Summary information and primary citation
- PDB-id
- 6ycq; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- transcription
- Method
- X-ray (1.65 Å)
- Summary
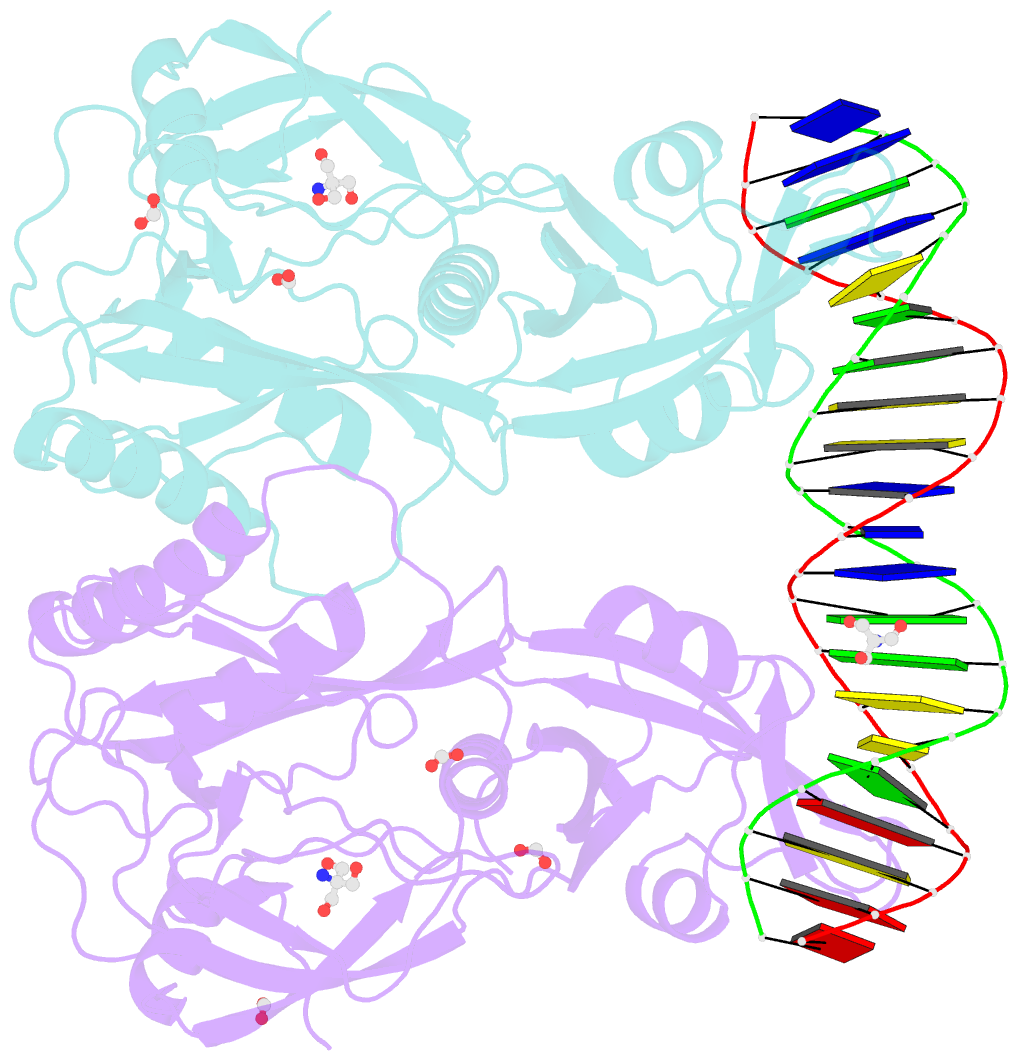
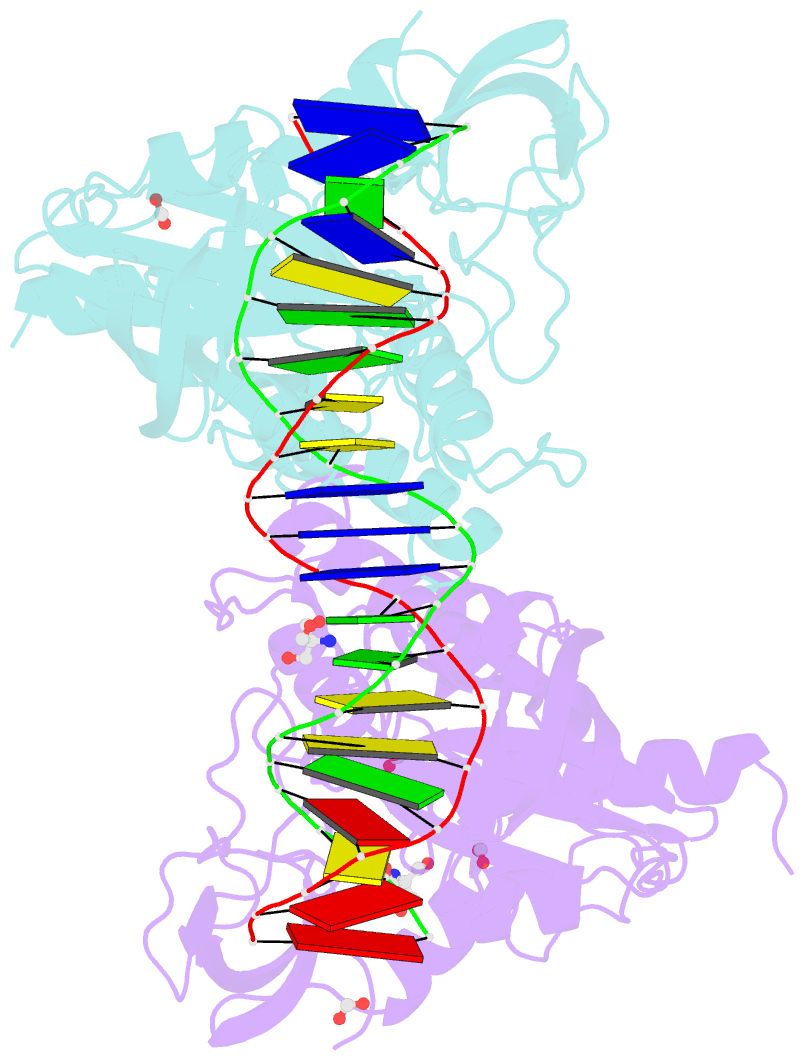
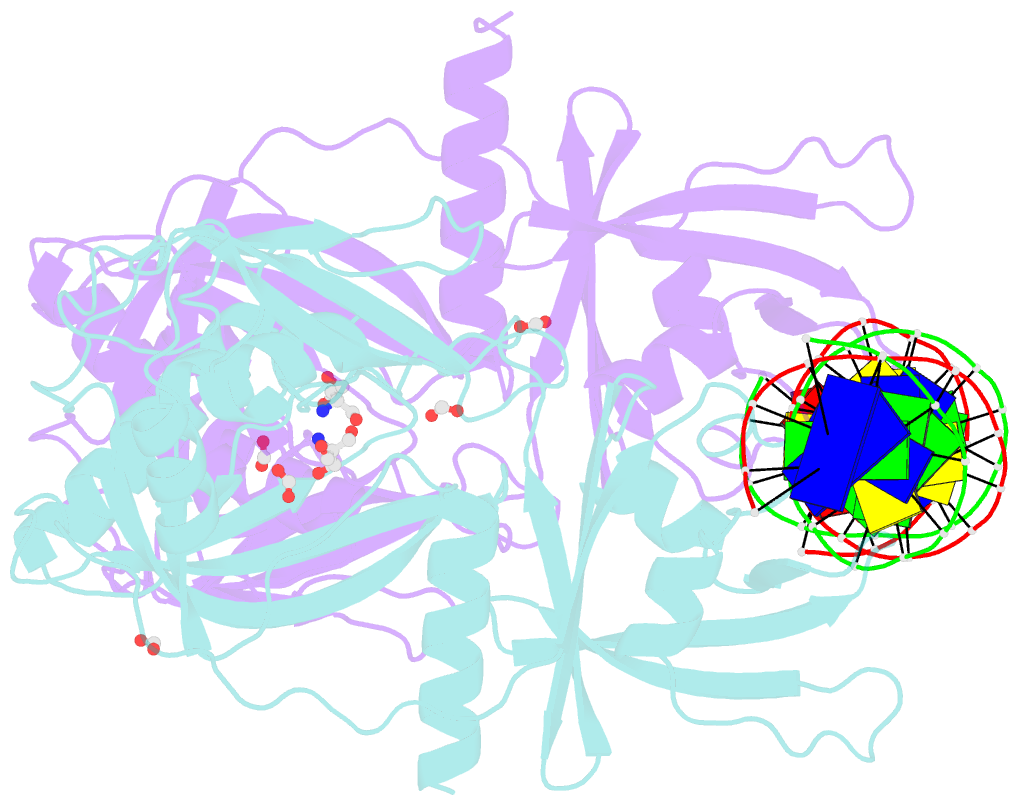
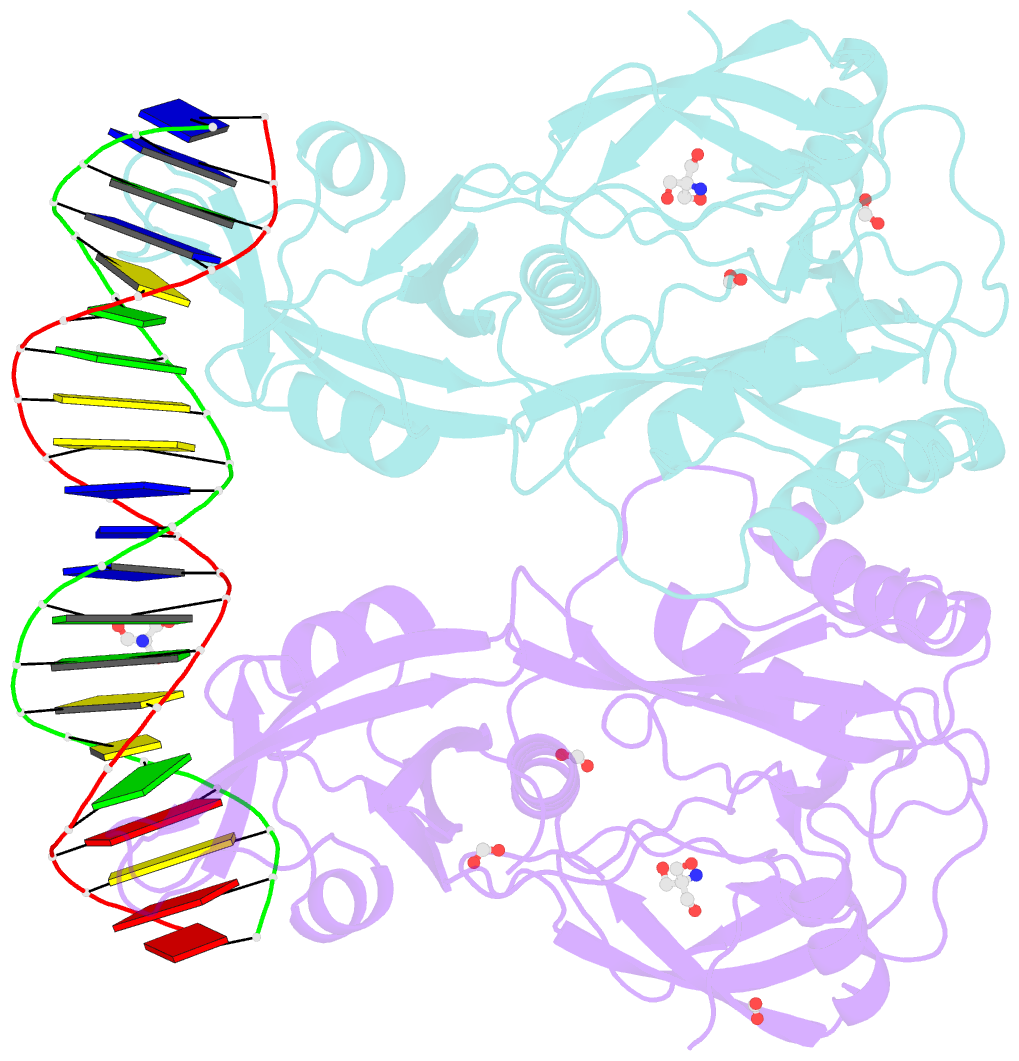
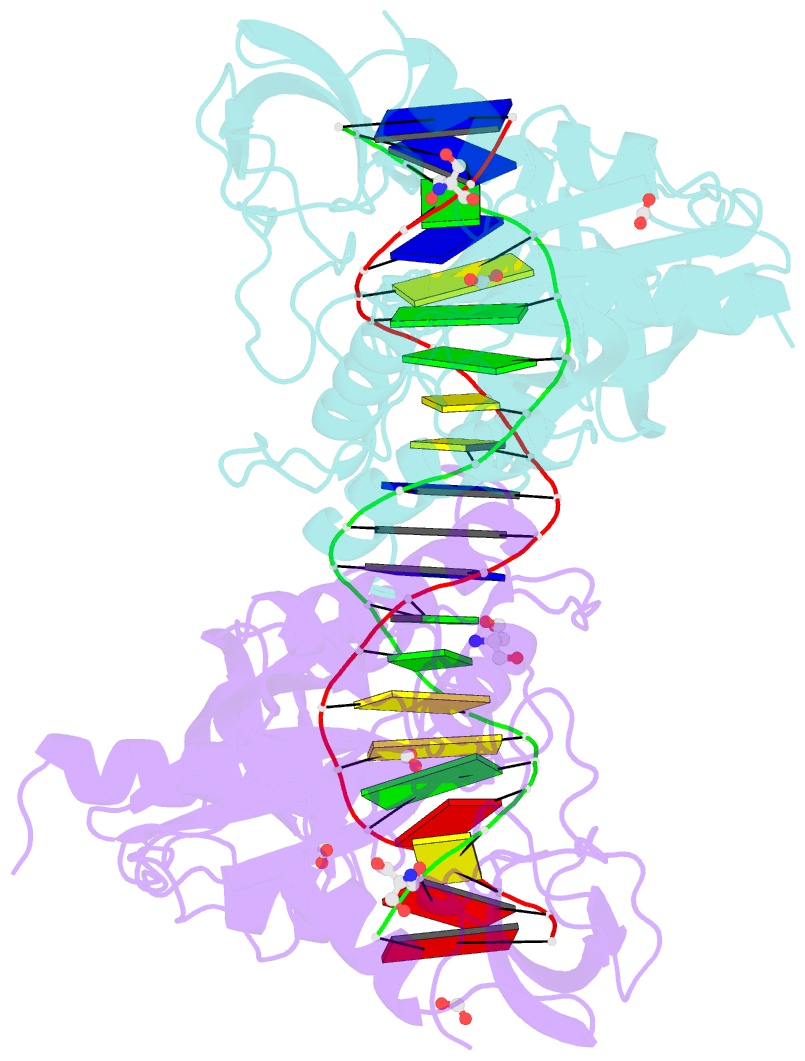
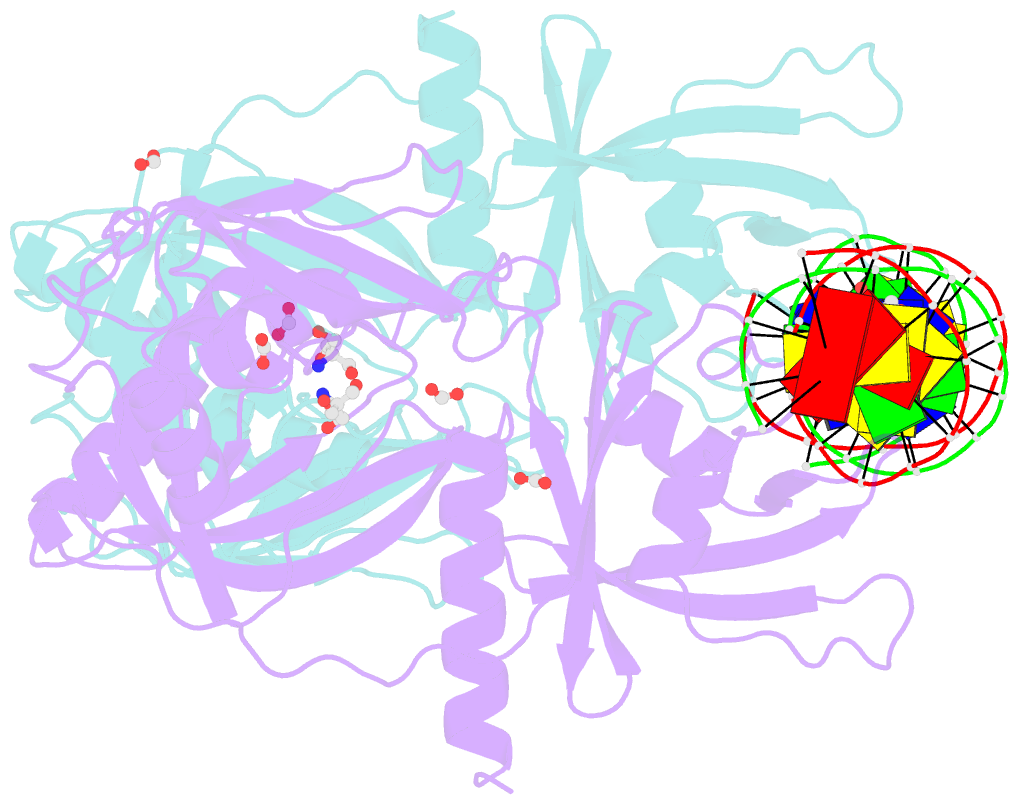
- Crystal structure of the DNA binding domain of arabidopsis thaliana auxin response factor 1 (atarf1) in complex with high affinity DNA
- Reference
- Freire-Rios A, Tanaka K, Crespo I, van der Wijk E, Sizentsova Y, Levitsky V, Lindhoud S, Fontana M, Hohlbein J, Boer DR, Mironova V, Weijers D (2020): "Architecture of DNA elements mediating ARF transcription factor binding and auxin-responsive gene expression in Arabidopsis ." Proc.Natl.Acad.Sci.USA, 117, 24557-24566. doi: 10.1073/pnas.2009554117.
- Abstract
- The hormone auxin controls many aspects of the plant life cycle by regulating the expression of thousands of genes. The transcriptional output of the nuclear auxin signaling pathway is determined by the activity of AUXIN RESPONSE transcription FACTORs (ARFs), through their binding to cis-regulatory elements in auxin-responsive genes. Crystal structures, in vitro, and heterologous studies have fueled a model in which ARF dimers bind with high affinity to distinctly spaced repeats of canonical AuxRE motifs. However, the relevance of this "caliper" model, and the mechanisms underlying the binding affinities in vivo, have remained elusive. Here we biochemically and functionally interrogate modes of ARF-DNA interaction. We show that a single additional hydrogen bond in Arabidopsis ARF1 confers high-affinity binding to individual DNA sites. We demonstrate the importance of AuxRE cooperativity within repeats in the Arabidopsis TMO5 and IAA11 promoters in vivo. Meta-analysis of transcriptomes further reveals strong genome-wide association of auxin response with both inverted (IR) and direct (DR) AuxRE repeats, which we experimentally validated. The association of these elements with auxin-induced up-regulation (DR and IR) or down-regulation (IR) was correlated with differential binding affinities of A-class and B-class ARFs, respectively, suggesting a mechanistic basis for the distinct activity of these repeats. Our results support the relevance of high-affinity binding of ARF transcription factors to uniquely spaced DNA elements in vivo, and suggest that differential binding affinities of ARF subfamilies underlie diversity in cis-element function.