Summary information and primary citation
- PDB-id
- 6uxw; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription-DNA
- Method
- cryo-EM (8.96 Å)
- Summary
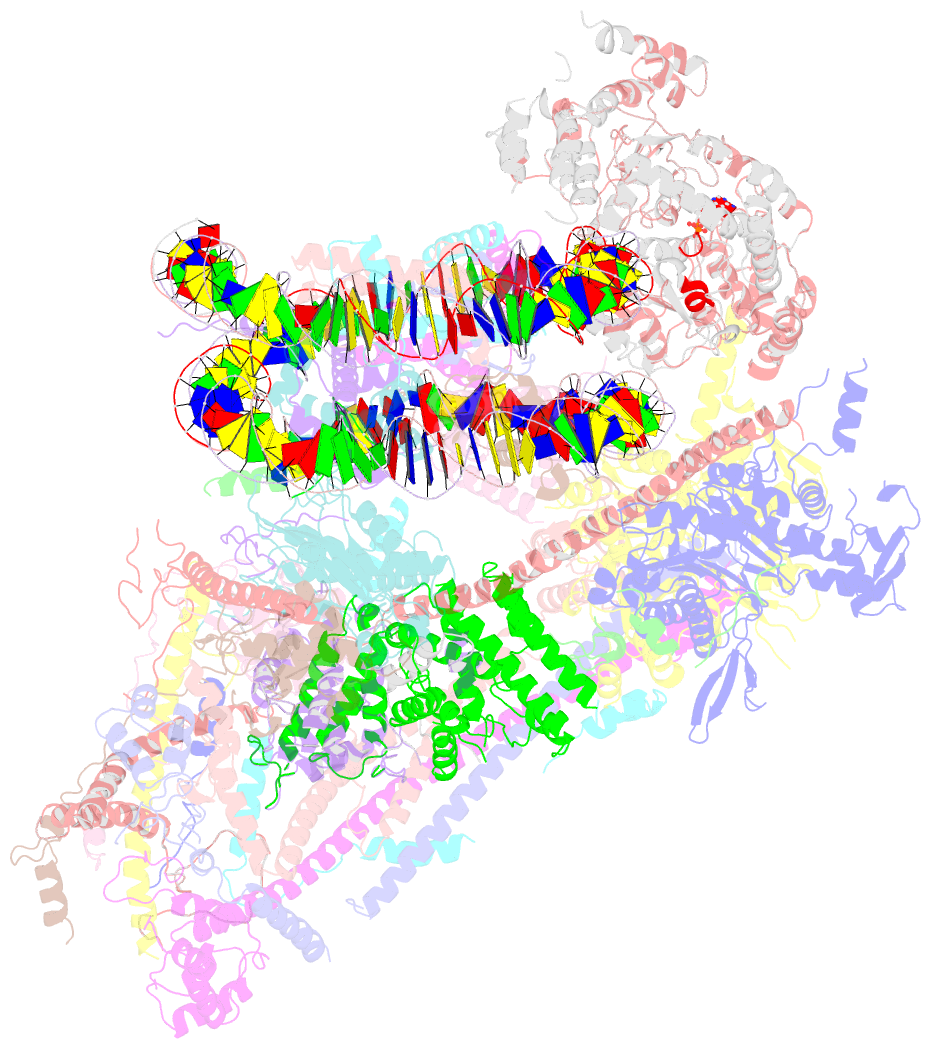
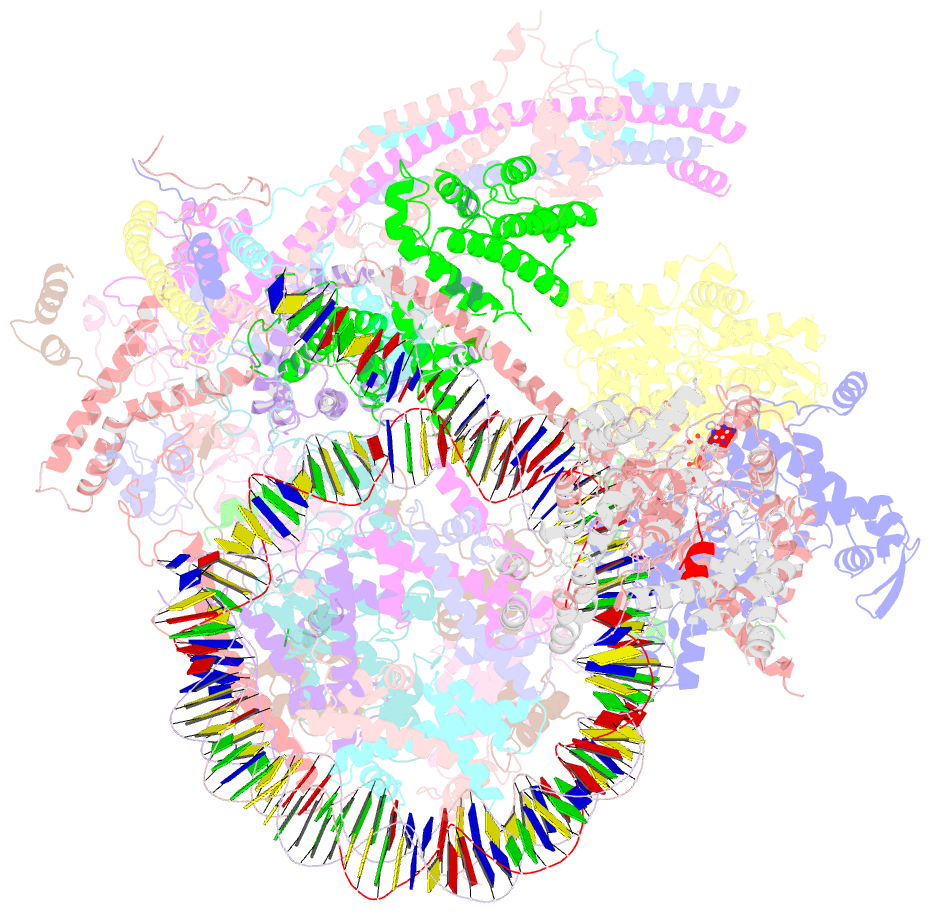
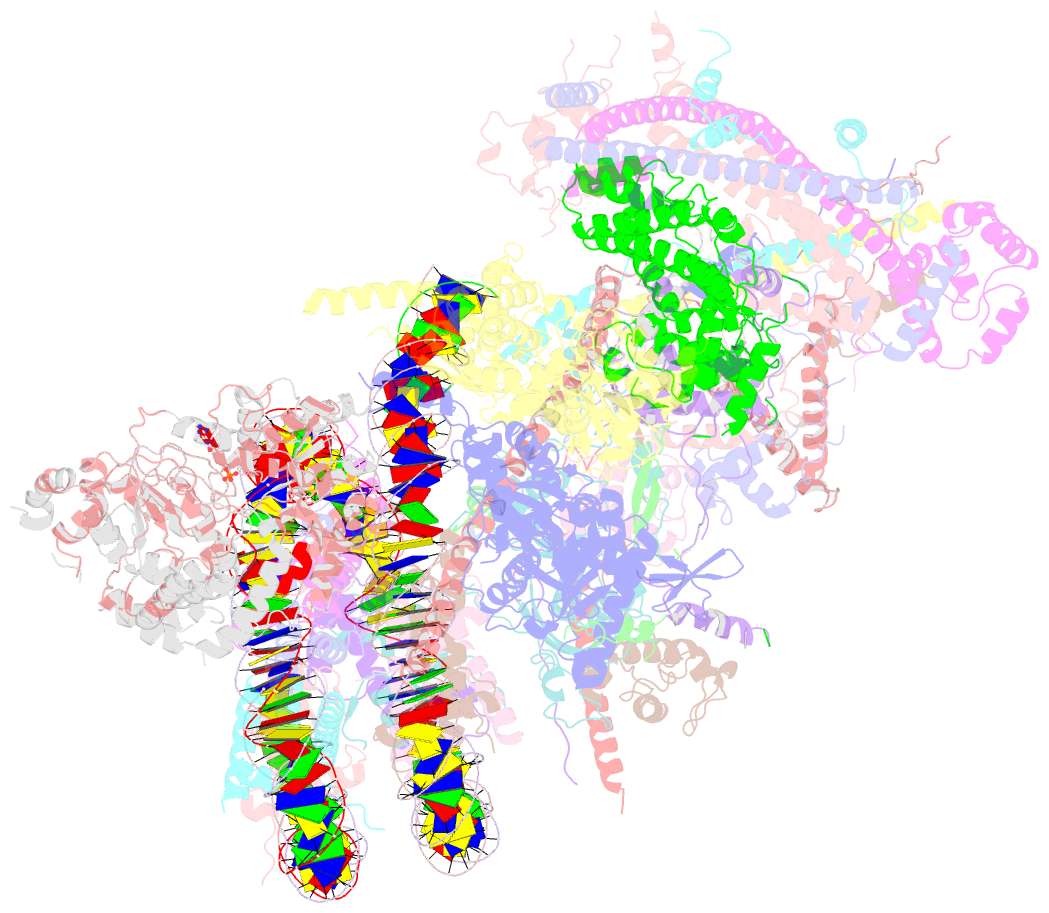
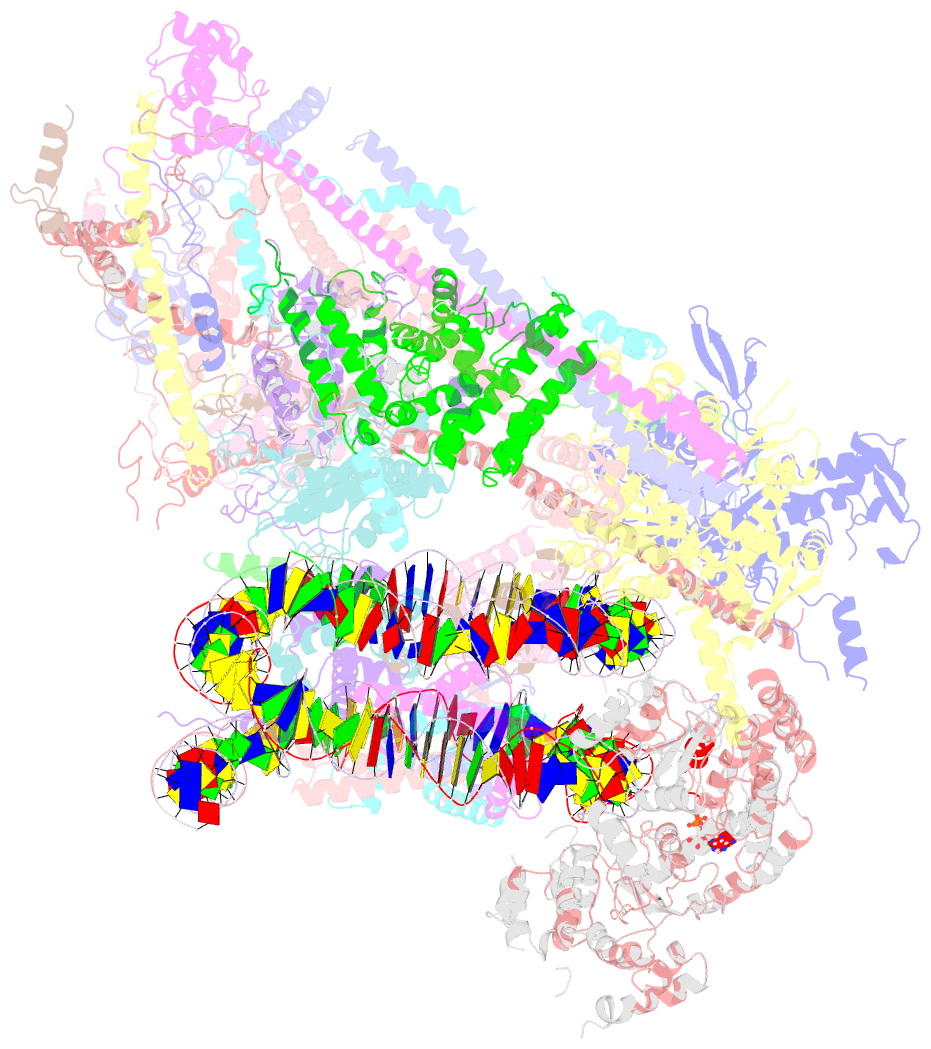
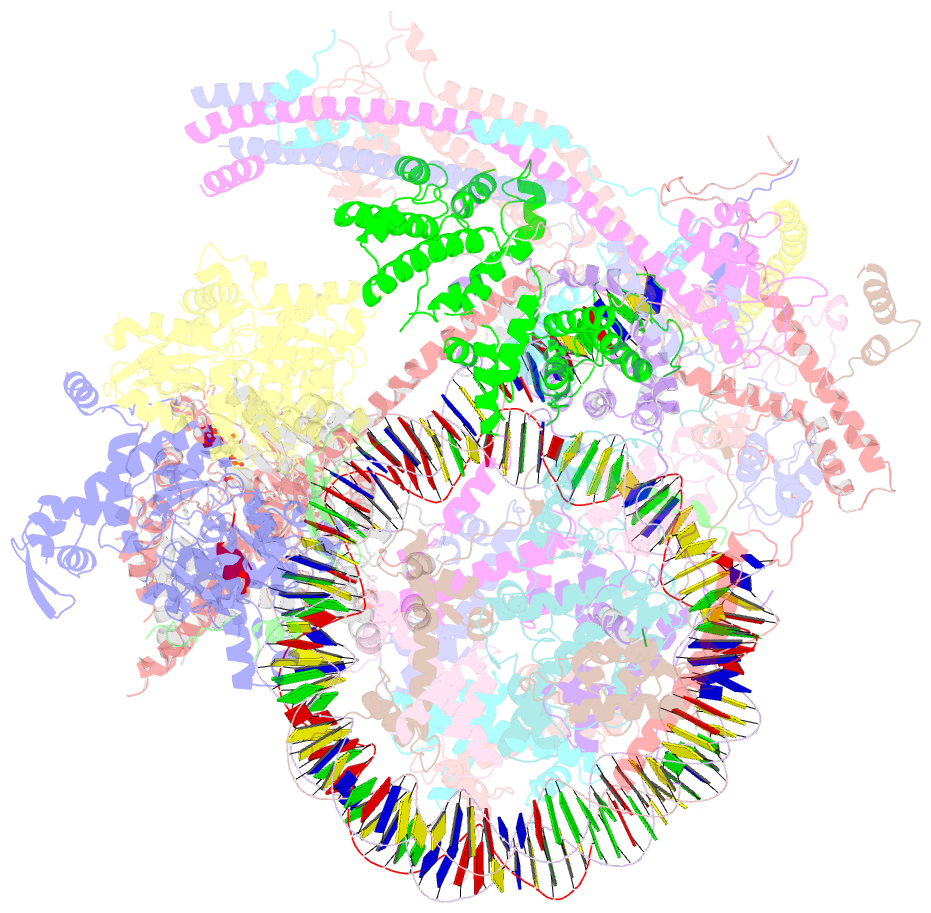
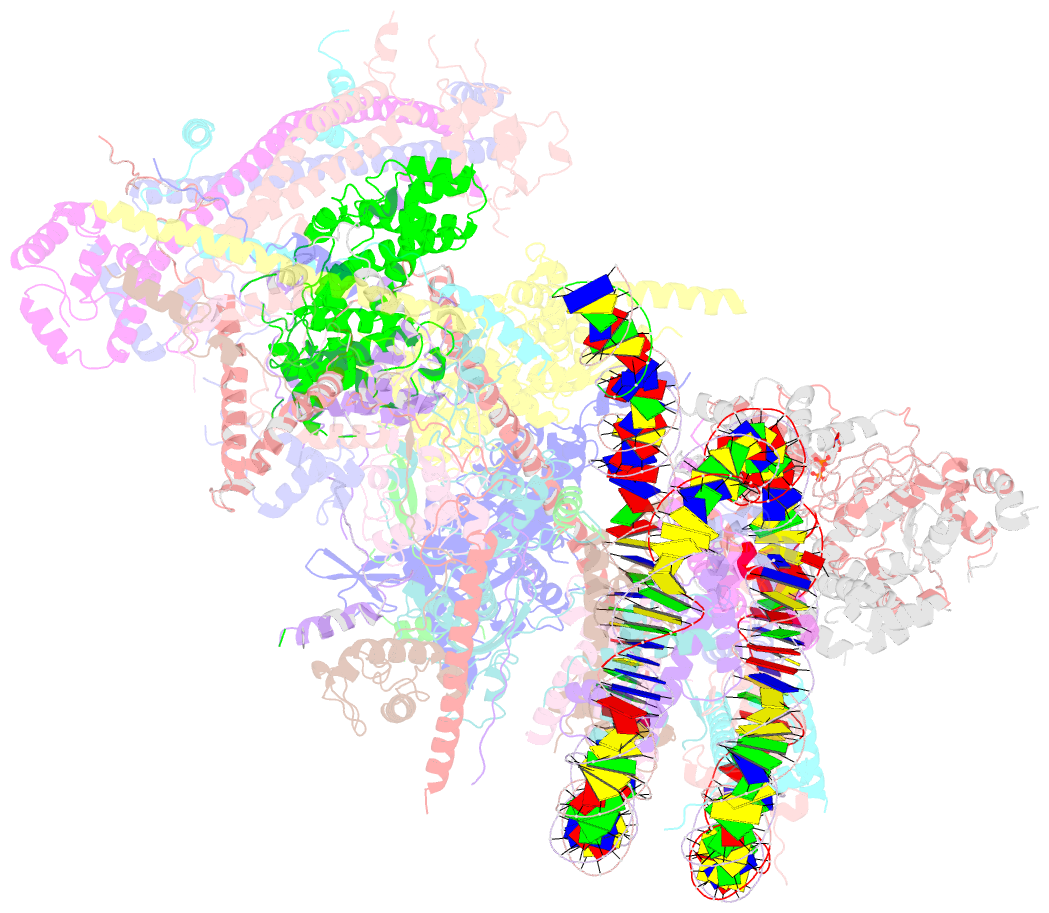
- Swi-snf nucleosome complex with adp-befx
- Reference
- Han Y, Reyes AA, Malik S, He Y (2020): "Cryo-EM structure of SWI/SNF complex bound to a nucleosome." Nature, 579, 452-455. doi: 10.1038/s41586-020-2087-1.
- Abstract
- The chromatin-remodelling complex SWI/SNF is highly conserved and has critical roles in various cellular processes, including transcription and DNA-damage repair1,2. It hydrolyses ATP to remodel chromatin structure by sliding and evicting histone octamers3-8, creating DNA regions that become accessible to other essential factors. However, our mechanistic understanding of the remodelling activity is hindered by the lack of a high-resolution structure of complexes from this family. Here we report the cryo-electron microscopy structure of Saccharomyces cerevisiae SWI/SNF bound to a nucleosome, at near-atomic resolution. In the structure, the actin-related protein (Arp) module is sandwiched between the ATPase and the rest of the complex, with the Snf2 helicase-SANT associated (HSA) domain connecting all modules. The body contains an assembly scaffold composed of conserved subunits Snf12 (also known as SMARCD or BAF60), Snf5 (also known as SMARCB1, BAF47 or INI1) and an asymmetric dimer of Swi3 (also known as SMARCC, BAF155 or BAF170). Another conserved subunit, Swi1 (also known as ARID1 or BAF250), resides in the core of SWI/SNF, acting as a molecular hub. We also observed interactions between Snf5 and the histones at the acidic patch, which could serve as an anchor during active DNA translocation. Our structure enables us to map and rationalize a subset of cancer-related mutations in the human SWI/SNF complex and to propose a model for how SWI/SNF recognizes and remodels the +1 nucleosome to generate nucleosome-depleted regions during gene activation9.