Summary information and primary citation
- PDB-id
- 6zww; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- RNA binding protein
- Method
- X-ray (3.16 Å)
- Summary
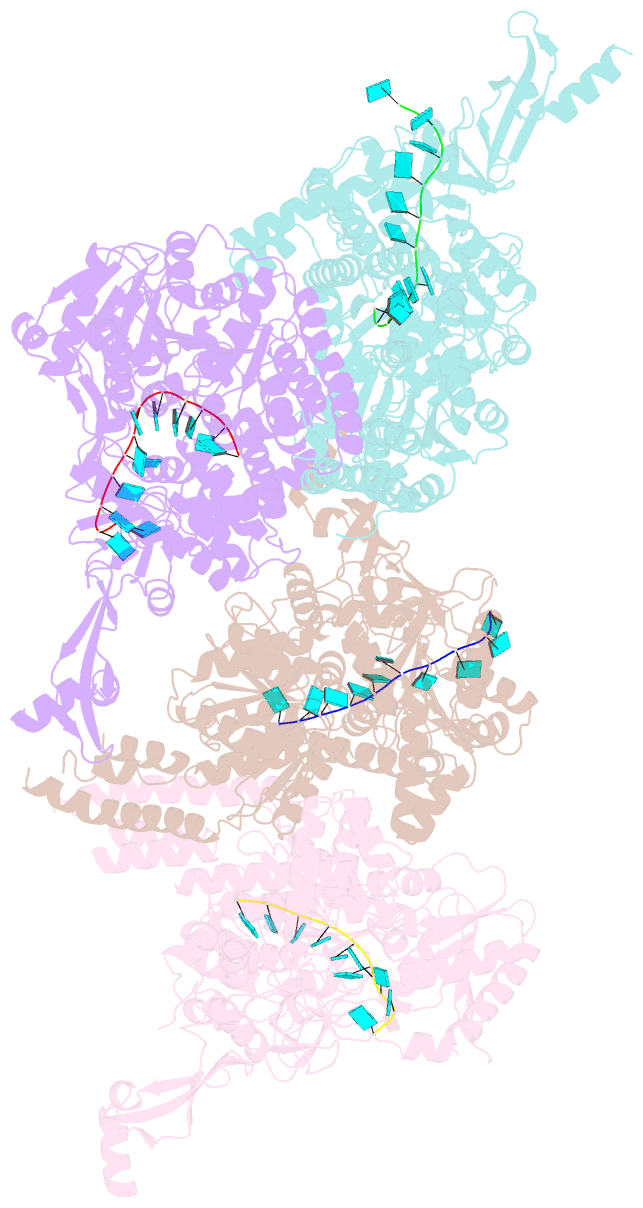
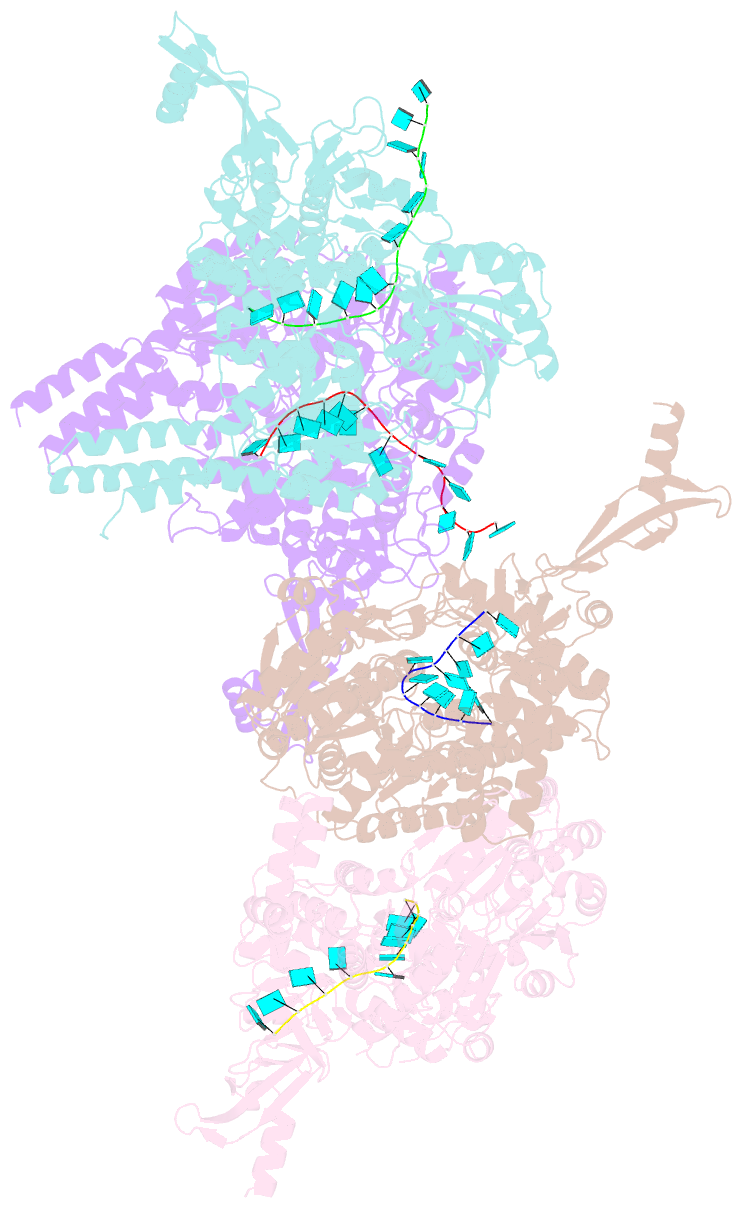
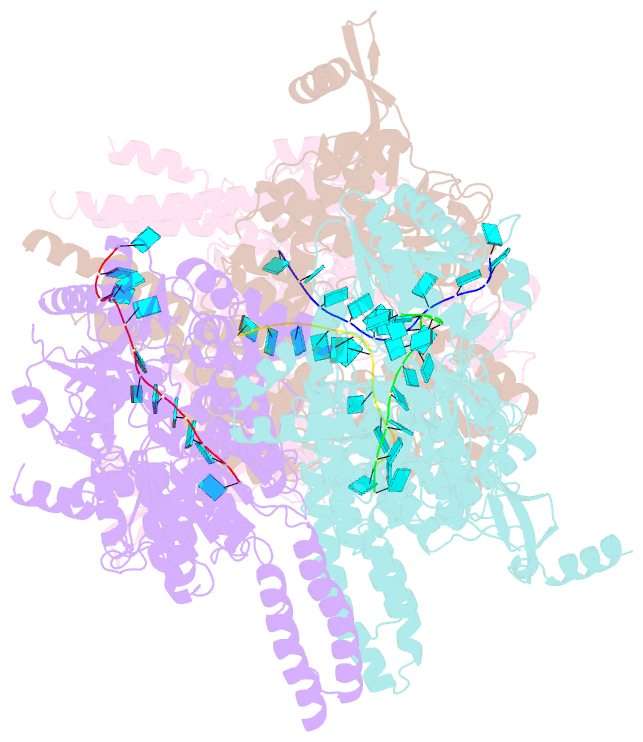
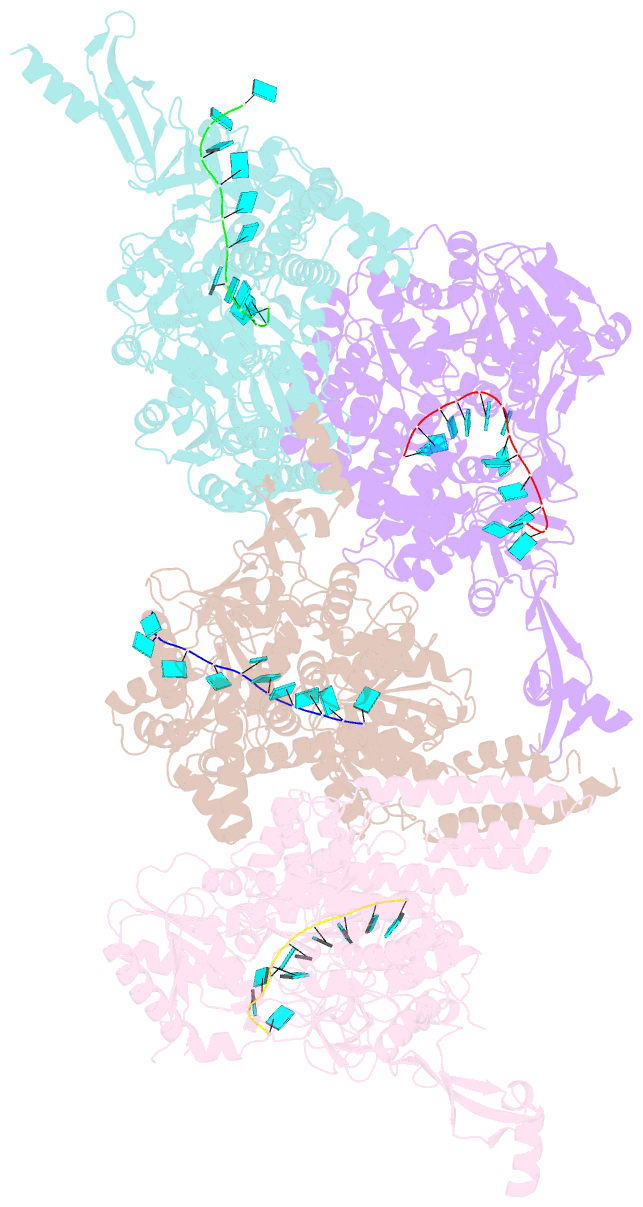
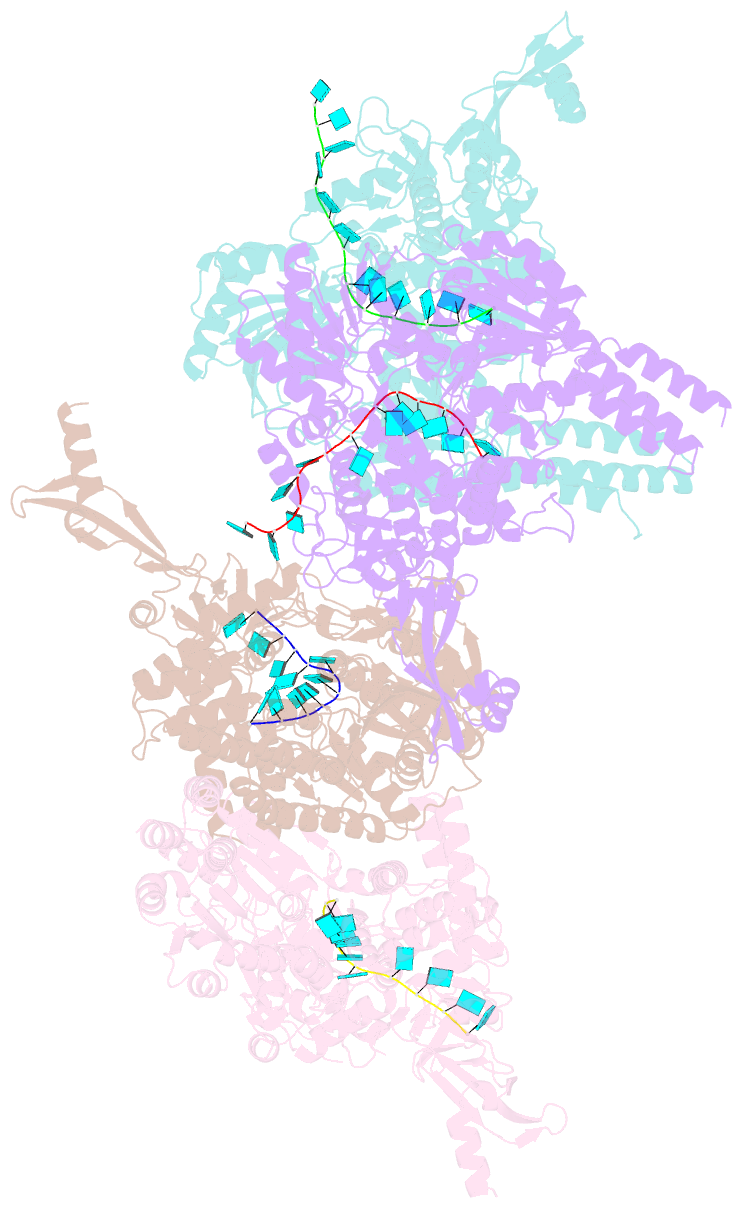
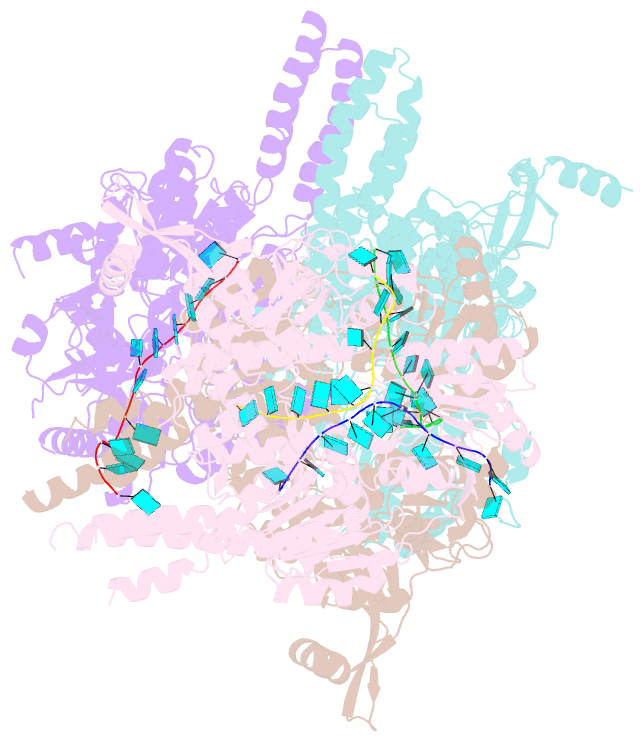
- Crystal structure of e. coli RNA helicase hrpa in complex with RNA
- Reference
- Grass LM, Wollenhaupt J, Barthel T, Parfentev I, Urlaub H, Loll B, Klauck E, Antelmann H, Wahl MC (2021): "Large-scale ratcheting in a bacterial DEAH/RHA-type RNA helicase that modulates antibiotics susceptibility." Proc.Natl.Acad.Sci.USA, 118. doi: 10.1073/pnas.2100370118.
- Abstract
- Many bacteria harbor RNA-dependent nucleoside-triphosphatases of the DEAH/RHA family, whose molecular mechanisms and cellular functions are poorly understood. Here, we show that the Escherichia coli DEAH/RHA protein, HrpA, is an ATP-dependent 3 to 5' RNA helicase and that the RNA helicase activity of HrpA influences bacterial survival under antibiotics treatment. Limited proteolysis, crystal structure analysis, and functional assays showed that HrpA contains an N-terminal DEAH/RHA helicase cassette preceded by a unique N-terminal domain and followed by a large C-terminal region that modulates the helicase activity. Structures of an expanded HrpA helicase cassette in the apo and RNA-bound states in combination with cross-linking/mass spectrometry revealed ratchet-like domain movements upon RNA engagement, much more pronounced than hitherto observed in related eukaryotic DEAH/RHA enzymes. Structure-based functional analyses delineated transient interdomain contact sites that support substrate loading and unwinding, suggesting that similar conformational changes support RNA translocation. Consistently, modeling studies showed that analogous dynamic intramolecular contacts are not possible in the related but helicase-inactive RNA-dependent nucleoside-triphosphatase, HrpB. Our results indicate that HrpA may be an interesting target to interfere with bacterial tolerance toward certain antibiotics and suggest possible interfering strategies.