Summary information and primary citation
- PDB-id
- 7abs; SNAP-derived features in text and JSON formats;
DNAproDB
- Class
- lyase
- Method
- X-ray (1.97 Å)
- Summary
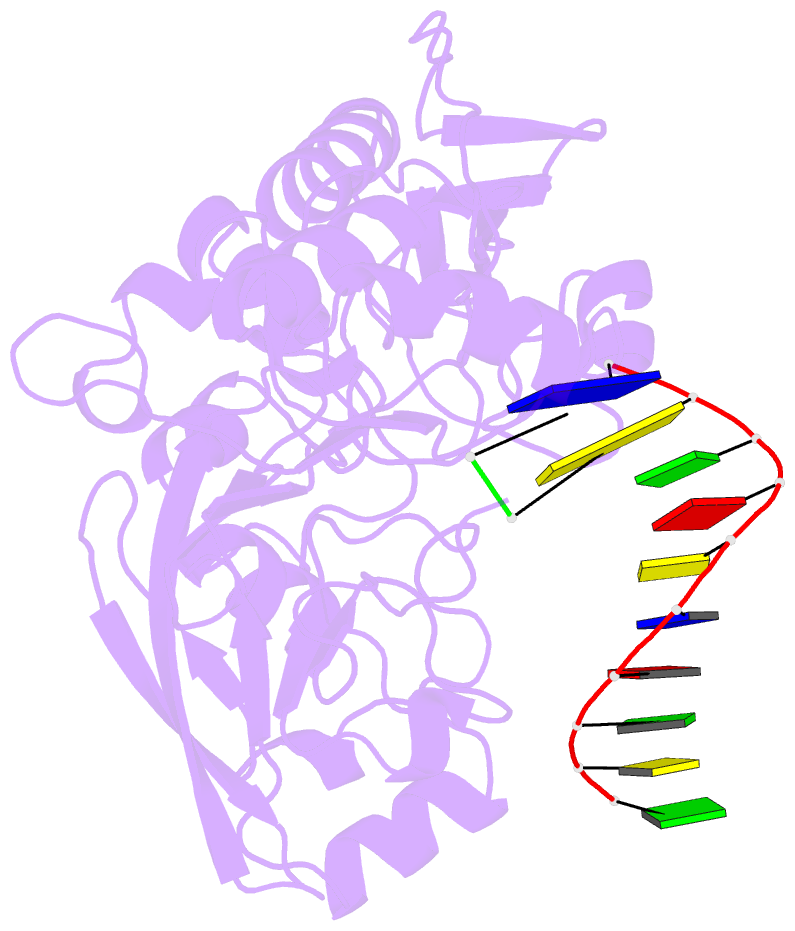
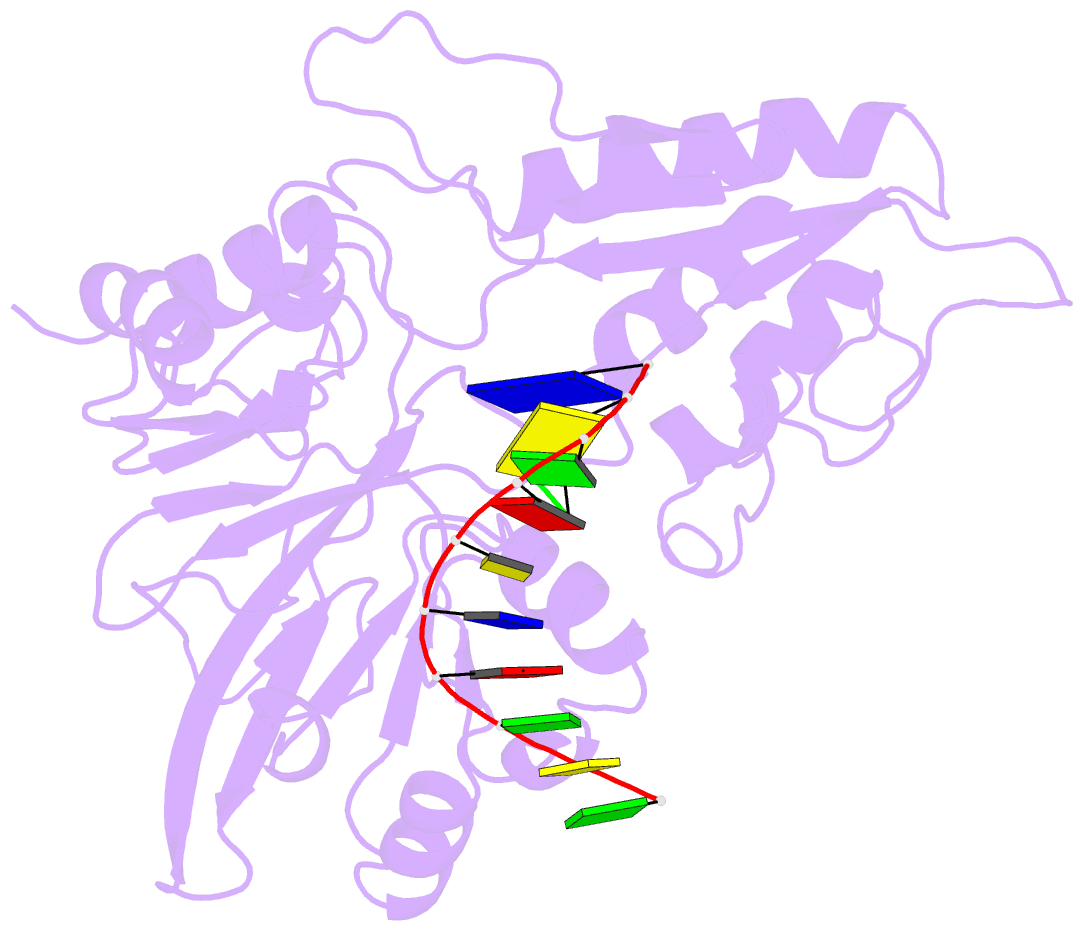
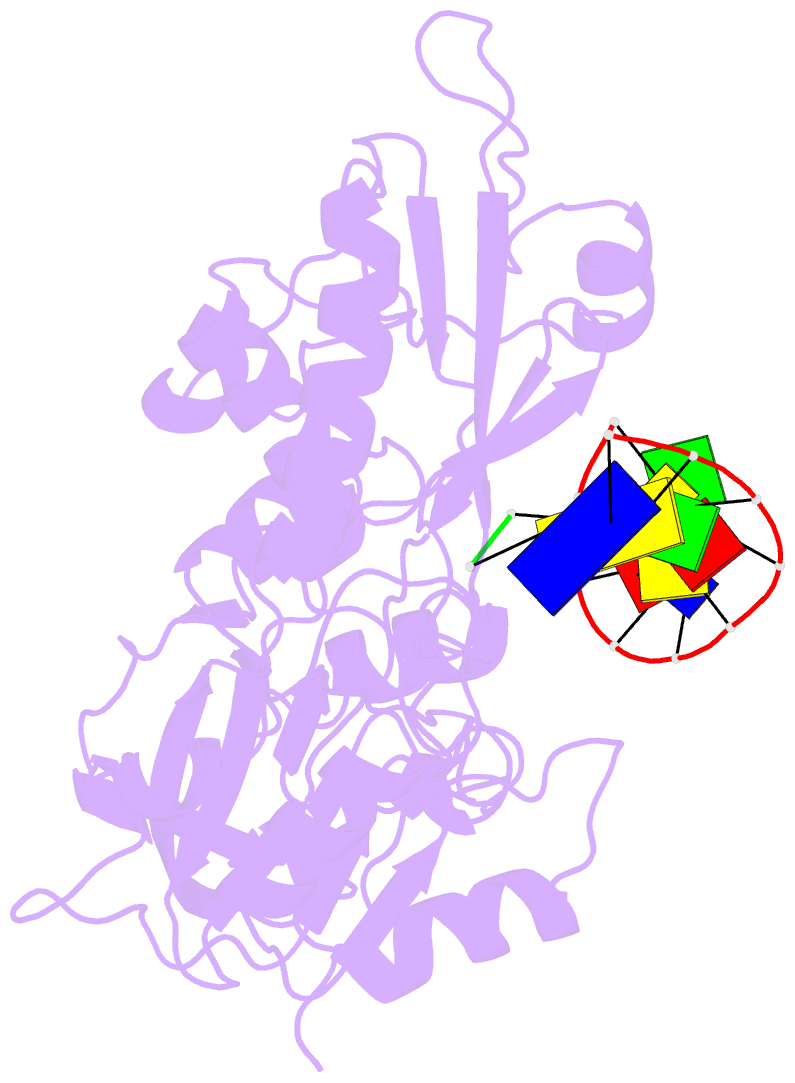
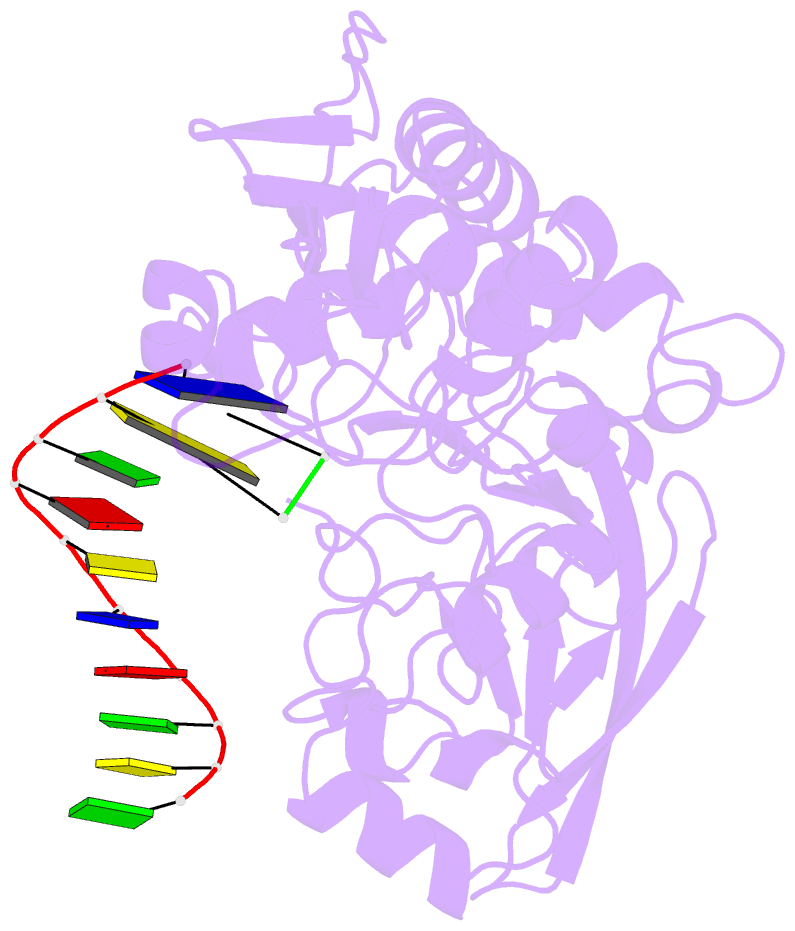
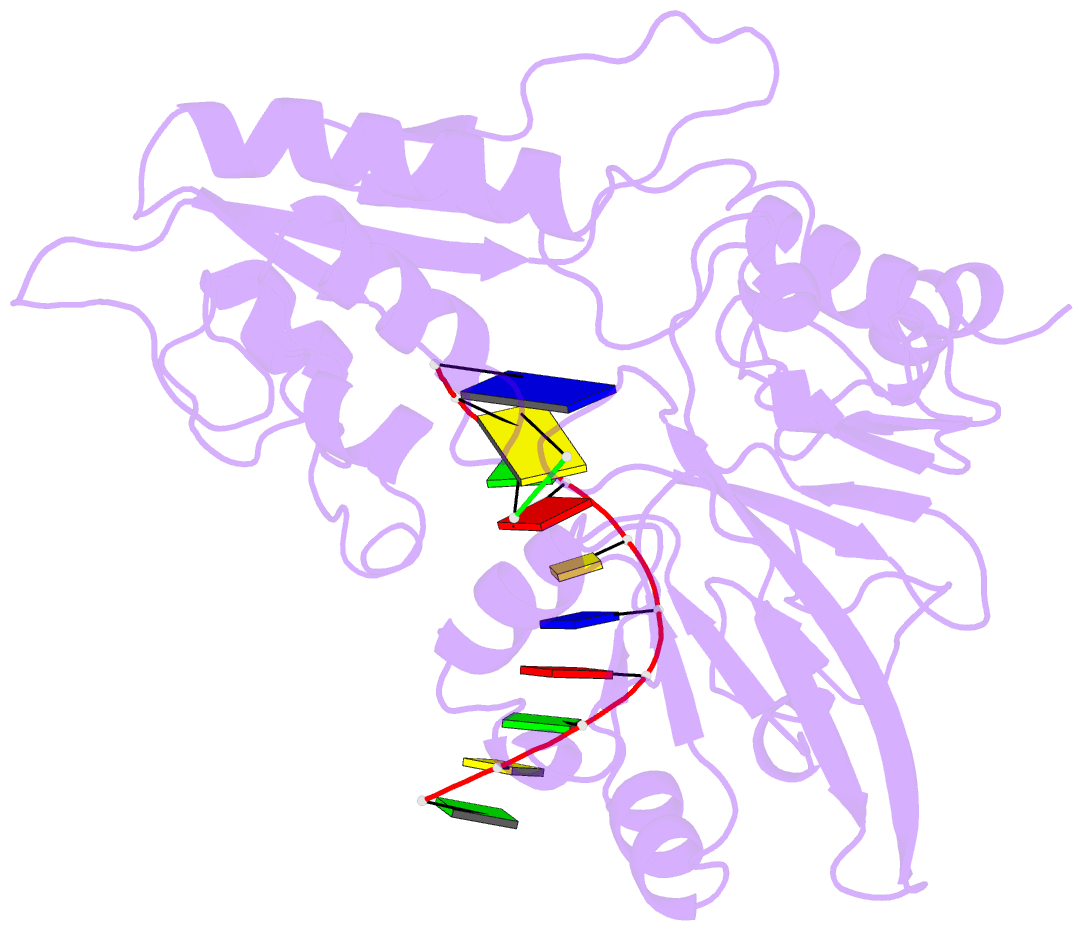
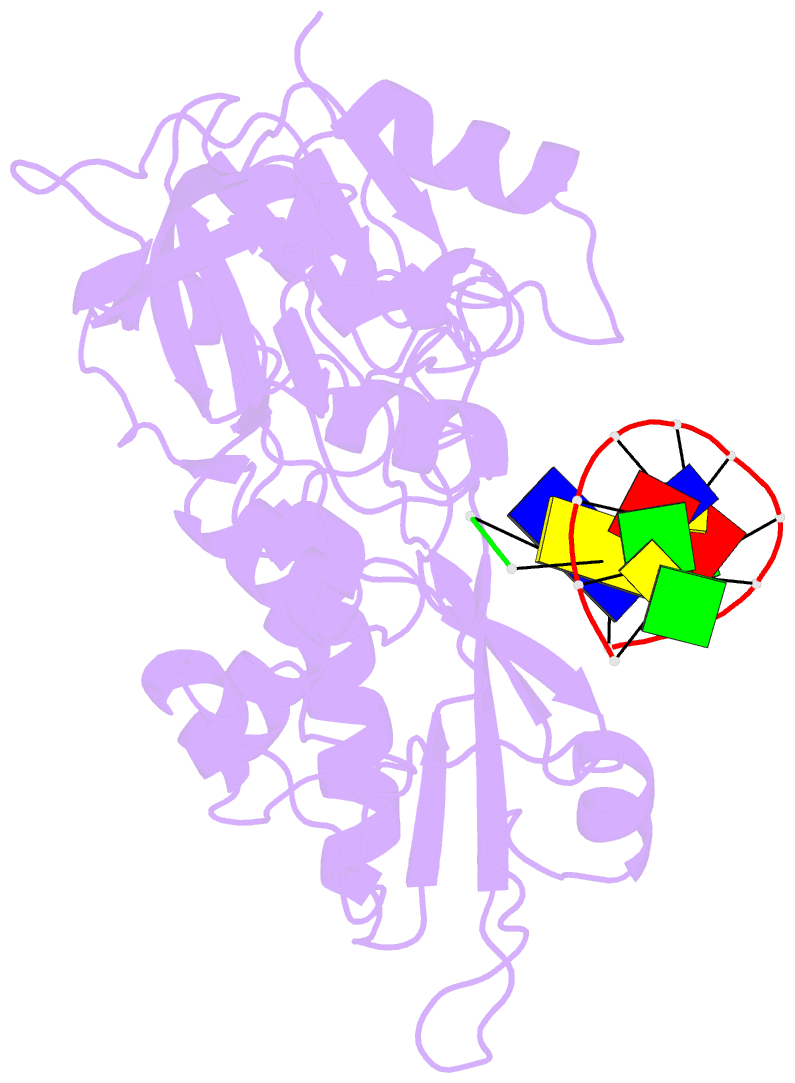
- Structure of human dclre1c-artemis in complex with DNA - re-evaluation of 6wo0
- Reference
- Yosaatmadja Y, Baddock HT, Newman JA, Bielinski M, Gavard AE, Mukhopadhyay SMM, Dannerfjord AA, Schofield CJ, McHugh PJ, Gileadi O (2021): "Structural and mechanistic insights into the Artemis endonuclease and strategies for its inhibition." Nucleic Acids Res., 49, 9310-9326. doi: 10.1093/nar/gkab693.
- Abstract
- Artemis (SNM1C/DCLRE1C) is an endonuclease that plays a key role in development of B- and T-lymphocytes and in dsDNA break repair by non-homologous end-joining (NHEJ). Artemis is phosphorylated by DNA-PKcs and acts to open DNA hairpin intermediates generated during V(D)J and class-switch recombination. Artemis deficiency leads to congenital radiosensitive severe acquired immune deficiency (RS-SCID). Artemis belongs to a superfamily of nucleases containing metallo-β-lactamase (MBL) and β-CASP (CPSF-Artemis-SNM1-Pso2) domains. We present crystal structures of the catalytic domain of wildtype and variant forms of Artemis, including one causing RS-SCID Omenn syndrome. The catalytic domain of the Artemis has similar endonuclease activity to the phosphorylated full-length protein. Our structures help explain the predominantly endonucleolytic activity of Artemis, which contrasts with the predominantly exonuclease activity of the closely related SNM1A and SNM1B MBL fold nucleases. The structures reveal a second metal binding site in its β-CASP domain unique to Artemis, which is amenable to inhibition by compounds including ebselen. By combining our structural data with that from a recently reported Artemis structure, we were able model the interaction of Artemis with DNA substrates. The structures, including one of Artemis with the cephalosporin ceftriaxone, will help enable the rational development of selective SNM1 nuclease inhibitors.