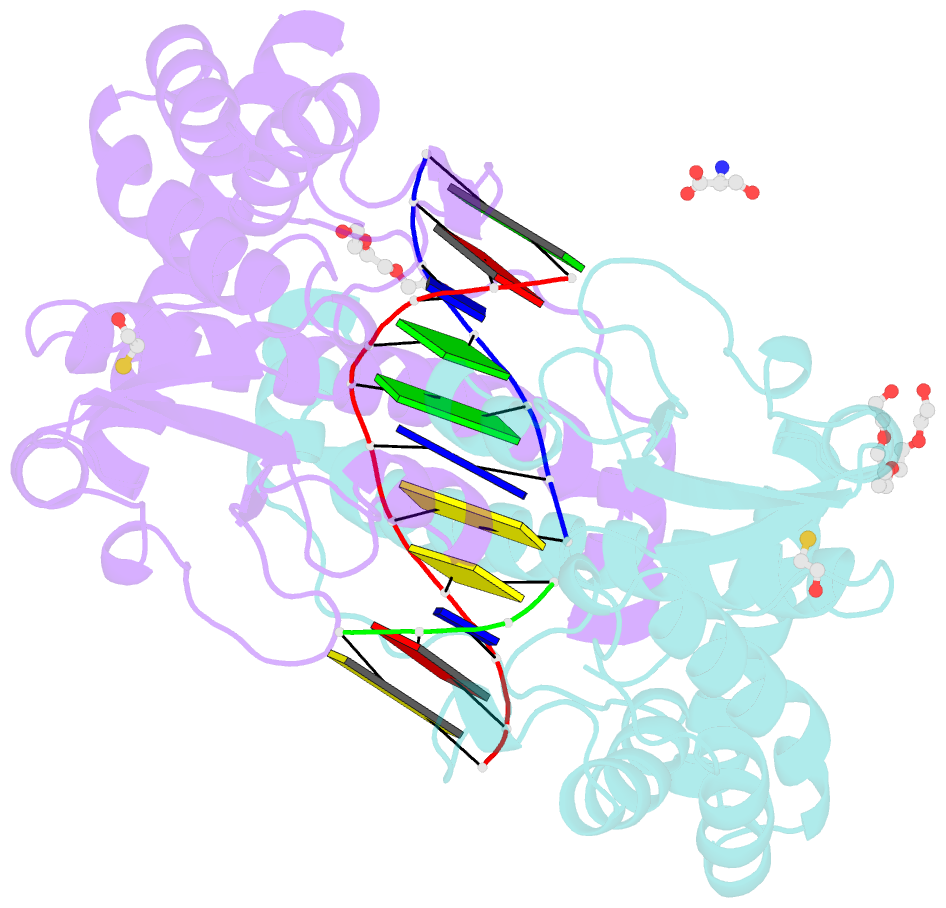
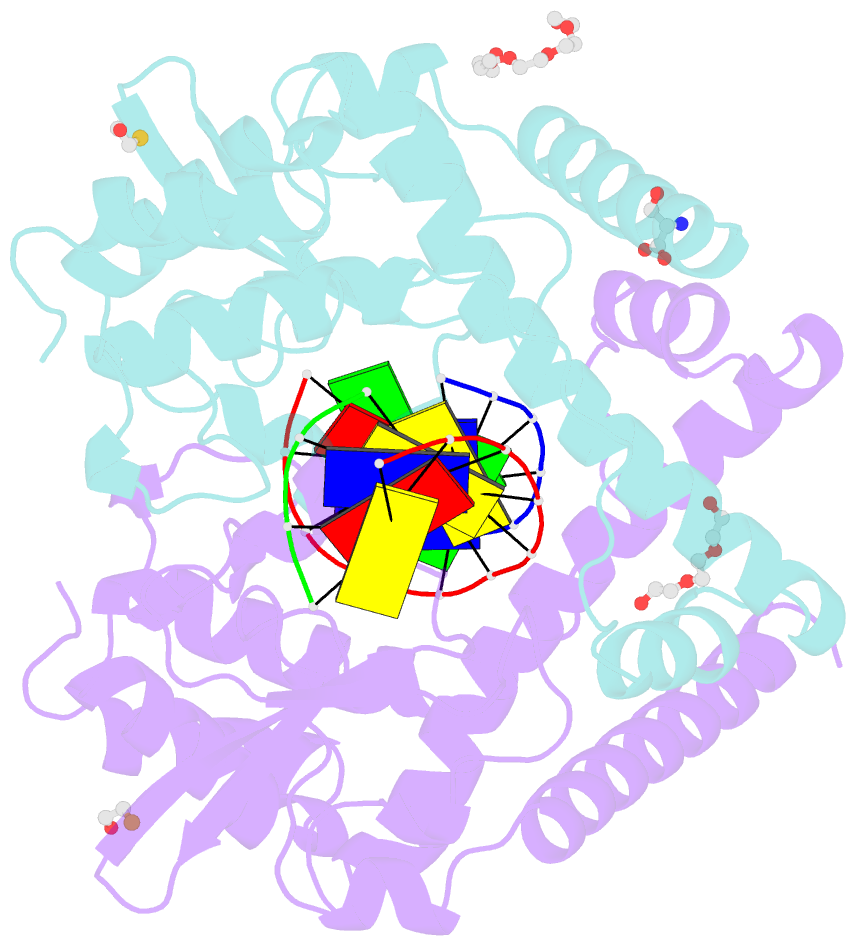
- Avaii restriction endonuclease in complex with partially cleaved dsDNA
- Reference
- Kisiala M, Kowalska M, Pastor M, Korza HJ, Czapinska H, Bochtler M (2020): "Restriction endonucleases that cleave RNA/DNA heteroduplexes bind dsDNA in A-like conformation." Nucleic Acids Res., 48, 6954-6969. doi: 10.1093/nar/gkaa403.
- Abstract
- Restriction endonucleases naturally target DNA duplexes. Systematic screening has identified a small minority of these enzymes that can also cleave RNA/DNA heteroduplexes and that may therefore be useful as tools for RNA biochemistry. We have chosen AvaII (G↓GWCC, where W stands for A or T) as a representative of this group of restriction endonucleases for detailed characterization. Here, we report crystal structures of AvaII alone, in specific complex with partially cleaved dsDNA, and in scanning complex with an RNA/DNA hybrid. The specific complex reveals a novel form of semi-specific dsDNA readout by a hexa-coordinated metal cation, most likely Ca2+ or Mg2+. Substitutions of residues anchoring this non-catalytic metal ion severely impair DNA binding and cleavage. The dsDNA in the AvaII complex is in the A-like form. This creates space for 2'-OH groups to be accommodated without intra-nucleic acid steric conflicts. PD-(D/E)XK restriction endonucleases of known structure that bind their dsDNA targets in the A-like form cluster into structurally similar groups. Most such enzymes, including some not previously studied in this respect, cleave RNA/DNA heteroduplexes. We conclude that A-form dsDNA binding is a good predictor for RNA/DNA cleavage activity.