Summary information and primary citation
- PDB-id
-
7dbp;
DSSR-derived features in text and
JSON formats; DNAproDB
- Class
- structural protein
- Method
- cryo-EM (4.5 Å)
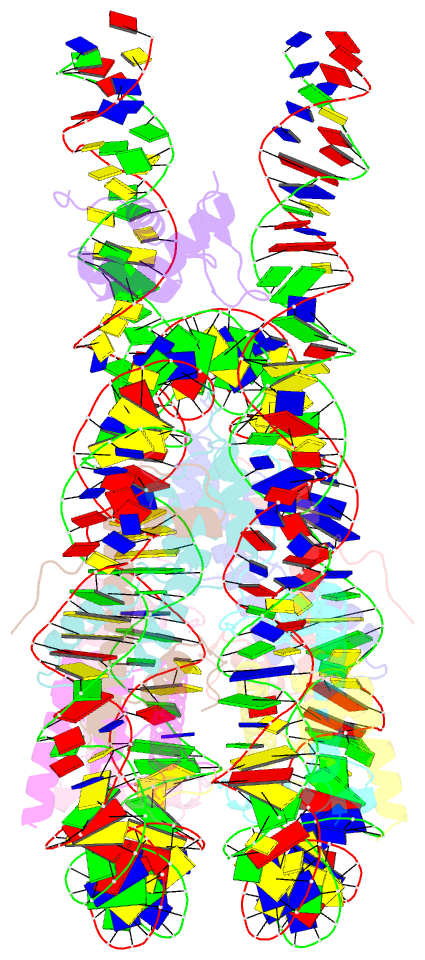
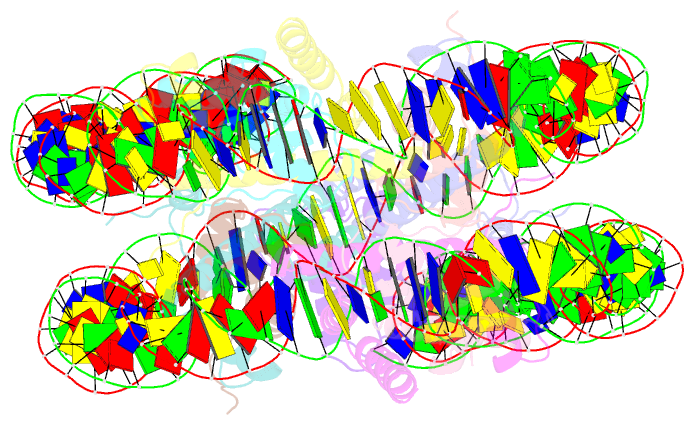
- Summary
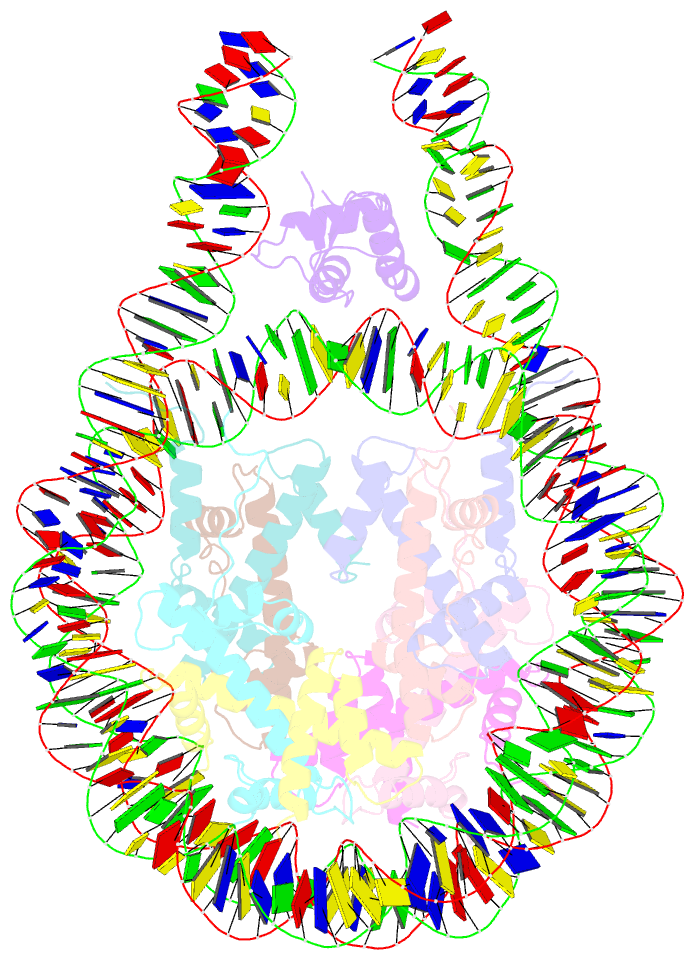
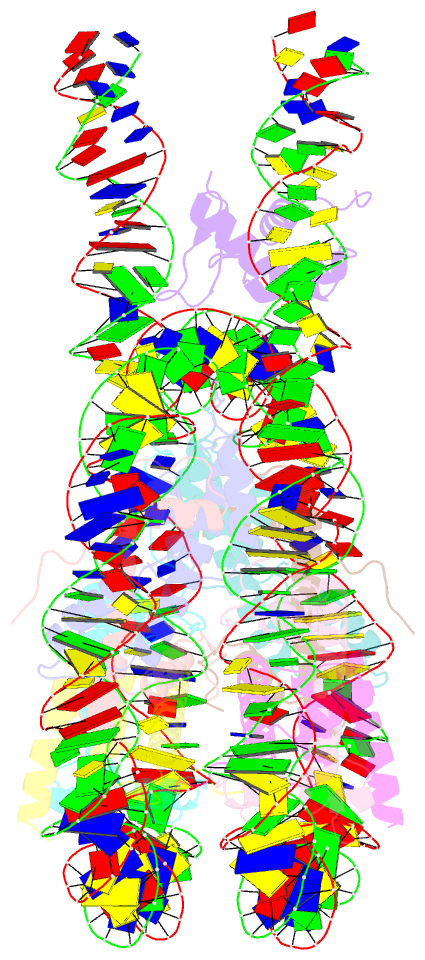
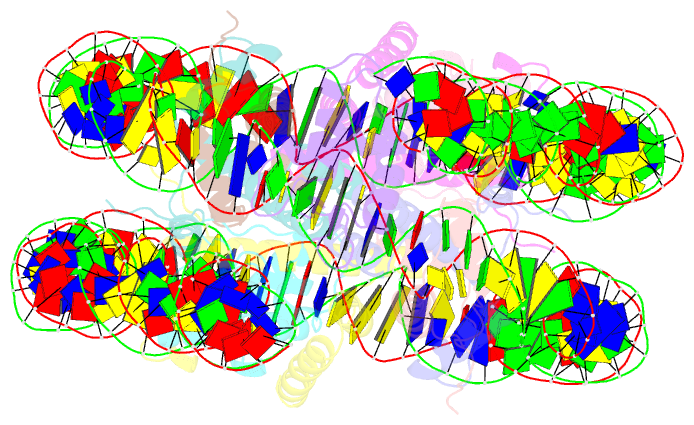
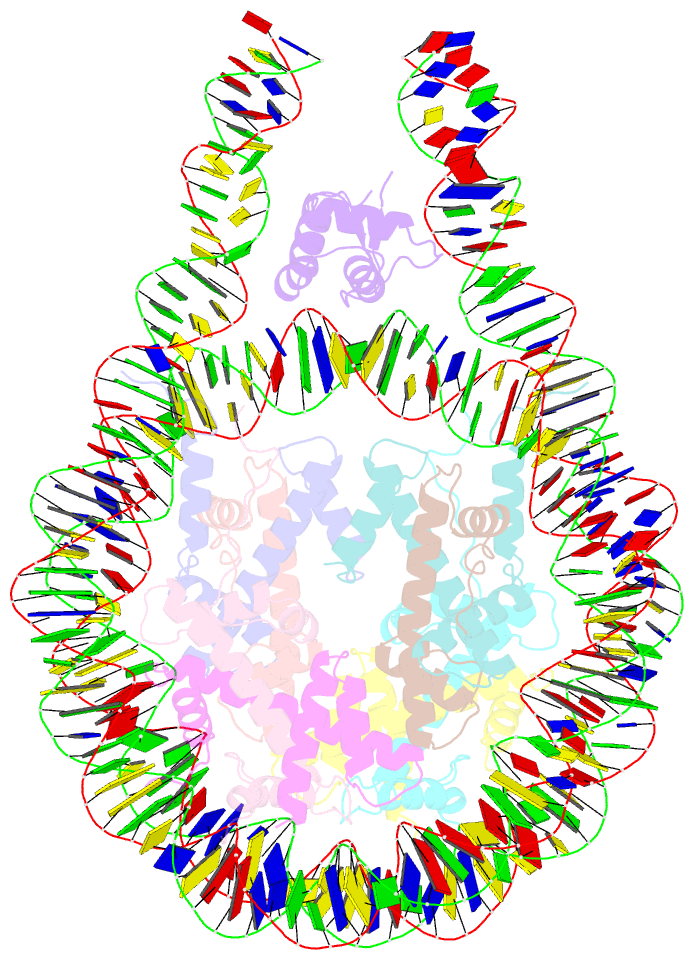
- Linker histone defines structure and self-association
behaviour of the 177 bp human chromosome
- Reference
-
Wang S, Vogirala VK, Soman A, Berezhnoy NV, Liu ZB, Wong
ASW, Korolev N, Su CJ, Sandin S, Nordenskiold L (2021):
"Linker
histone defines structure and self-association behaviour
of the 177 bp human chromatosome." Sci Rep,
11, 380. doi: 10.1038/s41598-020-79654-8.
- Abstract
- Linker histones play essential roles in the regulation
and maintenance of the dynamic chromatin structure of
higher eukaryotes. The influence of human histone H1.0 on
the nucleosome structure and biophysical properties of the
resulting chromatosome were investigated and compared with
the 177-bp nucleosome using Cryo-EM and SAXS. The
4.5 Å Cryo-EM chromatosome structure showed that the
linker histone binds at the nucleosome dyad interacting
with both linker DNA arms but in a tilted manner leaning
towards one of the linker sides. The chromatosome is
laterally compacted and rigid in the dyad and linker DNA
area, in comparison with the nucleosome where linker DNA
region is more flexible and displays structural
variability. In solution, the chromatosomes appear slightly
larger than the nucleosomes, with the volume increase
compared to the bound linker histone, according to solution
SAXS measurements. SAXS X-ray diffraction characterisation
of Mg-precipitated samples showed that the different shapes
of the 177 chromatosome enabled the formation of a highly
ordered lamello-columnar phase when precipitated by
Mg<sub>2+</sub>, indicating the influence of
linker histone on the nucleosome stacking. The biological
significance of linker histone, therefore, may be affected
by the change in the polyelectrolyte and DNA conformation
properties of the chromatosomes, in comparison to
nucleosomes.